![[OK]](Icons/tick.png) Basic Statistics
Basic Statistics
| Measure | Value |
|---|---|
| Filename | LW194_AGTCAA_L002_R1_001.fastq.gz |
| File type | Conventional base calls |
| Encoding | Sanger / Illumina 1.9 |
| Total Sequences | 6608247 |
| Filtered Sequences | 0 |
| Sequence length | 51 |
| %GC | 45 |
![[OK]](Icons/tick.png) Per base sequence quality
Per base sequence quality
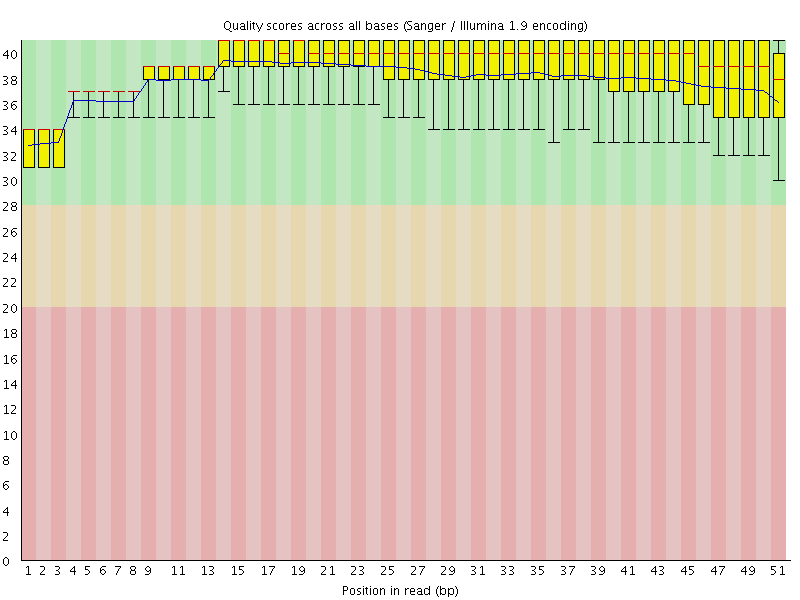
![[OK]](Icons/tick.png) Per sequence quality scores
Per sequence quality scores
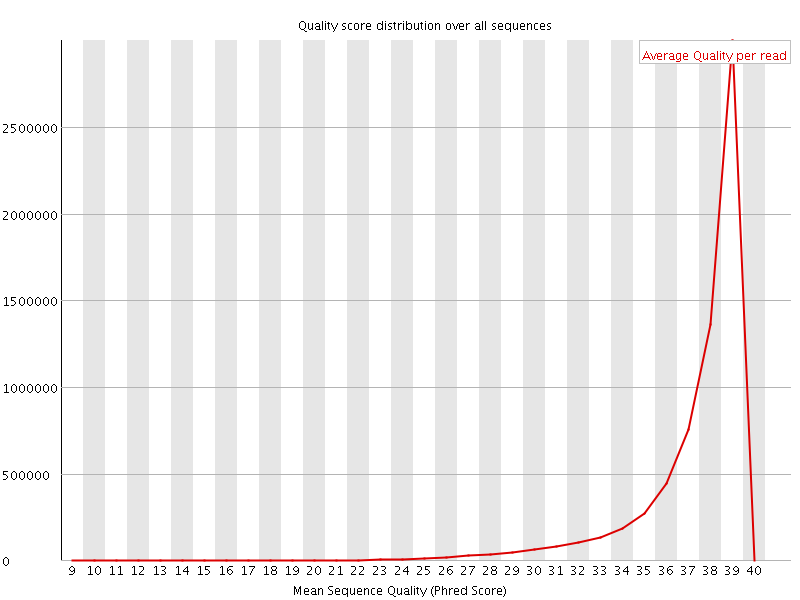
![[WARN]](Icons/warning.png) Per base sequence content
Per base sequence content
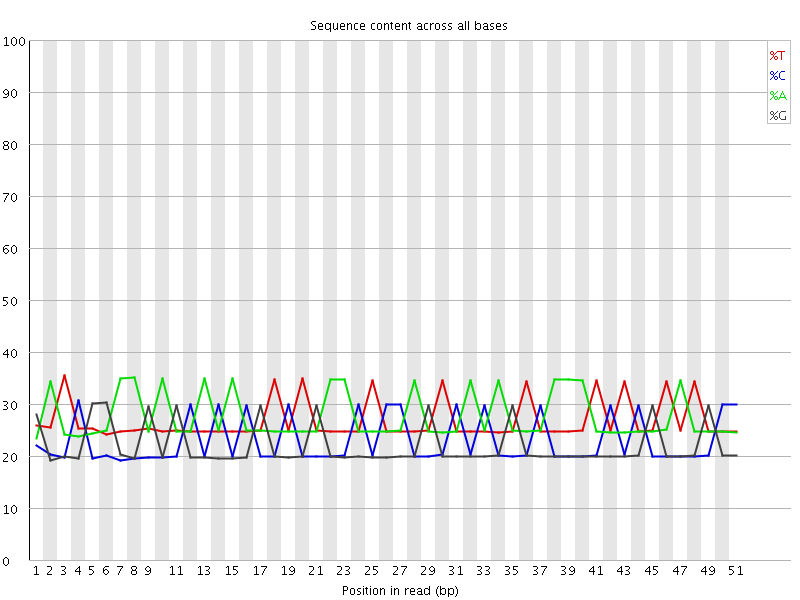
![[WARN]](Icons/warning.png) Per base GC content
Per base GC content
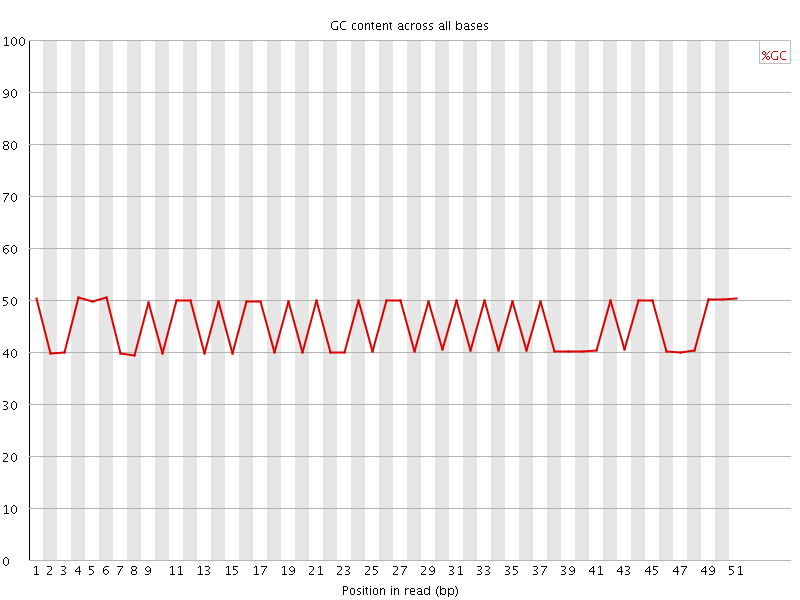
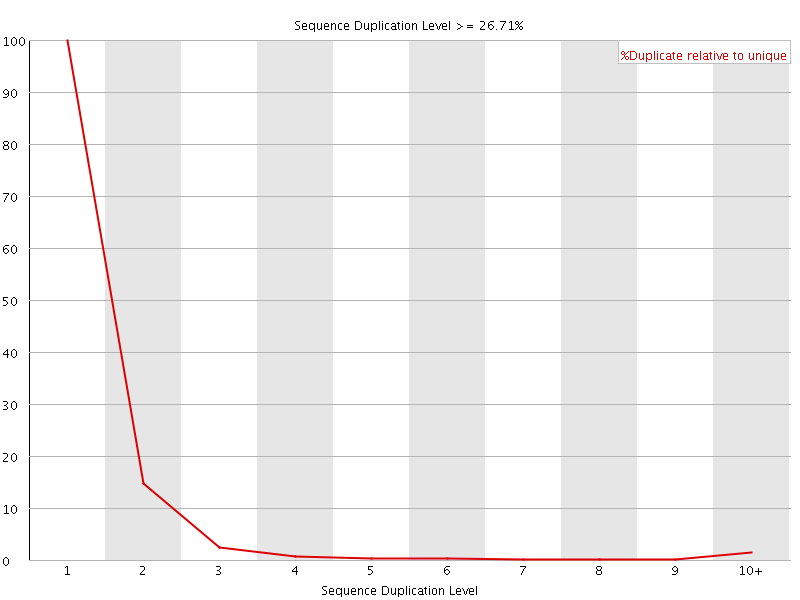
![[FAIL]](Icons/error.png) Per sequence GC content
Per sequence GC content
![[OK]](Icons/tick.png) Per base N content
Per base N content
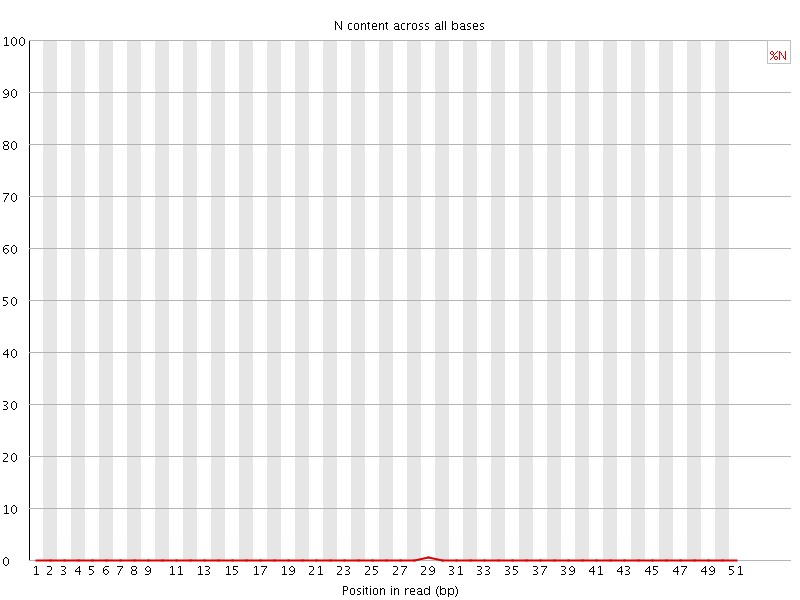
![[OK]](Icons/tick.png) Sequence Length Distribution
Sequence Length Distribution
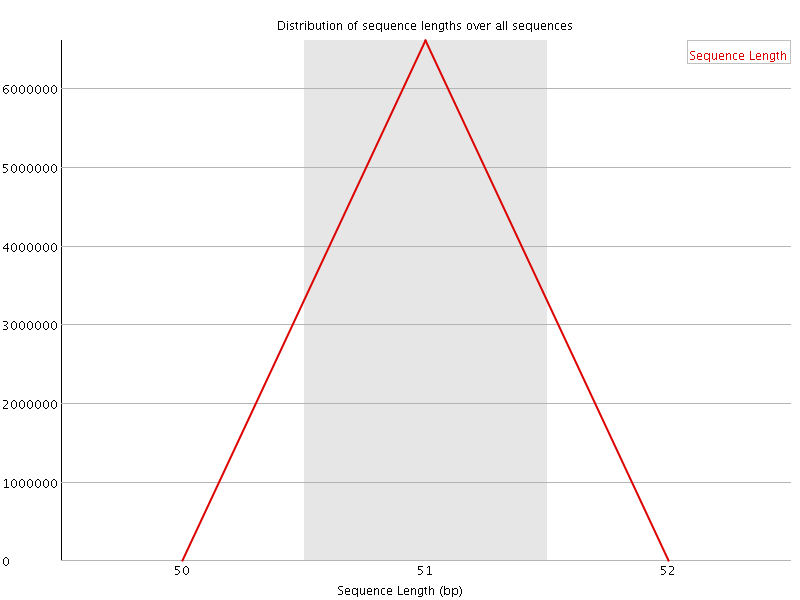
![[WARN]](Icons/warning.png) Sequence Duplication Levels
Sequence Duplication Levels
![[FAIL]](Icons/error.png) Overrepresented sequences
Overrepresented sequences
| Sequence | Count | Percentage | Possible Source |
|---|---|---|---|
| GATCGGAAGAGCACACGTCTGAACTCCAGTCACAGTCAAATCTCGTATGCC | 601977 | 9.109480925879435 | TruSeq Adapter, Index 8 (97% over 36bp) |
![[FAIL]](Icons/error.png) Kmer Content
Kmer Content
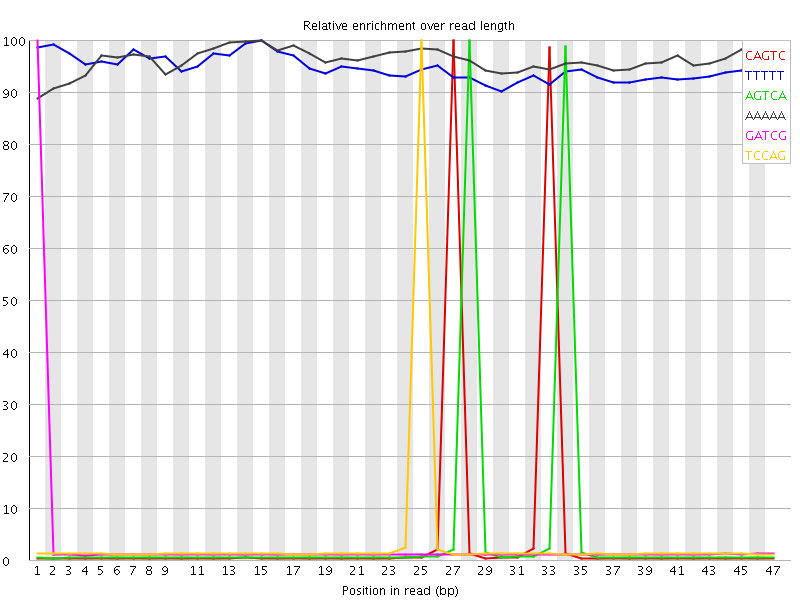
| Sequence | Count | Obs/Exp Overall | Obs/Exp Max | Max Obs/Exp Position |
|---|---|---|---|---|
| CAGTC | 1482680 | 5.3945565 | 111.49053 | 27 |
| TTTTT | 2222030 | 5.045308 | 5.326626 | 15 |
| AGTCA | 1519075 | 4.592379 | 92.53275 | 28 |
| AAAAA | 2200040 | 4.259908 | 4.4269285 | 15 |
| GATCG | 1066465 | 4.0514507 | 120.6892 | 1 |
| TCCAG | 1085330 | 3.9488451 | 112.765076 | 25 |
| GGAAG | 973255 | 3.739492 | 121.77352 | 5 |
| TCTCG | 993755 | 3.7326841 | 114.89874 | 41 |
| ATCGG | 968885 | 3.680749 | 120.636665 | 2 |
| CGGAA | 995195 | 3.6621697 | 116.9945 | 4 |
| ATTTT | 1663860 | 3.659493 | 3.884238 | 6 |
| GAGCA | 974600 | 3.5863836 | 116.03908 | 9 |
| TCGGA | 903815 | 3.433551 | 120.44403 | 3 |
| GAAGA | 1114630 | 3.408091 | 97.23402 | 6 |
| CTGAA | 1126940 | 3.4068992 | 94.63455 | 19 |
| GTCTG | 868590 | 3.406531 | 121.85279 | 17 |
| ATGCC | 917250 | 3.3373063 | 110.68144 | 47 |
| TTTTC | 1249265 | 3.3068044 | 3.6111765 | 1 |
| AAAAT | 1644555 | 3.287393 | 3.4963174 | 37 |
| CTCCA | 933785 | 3.2538655 | 107.58068 | 24 |
| AGAGC | 877290 | 3.228297 | 115.720314 | 8 |
| CGTCT | 857990 | 3.2227318 | 116.67905 | 16 |
| CTCGT | 857650 | 3.2214549 | 114.07886 | 42 |
| TCAAA | 1336535 | 3.2153885 | 73.585625 | 36 |
| CCAGT | 881140 | 3.205924 | 111.80622 | 26 |
| AGCAC | 905110 | 3.1898925 | 110.861404 | 10 |
| CGGCG | 567895 | 3.1604838 | 3.2894704 | 42 |
| ACGTC | 863855 | 3.1430347 | 113.26608 | 15 |
| GTCAC | 863465 | 3.1416159 | 111.63157 | 29 |
| TCTGA | 999670 | 3.1199584 | 97.4051 | 18 |
| GCACA | 880415 | 3.1028595 | 110.61982 | 11 |
| AAATC | 1271295 | 3.0584362 | 73.81023 | 38 |
| CACGT | 837350 | 3.0465994 | 113.30588 | 14 |
| CACAG | 862950 | 3.0413074 | 107.65087 | 31 |
| AATTT | 1416545 | 3.017873 | 3.1823146 | 5 |
| TGAAC | 980310 | 2.9636161 | 94.04492 | 20 |
| GAACT | 972955 | 2.9413805 | 93.967476 | 21 |
| CAAAT | 1219885 | 2.934756 | 73.60553 | 37 |
| ACACG | 826075 | 2.9113486 | 110.00678 | 13 |
| CACAC | 861235 | 2.9069715 | 105.60986 | 12 |
| ATCTC | 959405 | 2.8677316 | 91.20779 | 40 |
| AAGAG | 934995 | 2.8588398 | 96.51768 | 7 |
| ACTCC | 819485 | 2.8555763 | 107.62209 | 23 |
| GTCAA | 921445 | 2.7856588 | 91.602905 | 35 |
| TCACA | 927270 | 2.6847825 | 88.579 | 30 |
| AACTC | 920070 | 2.663936 | 89.860695 | 22 |
| ACAGT | 878460 | 2.655709 | 91.892525 | 32 |
| GTATG | 804195 | 2.620651 | 98.7228 | 45 |
| TATGC | 826710 | 2.5801523 | 94.51978 | 46 |
| TCGTA | 821050 | 2.5624874 | 94.632324 | 43 |
| CGTAT | 806835 | 2.5181227 | 94.53518 | 44 |
| AATCT | 997120 | 2.4764762 | 75.543884 | 39 |
| TGCCG | 458600 | 2.0967553 | 5.4471335 | 47 |