![[OK]](Icons/tick.png) Basic Statistics
Basic Statistics
| Measure | Value |
|---|---|
| Filename | LW199_GTCCGC_L003_R1_001.fastq.gz |
| File type | Conventional base calls |
| Encoding | Sanger / Illumina 1.9 |
| Total Sequences | 10364530 |
| Filtered Sequences | 0 |
| Sequence length | 51 |
| %GC | 41 |
![[OK]](Icons/tick.png) Per base sequence quality
Per base sequence quality
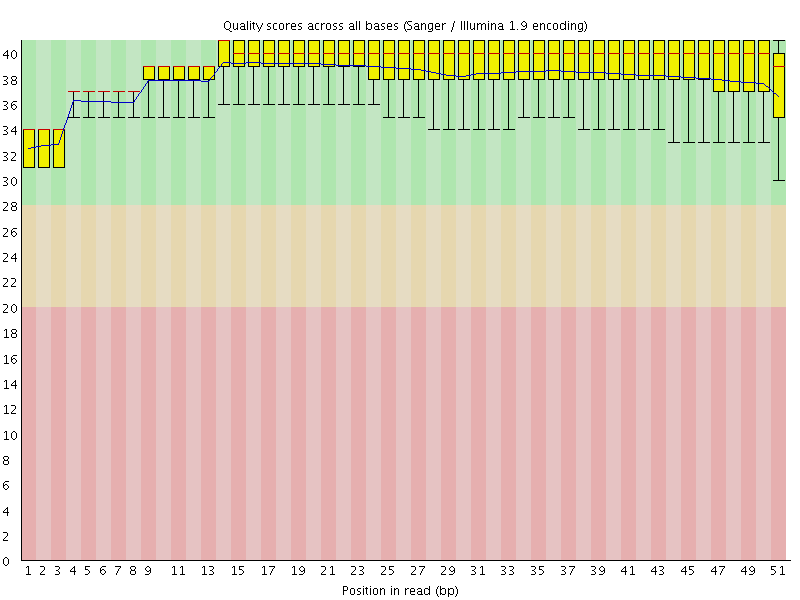
![[OK]](Icons/tick.png) Per sequence quality scores
Per sequence quality scores
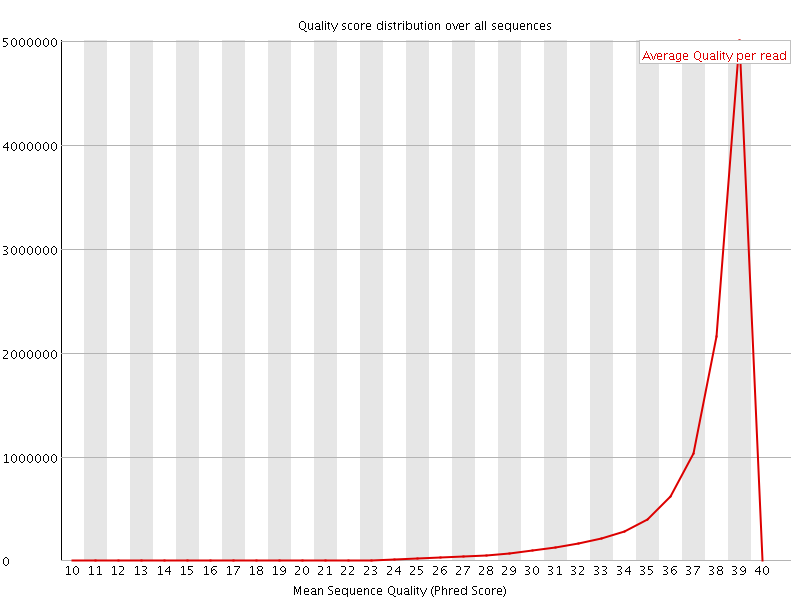
![[WARN]](Icons/warning.png) Per base sequence content
Per base sequence content
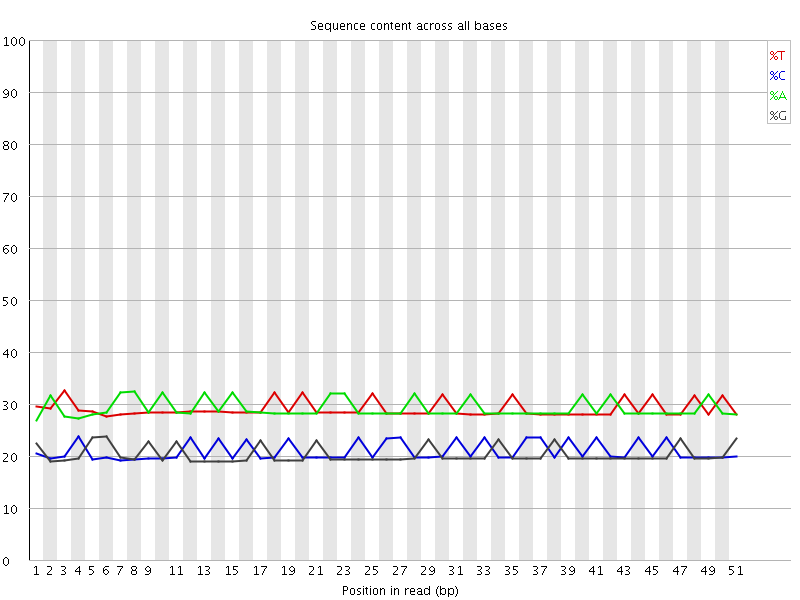
![[OK]](Icons/tick.png) Per base GC content
Per base GC content
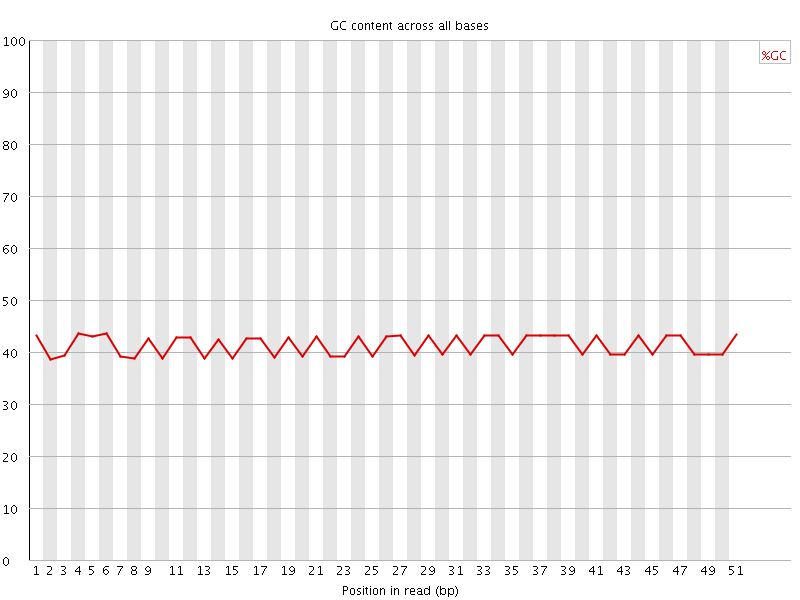
![[OK]](Icons/tick.png) Per sequence GC content
Per sequence GC content
![[OK]](Icons/tick.png) Per base N content
Per base N content
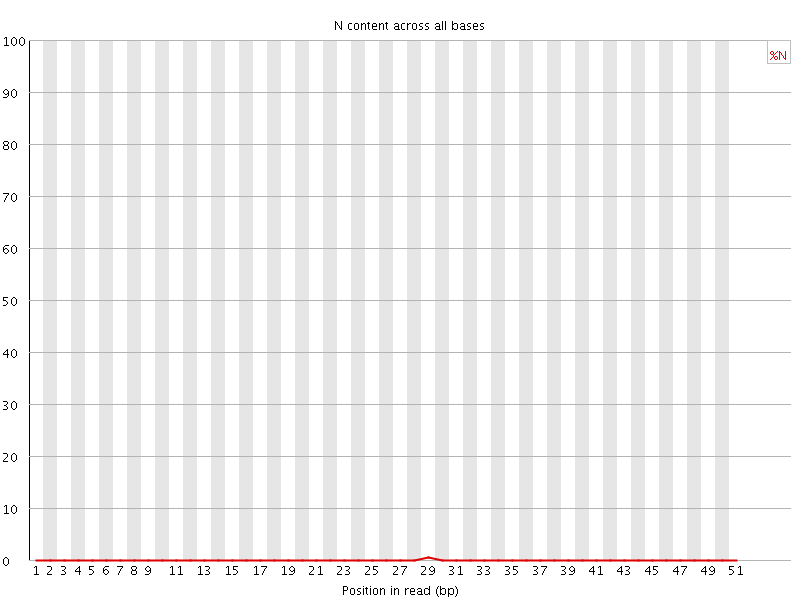
![[OK]](Icons/tick.png) Sequence Length Distribution
Sequence Length Distribution
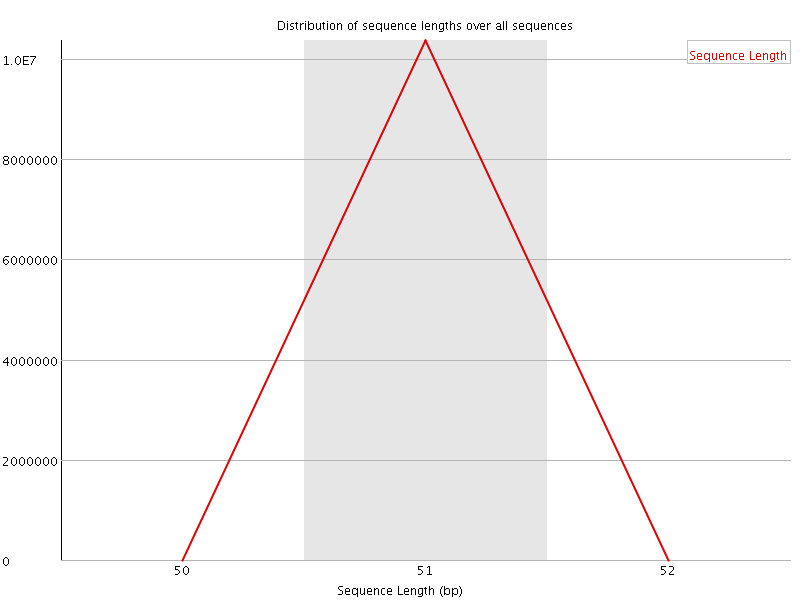
![[OK]](Icons/tick.png) Sequence Duplication Levels
Sequence Duplication Levels
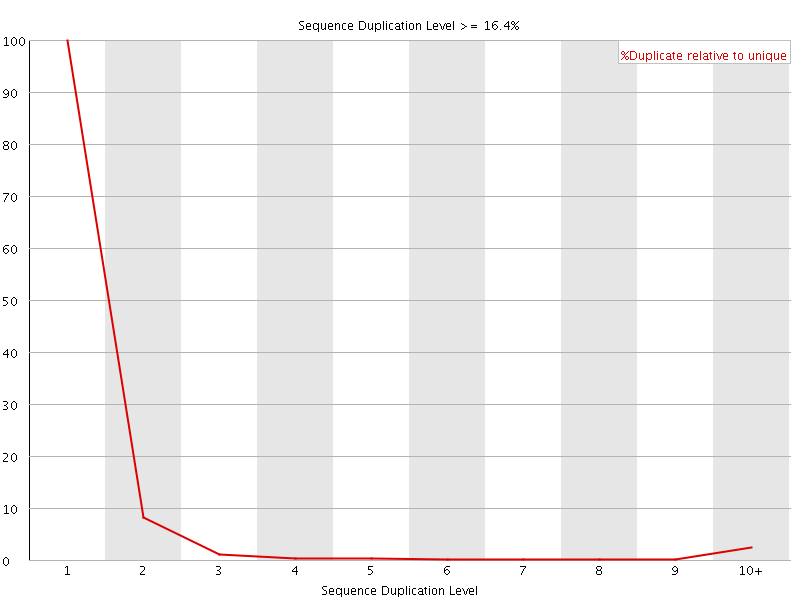
![[FAIL]](Icons/error.png) Overrepresented sequences
Overrepresented sequences
| Sequence | Count | Percentage | Possible Source |
|---|---|---|---|
| GATCGGAAGAGCACACGTCTGAACTCCAGTCACGTCCGCACATCTCGTATG | 334097 | 3.2234650292873868 | TruSeq Adapter, Index 1 (97% over 36bp) |
![[WARN]](Icons/warning.png) Kmer Content
Kmer Content
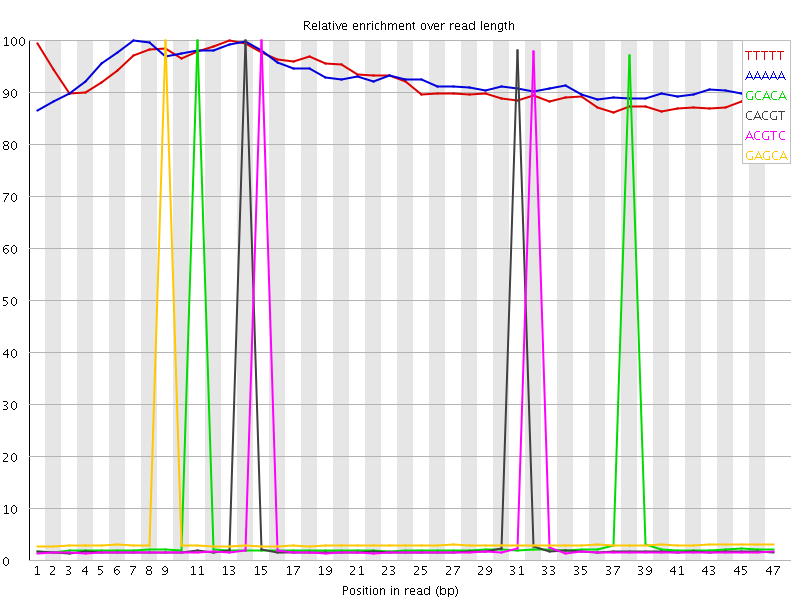
| Sequence | Count | Obs/Exp Overall | Obs/Exp Max | Max Obs/Exp Position |
|---|---|---|---|---|
| TTTTT | 3636305 | 3.5385218 | 3.8437088 | 13 |
| AAAAA | 3624585 | 3.4453661 | 3.718022 | 7 |
| GCACA | 1165435 | 3.068936 | 49.355145 | 11 |
| CACGT | 1106305 | 2.926925 | 49.434994 | 14 |
| ACGTC | 1084705 | 2.8697786 | 49.28051 | 15 |
| GAGCA | 961355 | 2.6108298 | 51.646446 | 9 |
| GGAAG | 928170 | 2.5996644 | 53.9132 | 5 |
| GAAGA | 1300995 | 2.5417206 | 38.23714 | 6 |
| CTCCA | 974255 | 2.499277 | 47.62727 | 24 |
| TCCGC | 673260 | 2.4760654 | 66.27257 | 35 |
| TCCAG | 926430 | 2.4510338 | 49.302223 | 25 |
| CGGAA | 893085 | 2.4254236 | 51.958218 | 4 |
| CCGCA | 652400 | 2.3881211 | 66.172386 | 36 |
| CTGAA | 1230320 | 2.3415973 | 36.53896 | 19 |
| TCTCG | 848950 | 2.256606 | 49.340023 | 43 |
| TCGGA | 807610 | 2.2036033 | 52.012238 | 3 |
| GTCCG | 580525 | 2.2018878 | 68.25729 | 34 |
| CGTCC | 594005 | 2.1845872 | 66.3313 | 33 |
| AGAGC | 794320 | 2.1571994 | 51.20716 | 8 |
| CGCAC | 585880 | 2.1446235 | 65.82079 | 37 |
| AGCAC | 809255 | 2.1310089 | 49.496098 | 10 |
| GATCG | 771980 | 2.106385 | 51.650932 | 1 |
| ATCGG | 766100 | 2.0903409 | 51.836075 | 2 |
| CACAC | 786925 | 2.0092692 | 47.70746 | 12 |
| TCTGA | 1032905 | 1.9751107 | 36.30529 | 18 |
| CCAGT | 740645 | 1.9595069 | 48.691864 | 26 |
| CTCGT | 733010 | 1.9484241 | 48.914856 | 44 |
| GTCTG | 704115 | 1.9302437 | 51.125347 | 17 |
| CGTCT | 725235 | 1.9277573 | 49.47733 | 16 |
| AAGAG | 973720 | 1.9023318 | 37.45284 | 7 |
| GTCAC | 709920 | 1.8782185 | 49.12649 | 29 |
| TCACG | 695815 | 1.8409013 | 48.82636 | 30 |
| TGAAC | 950315 | 1.8086798 | 35.89536 | 20 |
| ACTCC | 702635 | 1.8024845 | 47.161396 | 23 |
| ACACG | 682995 | 1.7985287 | 49.159794 | 13 |
| CAGTC | 667785 | 1.7667431 | 48.793804 | 27 |
| CATCT | 915685 | 1.6977829 | 34.16056 | 41 |
| ATCTC | 910300 | 1.6877985 | 34.389168 | 42 |
| GAACT | 874875 | 1.6650994 | 35.71342 | 21 |
| AACTC | 896685 | 1.6547755 | 34.45033 | 22 |
| CACAT | 877370 | 1.6191308 | 33.852093 | 39 |
| ACATC | 853850 | 1.5757262 | 33.855118 | 40 |
| AGTCA | 785935 | 1.4958249 | 35.43502 | 28 |
| TCGTA | 734435 | 1.4043792 | 35.19042 | 45 |
| GTATG | 654030 | 1.2898037 | 36.004116 | 47 |
| CGTAT | 673105 | 1.2871046 | 35.041855 | 46 |