![[OK]](Icons/tick.png) Basic Statistics
Basic Statistics
| Measure | Value |
|---|---|
| Filename | LW196_ATGTCA_L002_R1_001.fastq.gz |
| File type | Conventional base calls |
| Encoding | Sanger / Illumina 1.9 |
| Total Sequences | 13045884 |
| Filtered Sequences | 0 |
| Sequence length | 51 |
| %GC | 42 |
![[OK]](Icons/tick.png) Per base sequence quality
Per base sequence quality
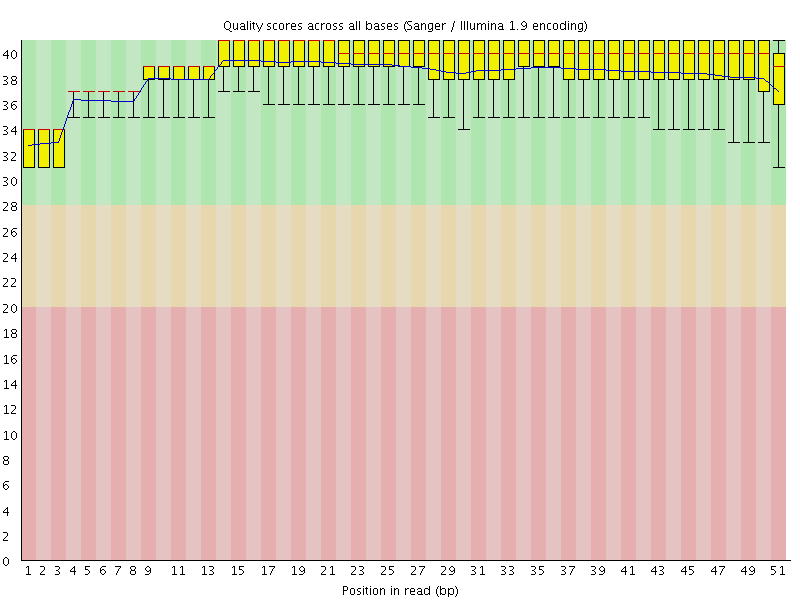
![[OK]](Icons/tick.png) Per sequence quality scores
Per sequence quality scores
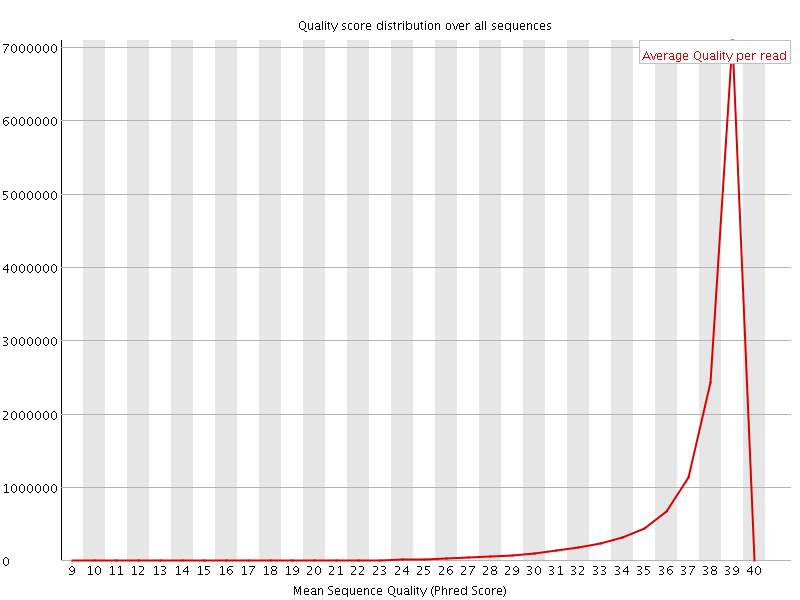
![[WARN]](Icons/warning.png) Per base sequence content
Per base sequence content
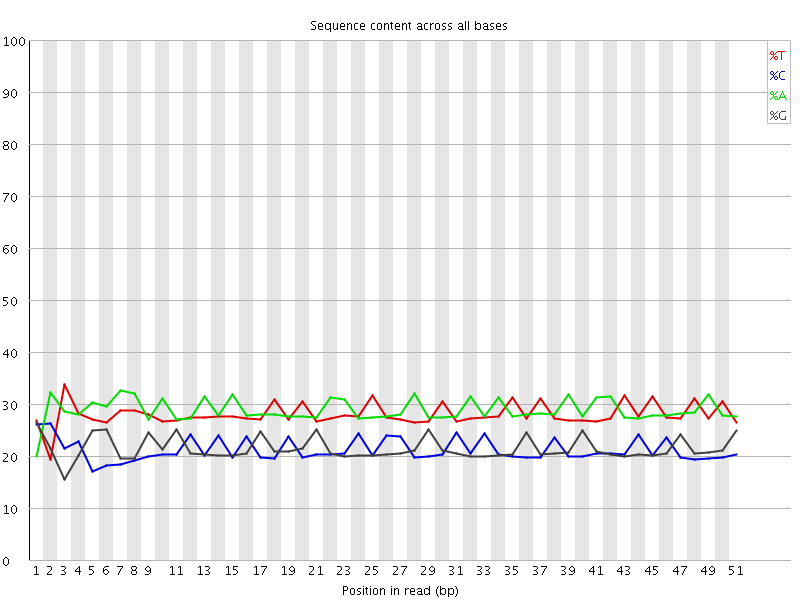
![[FAIL]](Icons/error.png) Per base GC content
Per base GC content
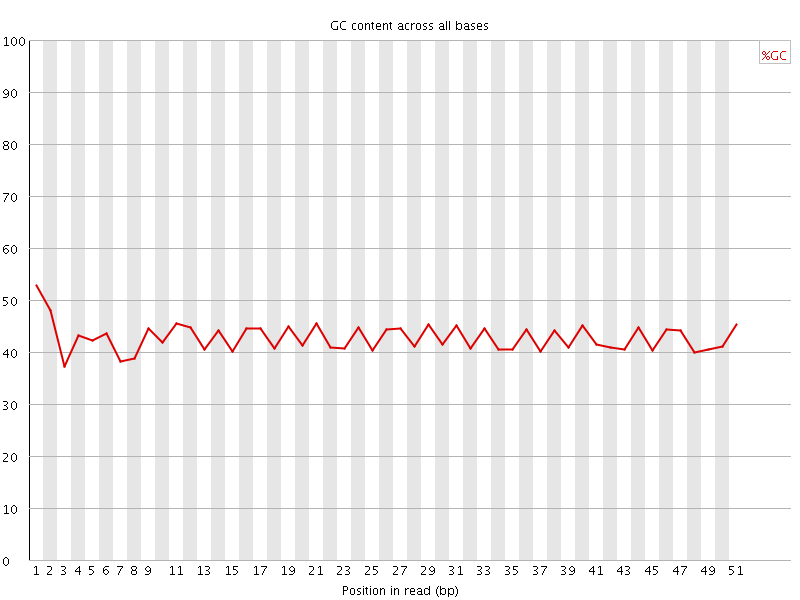
![[FAIL]](Icons/error.png) Per sequence GC content
Per sequence GC content
![[OK]](Icons/tick.png) Per base N content
Per base N content
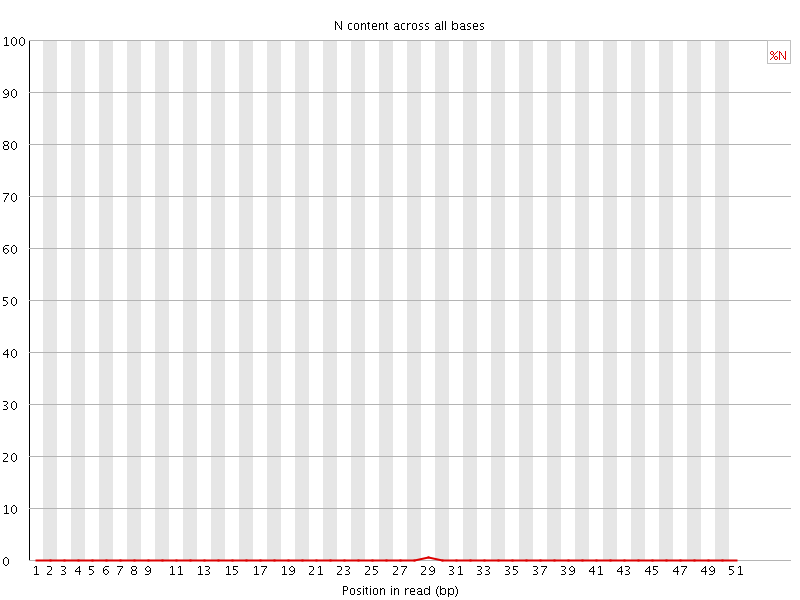
![[OK]](Icons/tick.png) Sequence Length Distribution
Sequence Length Distribution
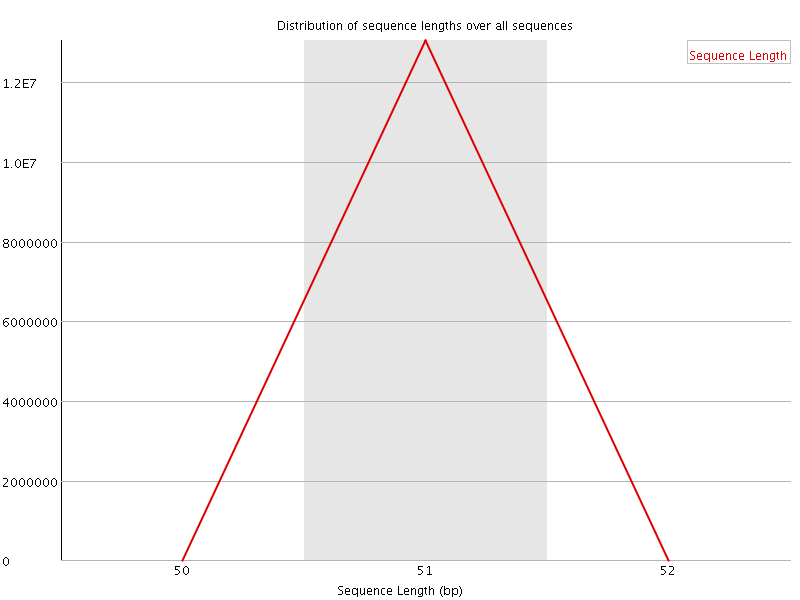
![[OK]](Icons/tick.png) Sequence Duplication Levels
Sequence Duplication Levels
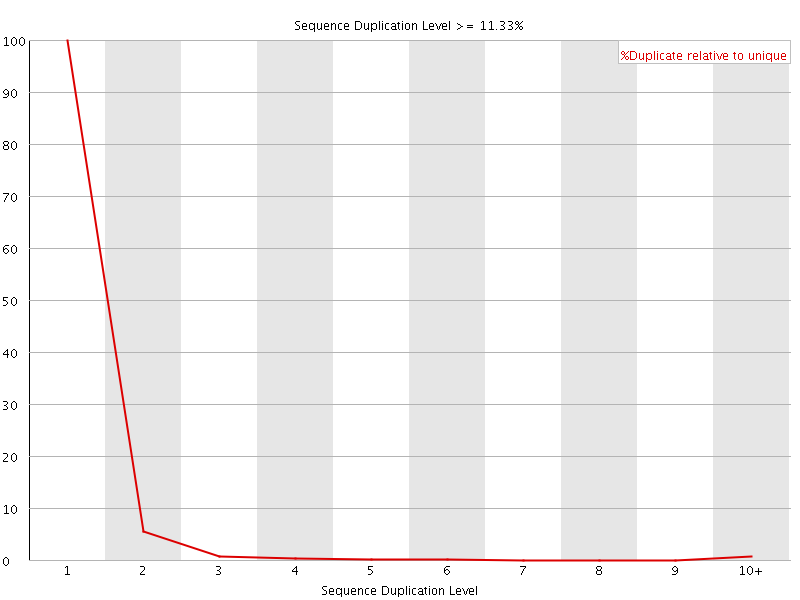
![[FAIL]](Icons/error.png) Overrepresented sequences
Overrepresented sequences
| Sequence | Count | Percentage | Possible Source |
|---|---|---|---|
| GATCGGAAGAGCACACGTCTGAACTCCAGTCACATGTCAGAATCTCGTATG | 449918 | 3.448735248604081 | TruSeq Adapter, Index 1 (97% over 35bp) |
![[FAIL]](Icons/error.png) Kmer Content
Kmer Content
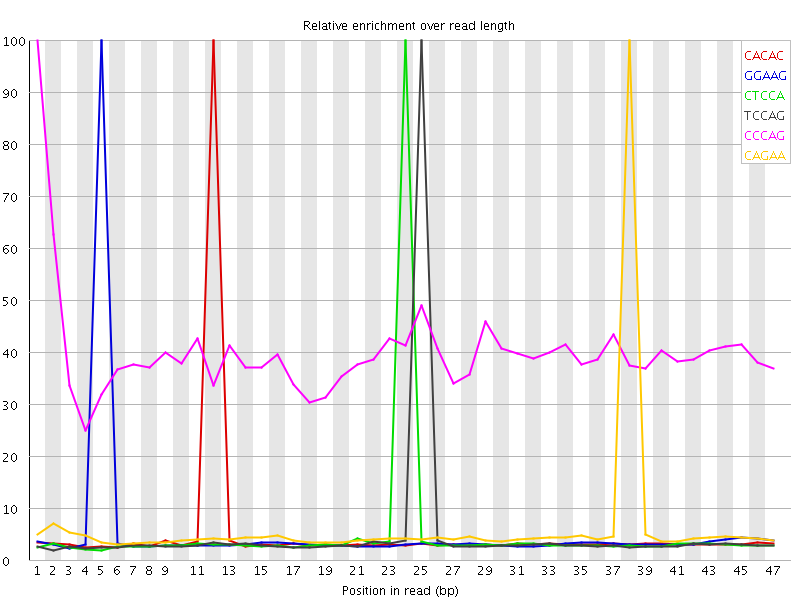
| Sequence | Count | Obs/Exp Overall | Obs/Exp Max | Max Obs/Exp Position |
|---|---|---|---|---|
| CACAC | 1309485 | 2.6400266 | 50.245804 | 12 |
| GGAAG | 1359570 | 2.6285522 | 49.61455 | 5 |
| CTCCA | 1253335 | 2.598445 | 50.986355 | 24 |
| TCCAG | 1238905 | 2.532912 | 50.439507 | 25 |
| CCCAG | 855780 | 2.312535 | 5.759784 | 1 |
| CAGAA | 1554300 | 2.273504 | 35.923508 | 38 |
| GAGCA | 1130280 | 2.2159772 | 49.113754 | 9 |
| GCACA | 1114470 | 2.215705 | 49.57941 | 11 |
| CCAGT | 1067290 | 2.1820495 | 49.926247 | 26 |
| GAAGA | 1511395 | 2.1800907 | 37.096004 | 6 |
| AGAGC | 1109345 | 2.1749332 | 49.143208 | 8 |
| GTCTG | 1046030 | 2.1687057 | 51.38361 | 17 |
| ACTCC | 1022630 | 2.1201417 | 50.57619 | 23 |
| AGCAC | 1058045 | 2.103525 | 49.47067 | 10 |
| CATGT | 1358920 | 2.101996 | 38.20812 | 33 |
| GTCAG | 1029775 | 2.076157 | 48.764736 | 36 |
| AAGAG | 1433540 | 2.06779 | 36.912148 | 7 |
| ACATG | 1366995 | 2.0562048 | 37.289566 | 32 |
| CTGAA | 1365305 | 2.0536628 | 37.678005 | 19 |
| TCACA | 1343465 | 2.049227 | 37.797047 | 30 |
| CAGTC | 981155 | 2.0059485 | 49.891933 | 27 |
| TCTGA | 1285160 | 1.9879031 | 38.702023 | 18 |
| CACAT | 1301875 | 1.9857885 | 37.618828 | 31 |
| GTCAC | 955930 | 1.9543762 | 49.974567 | 29 |
| TCAGA | 1291170 | 1.9421505 | 36.745304 | 37 |
| TGTCA | 1218525 | 1.8848312 | 37.732063 | 35 |
| ATCTC | 1198230 | 1.8795003 | 38.18554 | 42 |
| TGAAC | 1177100 | 1.7705686 | 37.38006 | 20 |
| GATCG | 873325 | 1.760734 | 50.234062 | 1 |
| GAATC | 1145815 | 1.7235105 | 36.36834 | 40 |
| ATGTC | 1090855 | 1.6873494 | 37.599483 | 34 |
| AACTC | 1106025 | 1.6870526 | 37.519363 | 22 |
| AGAAT | 1513850 | 1.6753271 | 27.359688 | 39 |
| TCTCG | 793965 | 1.6692526 | 50.467606 | 43 |
| AGTCA | 1097455 | 1.6507683 | 37.016956 | 28 |
| GAACT | 1082465 | 1.6282208 | 37.21267 | 21 |
| CGGAA | 814755 | 1.5973731 | 48.91448 | 4 |
| CGTCT | 750330 | 1.5775133 | 51.313244 | 16 |
| ATCGG | 781185 | 1.5749679 | 50.156097 | 2 |
| CACGT | 759765 | 1.5533218 | 50.055565 | 14 |
| ACACG | 754980 | 1.5009941 | 48.77621 | 13 |
| ACGTC | 730350 | 1.4931835 | 49.95744 | 15 |
| CTCGT | 710075 | 1.4928801 | 50.11063 | 44 |
| TCGGA | 737860 | 1.4876194 | 50.115402 | 3 |
| AATCT | 1271720 | 1.4676112 | 28.084898 | 41 |
| GTATG | 948805 | 1.447274 | 36.616795 | 47 |
| CGTAT | 700660 | 1.0837904 | 36.834682 | 46 |
| TCGTA | 688520 | 1.0650122 | 36.88063 | 45 |