![[OK]](Icons/tick.png) Basic Statistics
Basic Statistics
| Measure | Value |
|---|---|
| Filename | LW193_CTTGTA_L002_R1_001.fastq.gz |
| File type | Conventional base calls |
| Encoding | Sanger / Illumina 1.9 |
| Total Sequences | 6284938 |
| Filtered Sequences | 0 |
| Sequence length | 51 |
| %GC | 38 |
![[OK]](Icons/tick.png) Per base sequence quality
Per base sequence quality
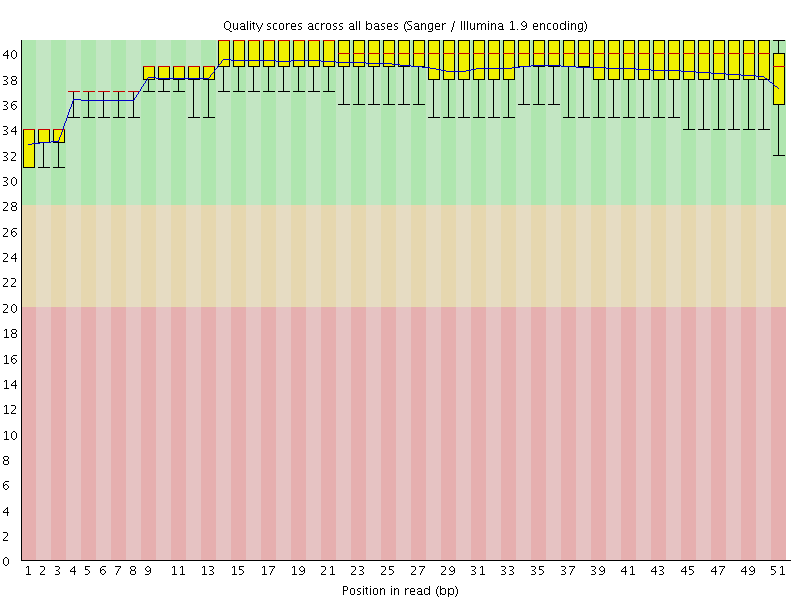
![[OK]](Icons/tick.png) Per sequence quality scores
Per sequence quality scores
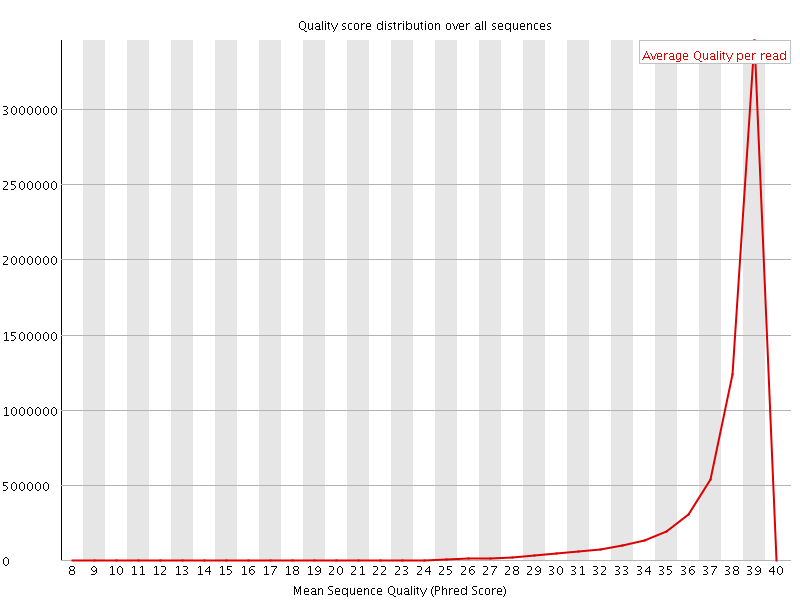
![[WARN]](Icons/warning.png) Per base sequence content
Per base sequence content
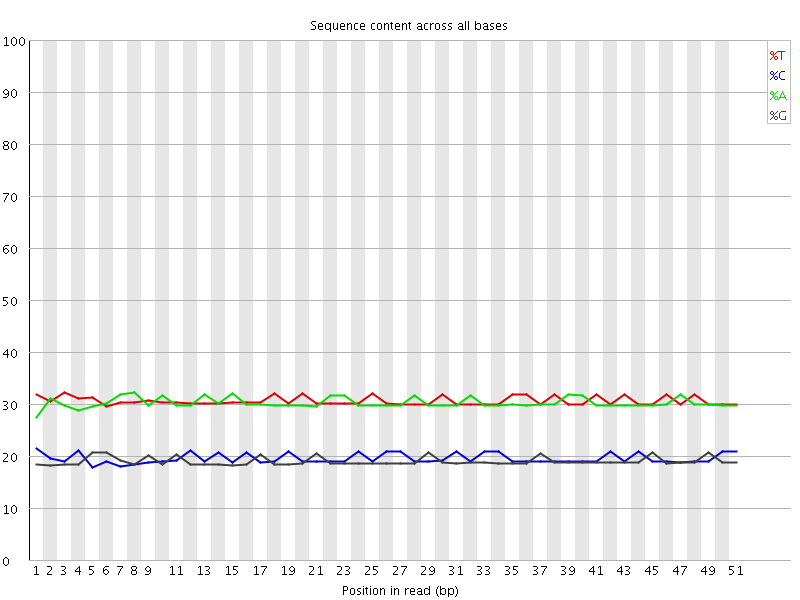
![[OK]](Icons/tick.png) Per base GC content
Per base GC content
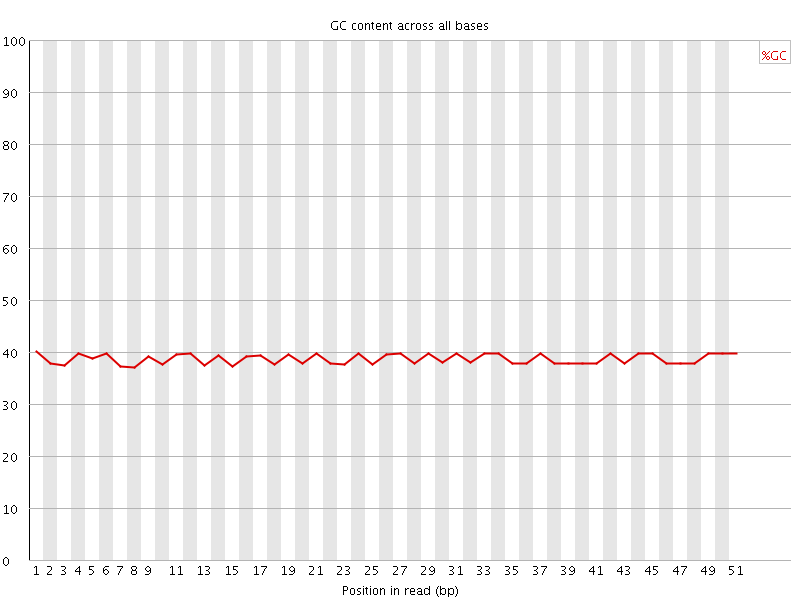
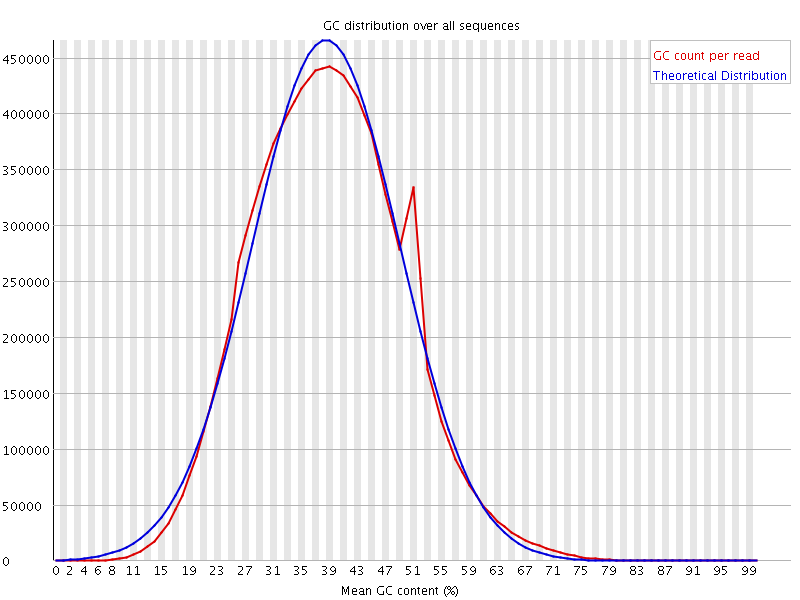
![[OK]](Icons/tick.png) Per sequence GC content
Per sequence GC content
![[OK]](Icons/tick.png) Per base N content
Per base N content
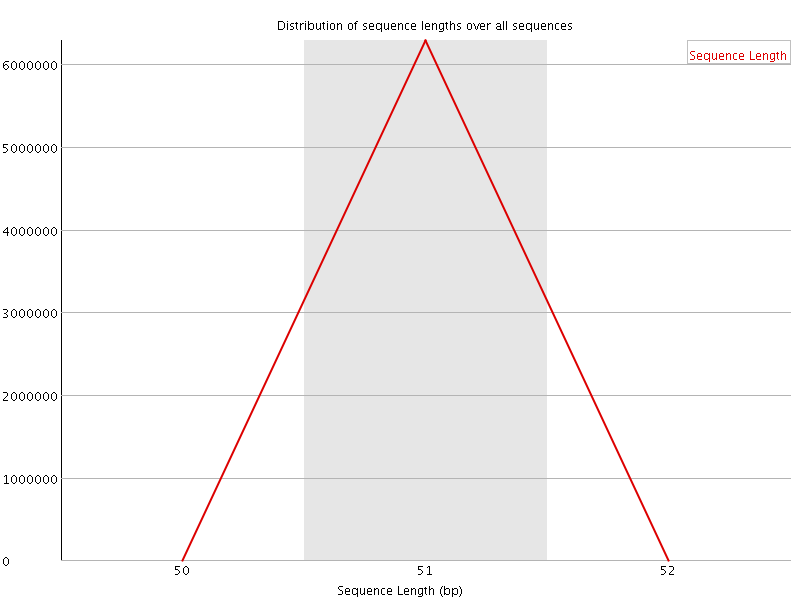
![[OK]](Icons/tick.png) Sequence Length Distribution
Sequence Length Distribution
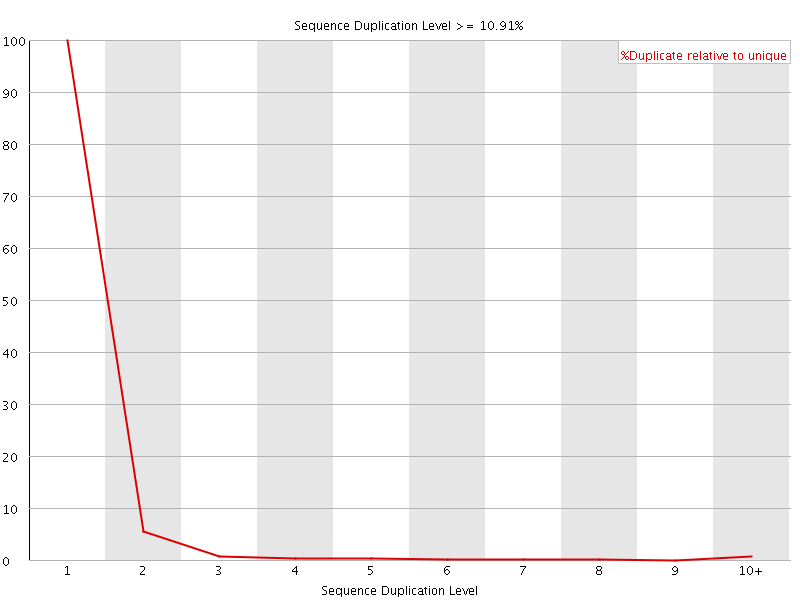
![[OK]](Icons/tick.png) Sequence Duplication Levels
Sequence Duplication Levels
![[FAIL]](Icons/error.png) Overrepresented sequences
Overrepresented sequences
| Sequence | Count | Percentage | Possible Source |
|---|---|---|---|
| GATCGGAAGAGCACACGTCTGAACTCCAGTCACCTTGTAATCTCGTATGCC | 111405 | 1.7725711852686534 | TruSeq Adapter, Index 12 (100% over 51bp) |
![[WARN]](Icons/warning.png) Kmer Content
Kmer Content
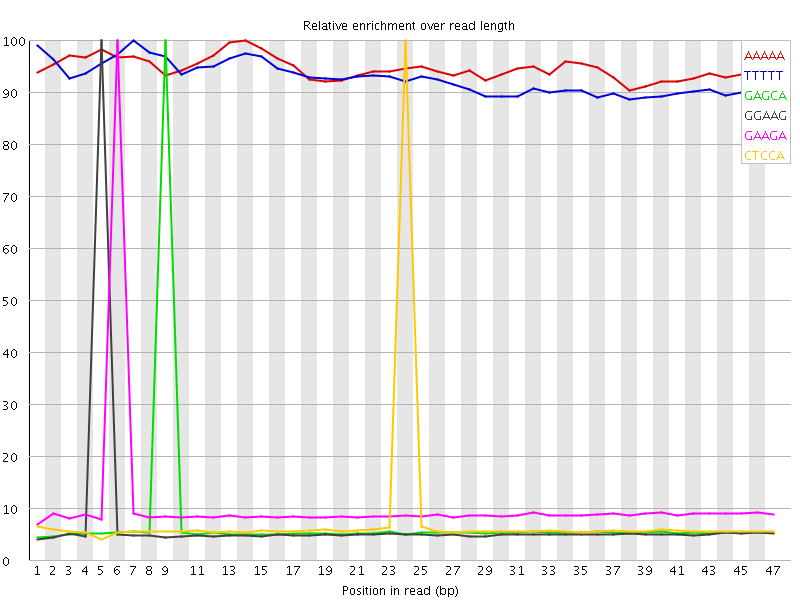
| Sequence | Count | Obs/Exp Overall | Obs/Exp Max | Max Obs/Exp Position |
|---|---|---|---|---|
| AAAAA | 2681655 | 3.500712 | 3.6969447 | 14 |
| TTTTT | 2788620 | 3.4362564 | 3.705472 | 7 |
| GAGCA | 448445 | 2.2789516 | 30.901115 | 9 |
| GGAAG | 428600 | 2.2449975 | 31.992813 | 5 |
| GAAGA | 673760 | 2.220916 | 20.875399 | 6 |
| CTCCA | 467010 | 2.208337 | 28.521734 | 24 |
| CGGAA | 429990 | 2.1851654 | 31.074097 | 4 |
| TCCAG | 431530 | 2.1032352 | 29.167562 | 25 |
| CTGAA | 663665 | 2.0981023 | 19.742638 | 19 |
| TCTCG | 388135 | 1.8700285 | 28.391907 | 41 |
| TCGGA | 370410 | 1.8607893 | 30.393286 | 3 |
| AGAGC | 351570 | 1.7866429 | 30.4299 | 8 |
| TCACC | 373985 | 1.7684526 | 27.666286 | 30 |
| GCACA | 355555 | 1.7530535 | 29.563677 | 11 |
| CACAC | 363155 | 1.7371715 | 28.690605 | 12 |
| AGCAC | 346270 | 1.7072737 | 29.4977 | 10 |
| TCTGA | 532175 | 1.6631094 | 19.136799 | 18 |
| ATCGG | 330955 | 1.6625836 | 30.045404 | 2 |
| CCAGT | 325615 | 1.5870159 | 28.62371 | 26 |
| AAGAG | 473235 | 1.5599251 | 20.123272 | 7 |
| CTCGT | 322180 | 1.5522583 | 27.997477 | 42 |
| CGTCT | 316210 | 1.5234951 | 28.508528 | 16 |
| CACGT | 311655 | 1.5189762 | 28.77828 | 14 |
| GATCG | 300465 | 1.5094142 | 29.727627 | 1 |
| GTCTG | 303905 | 1.5091796 | 29.298162 | 17 |
| GTCAC | 307170 | 1.4971168 | 28.407068 | 29 |
| TGAAC | 473430 | 1.496696 | 19.234814 | 20 |
| CACCT | 312375 | 1.4771191 | 27.319038 | 31 |
| CCTTG | 306410 | 1.4762787 | 27.837545 | 33 |
| ACACG | 297615 | 1.467382 | 29.1771 | 13 |
| ACTCC | 306995 | 1.4516789 | 27.931623 | 23 |
| ATGCC | 295880 | 1.4420905 | 28.251486 | 47 |
| ACGTC | 290340 | 1.4150891 | 28.74215 | 15 |
| CAGTC | 283605 | 1.3822634 | 28.382874 | 27 |
| AACTC | 441180 | 1.3531812 | 18.560974 | 22 |
| GAACT | 419040 | 1.3247479 | 19.012886 | 21 |
| CTTGT | 421070 | 1.300796 | 18.150158 | 34 |
| ATCTC | 427275 | 1.2954963 | 17.980946 | 40 |
| AGTCA | 367820 | 1.1628218 | 18.630625 | 28 |
| AATCT | 556525 | 1.0944964 | 11.864416 | 39 |
| ACCTT | 346830 | 1.0515875 | 17.5958 | 32 |
| TTGTA | 512615 | 1.0271815 | 11.931511 | 35 |
| TCGTA | 328240 | 1.0257885 | 18.224747 | 43 |
| TATGC | 301800 | 0.9431605 | 18.111706 | 46 |
| GTATG | 287440 | 0.9258725 | 18.620962 | 45 |
| CGTAT | 294115 | 0.91914403 | 18.109377 | 44 |
| TGTAA | 449335 | 0.91083044 | 11.950182 | 36 |
| GTAAT | 387200 | 0.78487885 | 11.830854 | 37 |
| TAATC | 381690 | 0.75065506 | 11.432849 | 38 |