![[OK]](Icons/tick.png) Basic Statistics
Basic Statistics
| Measure | Value |
|---|---|
| Filename | LW197_CCGTCC_L003_R1_001.fastq.gz |
| File type | Conventional base calls |
| Encoding | Sanger / Illumina 1.9 |
| Total Sequences | 9839031 |
| Filtered Sequences | 0 |
| Sequence length | 51 |
| %GC | 45 |
![[OK]](Icons/tick.png) Per base sequence quality
Per base sequence quality
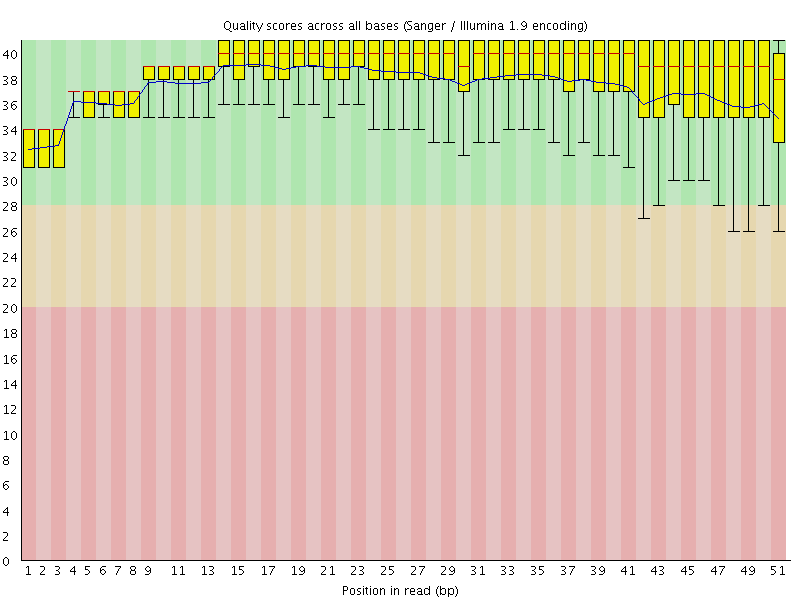
![[OK]](Icons/tick.png) Per sequence quality scores
Per sequence quality scores
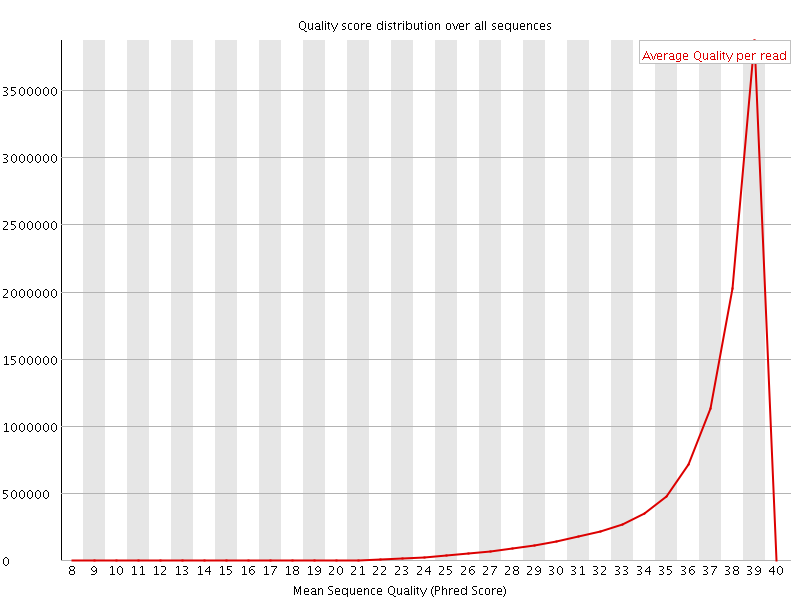
![[FAIL]](Icons/error.png) Per base sequence content
Per base sequence content
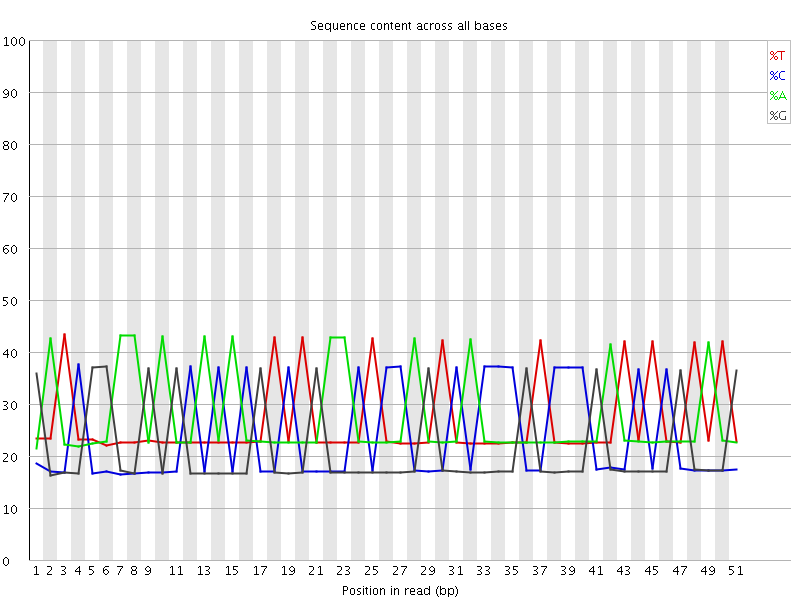
![[FAIL]](Icons/error.png) Per base GC content
Per base GC content
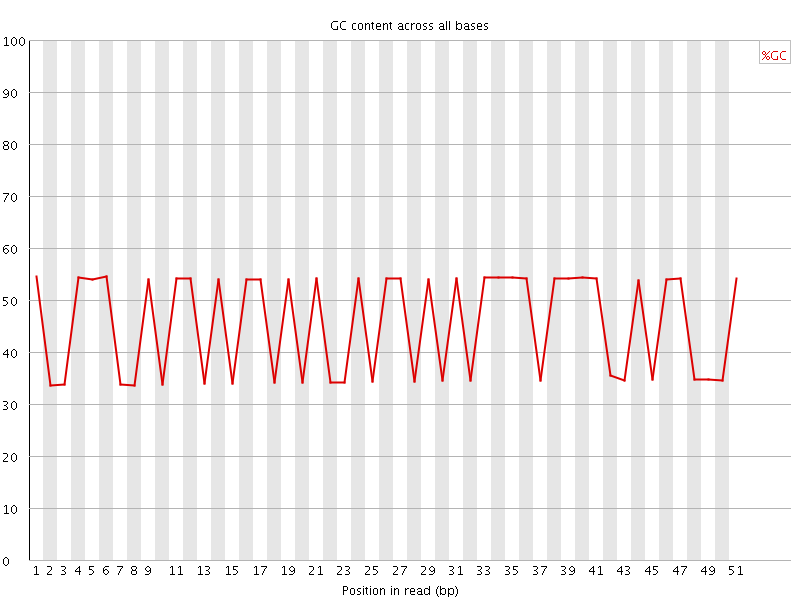
![[FAIL]](Icons/error.png) Per sequence GC content
Per sequence GC content
![[OK]](Icons/tick.png) Per base N content
Per base N content
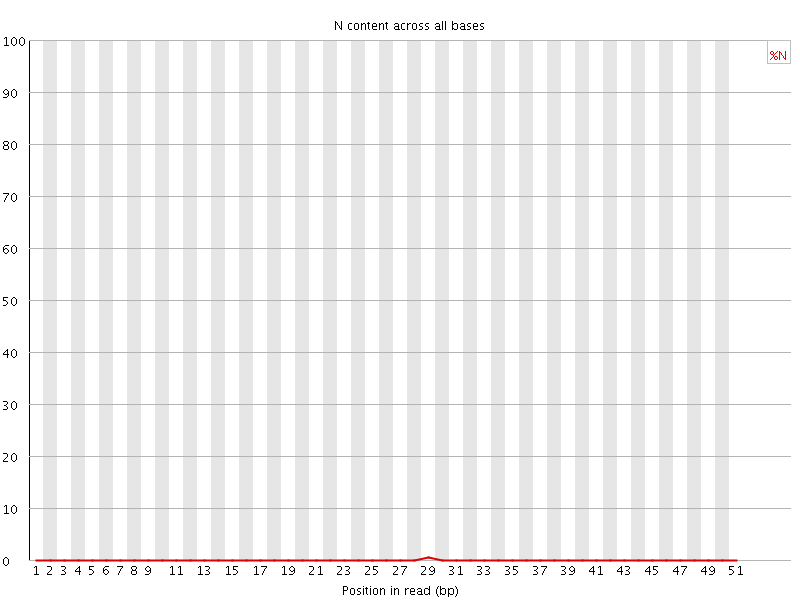
![[OK]](Icons/tick.png) Sequence Length Distribution
Sequence Length Distribution
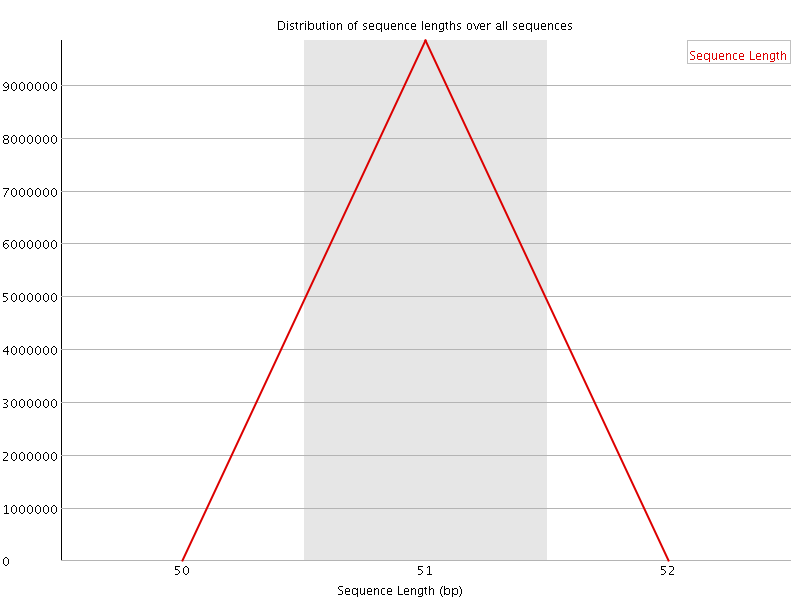
![[WARN]](Icons/warning.png) Sequence Duplication Levels
Sequence Duplication Levels
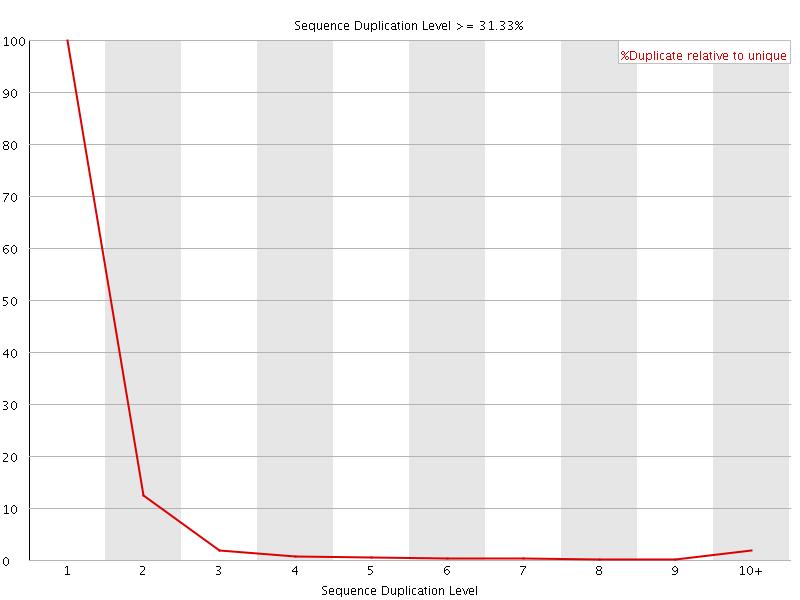
![[FAIL]](Icons/error.png) Overrepresented sequences
Overrepresented sequences
| Sequence | Count | Percentage | Possible Source |
|---|---|---|---|
| GATCGGAAGAGCACACGTCTGAACTCCAGTCACCCGTCCCGATCTCGTATG | 1611511 | 16.378757217047085 | TruSeq Adapter, Index 7 (97% over 36bp) |
| GATCGGAAGAGCACACGTCTGAACTCCAGTCACCCGTCCCGCTCTCGTATG | 29876 | 0.303647788079944 | TruSeq Adapter, Index 7 (97% over 36bp) |
| GATCGGAAGAGCACACGTCTGAACTCCAGTCACCCGTCCCGATCTCGTCTG | 13543 | 0.13764566856228017 | TruSeq Adapter, Index 7 (97% over 36bp) |
| GATCGGAAGAGCACACGTCTGAACTCCAGTCACCCGTCCCGGTCTCGTATG | 10653 | 0.10827285735759956 | TruSeq Adapter, Index 7 (97% over 36bp) |
![[FAIL]](Icons/error.png) Kmer Content
Kmer Content
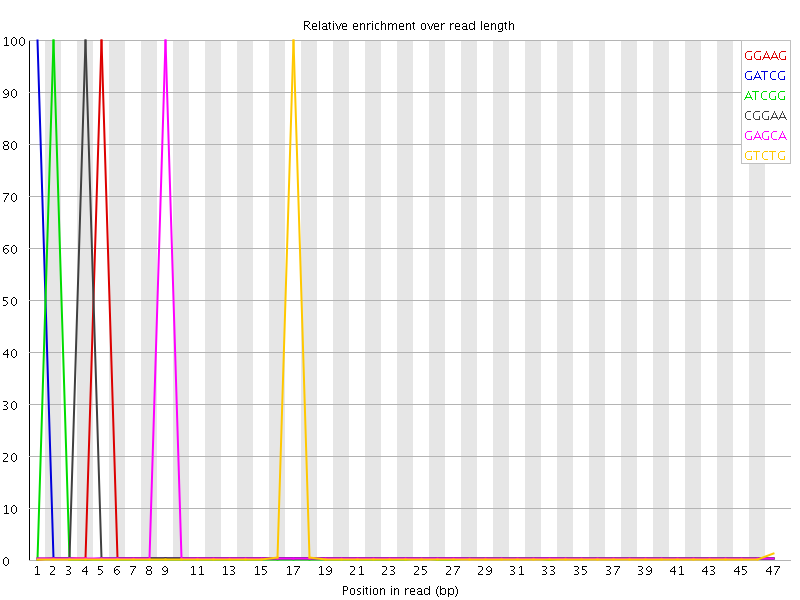
| Sequence | Count | Obs/Exp Overall | Obs/Exp Max | Max Obs/Exp Position |
|---|---|---|---|---|
| GGAAG | 2393805 | 6.6485233 | 260.89066 | 5 |
| GATCG | 2430760 | 6.303123 | 242.97726 | 1 |
| ATCGG | 2350565 | 6.095172 | 243.46045 | 2 |
| CGGAA | 2421720 | 6.0948005 | 236.54973 | 4 |
| GAGCA | 2403865 | 6.049864 | 235.4513 | 9 |
| GTCTG | 2261170 | 6.041226 | 247.89514 | 17 |
| TCGGA | 2323215 | 6.0242515 | 243.66014 | 3 |
| CCGTC | 2221030 | 5.989791 | 243.96378 | 34 |
| TCCCG | 2170870 | 5.8545175 | 240.33327 | 37 |
| GAAGA | 2643145 | 5.7957826 | 206.35019 | 6 |
| TCCAG | 2464320 | 5.790418 | 216.35825 | 25 |
| CGTCC | 2143740 | 5.7813516 | 242.81973 | 35 |
| CCCGT | 2134585 | 5.756661 | 244.37967 | 33 |
| AGAGC | 2279870 | 5.737803 | 235.2563 | 8 |
| TCTCG | 2324730 | 5.628122 | 212.65625 | 43 |
| ACCCG | 2143745 | 5.611154 | 238.47253 | 32 |
| GTCCC | 2076320 | 5.5995297 | 241.67328 | 36 |
| CGTCT | 2277155 | 5.5129437 | 224.44977 | 16 |
| CCCGA | 2073610 | 5.427579 | 223.63826 | 38 |
| CTGAA | 2611260 | 5.3458714 | 190.79582 | 19 |
| CGATC | 2267930 | 5.32896 | 199.65182 | 40 |
| ACGTC | 2246225 | 5.27796 | 218.22797 | 15 |
| GTCAC | 2241605 | 5.267104 | 215.63582 | 29 |
| AAGAG | 2399375 | 5.2612534 | 205.55893 | 7 |
| CCAGT | 2234555 | 5.250539 | 215.09274 | 26 |
| CTCGT | 2163805 | 5.2385263 | 211.33676 | 44 |
| CACGT | 2229400 | 5.2384257 | 218.26448 | 14 |
| AGCAC | 2292025 | 5.2270184 | 212.89005 | 10 |
| GCACA | 2280890 | 5.2016244 | 212.73262 | 11 |
| TCTGA | 2444760 | 5.1568294 | 196.27937 | 18 |
| CCGAT | 2185020 | 5.134146 | 199.61938 | 39 |
| CACCC | 2157460 | 5.117065 | 216.29433 | 31 |
| CAGTC | 2173265 | 5.1065254 | 214.9345 | 27 |
| ACACG | 2212515 | 5.0456934 | 212.13402 | 13 |
| CTCCA | 2368130 | 5.0421705 | 195.88911 | 24 |
| TCACC | 2362490 | 5.030161 | 194.82397 | 30 |
| TGAAC | 2408355 | 4.9304767 | 190.24455 | 20 |
| TTTTT | 3084325 | 4.894986 | 5.1862173 | 2 |
| GAACT | 2386835 | 4.8864193 | 189.99324 | 21 |
| CACAC | 2294715 | 4.7420106 | 192.39705 | 12 |
| GTATG | 2022625 | 4.708286 | 198.65768 | 47 |
| ACTCC | 2199975 | 4.6841373 | 196.53253 | 23 |
| GATCT | 2204055 | 4.649101 | 179.55598 | 41 |
| AGTCA | 2243085 | 4.592129 | 187.5918 | 28 |
| TCGTA | 2080645 | 4.3887877 | 181.2485 | 45 |
| AACTC | 2352415 | 4.363967 | 171.92998 | 22 |
| CGTAT | 2052020 | 4.3284073 | 180.9538 | 46 |
| ATCTC | 2244835 | 4.290719 | 162.88158 | 42 |
| AAAAA | 3130130 | 4.2782173 | 4.5434365 | 13 |
| ATTTT | 2252465 | 3.4695342 | 3.6419764 | 12 |
| AAAAT | 2266705 | 3.1920793 | 3.372016 | 15 |
| TTTTC | 1719485 | 3.0398767 | 3.4088728 | 2 |
| CTCTC | 480555 | 1.0542247 | 6.4040594 | 42 |
| CCCGC | 291980 | 0.87715554 | 8.242802 | 38 |
| CGCTC | 314255 | 0.8474995 | 7.191309 | 40 |
| CCGCT | 300590 | 0.8106469 | 7.0391903 | 39 |
| GCTCT | 334465 | 0.8097326 | 6.530021 | 41 |