![[OK]](Icons/tick.png) Basic Statistics
Basic Statistics
| Measure | Value |
|---|---|
| Filename | LW192_GGCTAC_L002_R1_001.fastq.gz |
| File type | Conventional base calls |
| Encoding | Sanger / Illumina 1.9 |
| Total Sequences | 6259067 |
| Filtered Sequences | 0 |
| Sequence length | 51 |
| %GC | 37 |
![[OK]](Icons/tick.png) Per base sequence quality
Per base sequence quality
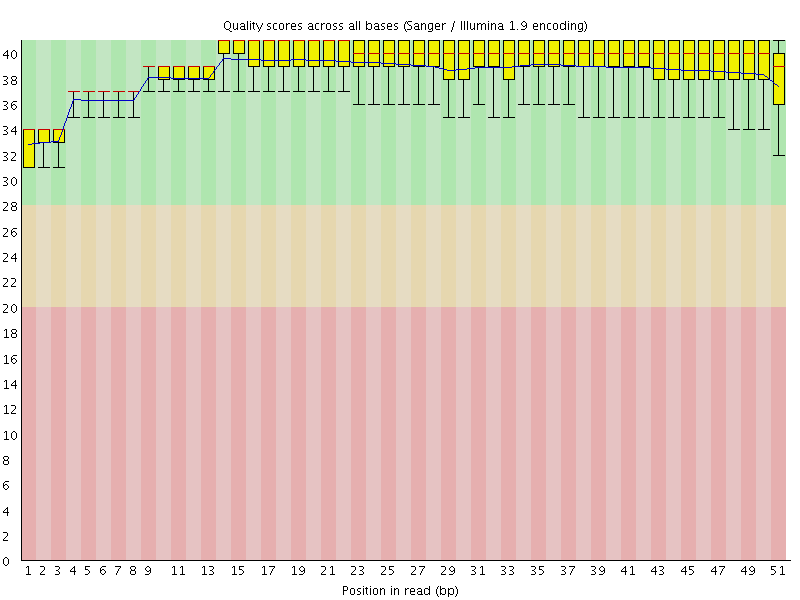
![[OK]](Icons/tick.png) Per sequence quality scores
Per sequence quality scores
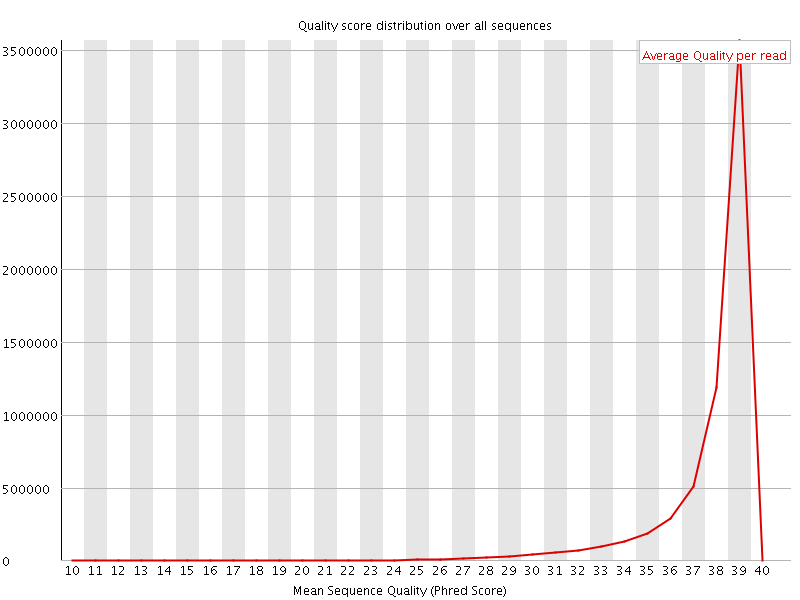
![[WARN]](Icons/warning.png) Per base sequence content
Per base sequence content
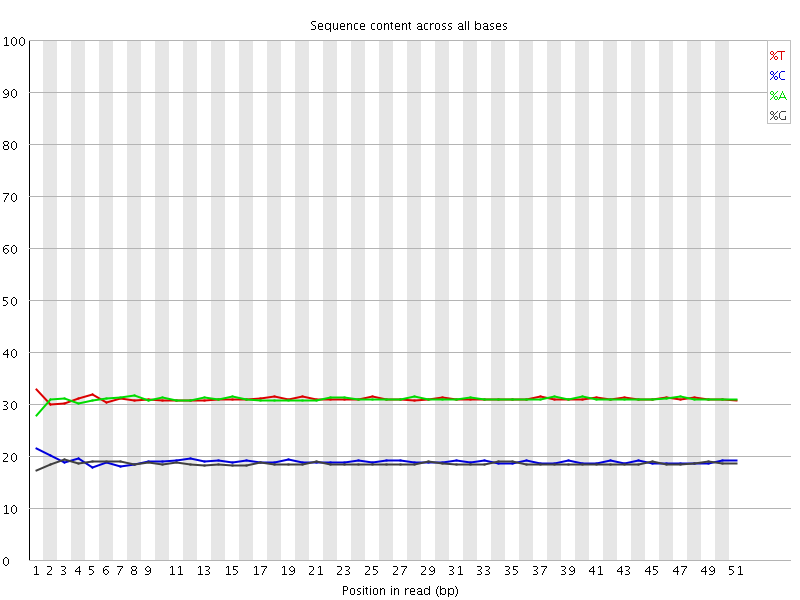
![[OK]](Icons/tick.png) Per base GC content
Per base GC content
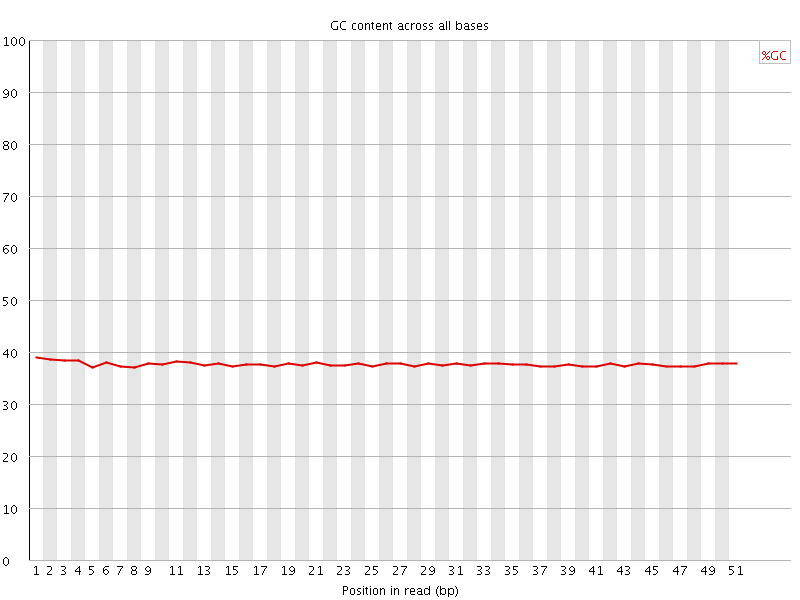
![[OK]](Icons/tick.png) Per sequence GC content
Per sequence GC content
![[OK]](Icons/tick.png) Per base N content
Per base N content
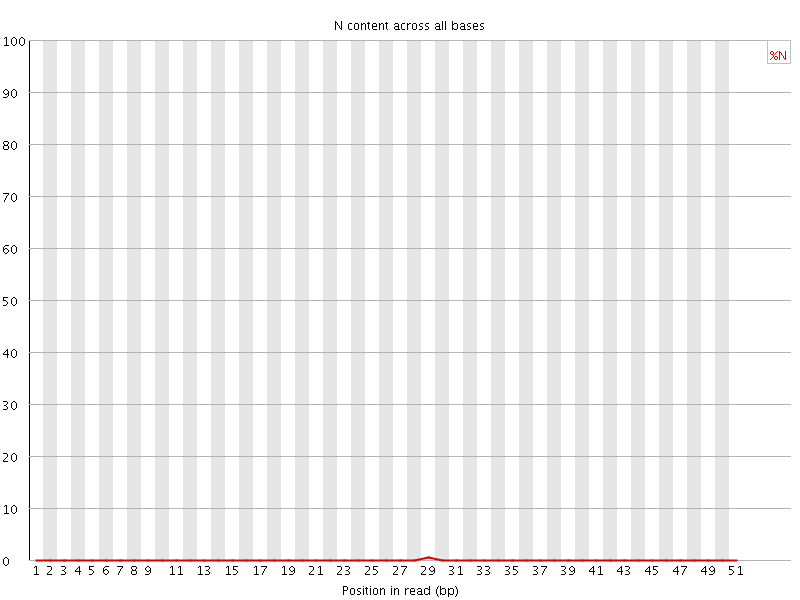
![[OK]](Icons/tick.png) Sequence Length Distribution
Sequence Length Distribution
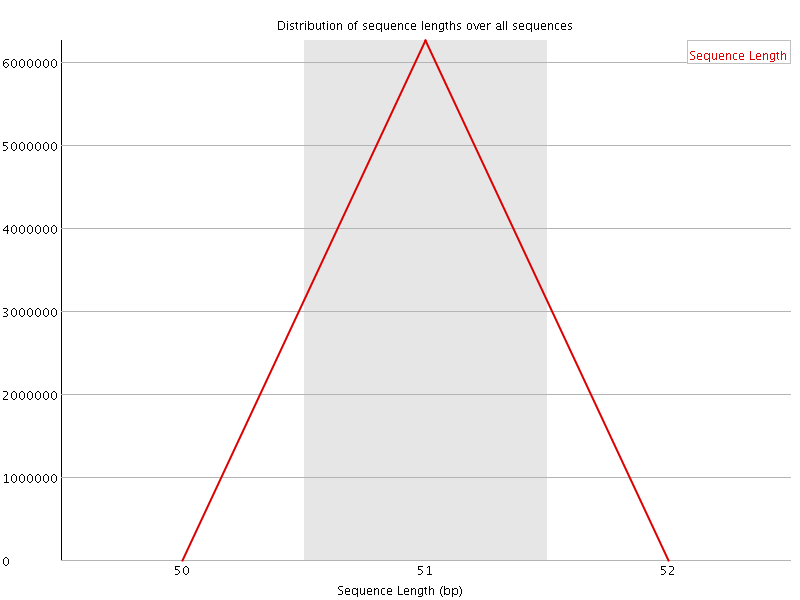
![[OK]](Icons/tick.png) Sequence Duplication Levels
Sequence Duplication Levels
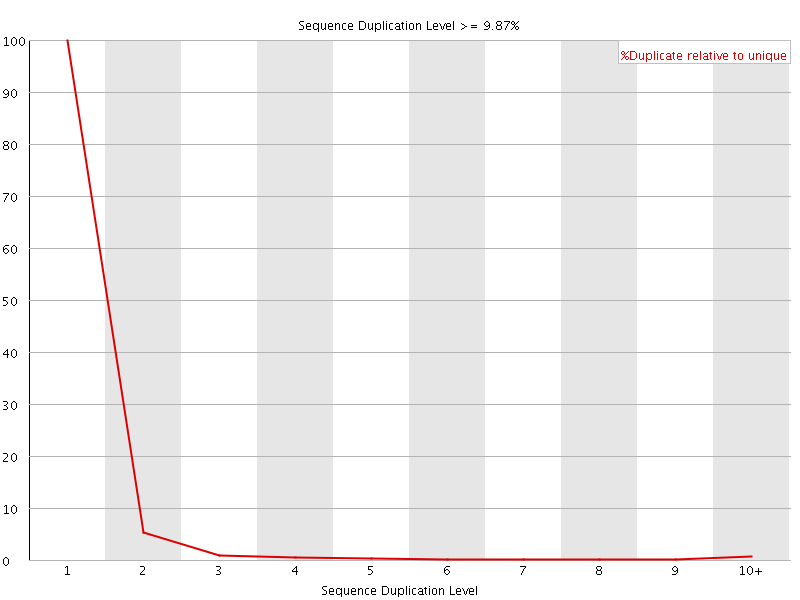
![[WARN]](Icons/warning.png) Overrepresented sequences
Overrepresented sequences
| Sequence | Count | Percentage | Possible Source |
|---|---|---|---|
| GATCGGAAGAGCACACGTCTGAACTCCAGTCACGGCTACATCTCGTATGCC | 26567 | 0.42445623285387424 | TruSeq Adapter, Index 11 (100% over 51bp) |
![[WARN]](Icons/warning.png) Kmer Content
Kmer Content
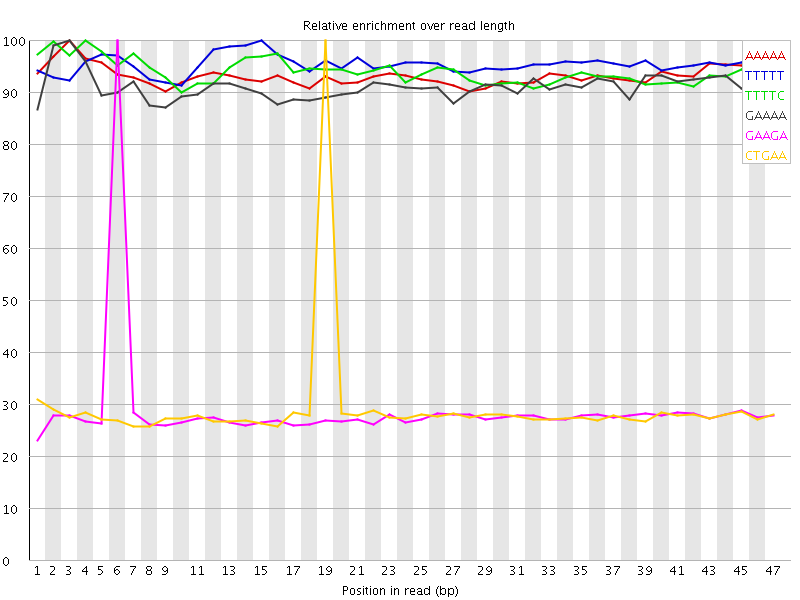
| Sequence | Count | Obs/Exp Overall | Obs/Exp Max | Max Obs/Exp Position |
|---|---|---|---|---|
| AAAAA | 3102545 | 3.638159 | 3.903042 | 3 |
| TTTTT | 3095065 | 3.6122901 | 3.785008 | 15 |
| TTTTC | 1613325 | 3.0661404 | 3.2621236 | 4 |
| GAAAA | 1570575 | 3.0622163 | 3.3603585 | 3 |
| GAAGA | 583510 | 1.8916428 | 6.547647 | 6 |
| CTGAA | 585210 | 1.8544964 | 6.3411713 | 19 |
| CGGAA | 346555 | 1.8277217 | 9.535324 | 4 |
| CTCCA | 360520 | 1.8185464 | 8.92329 | 24 |
| GAGCA | 339730 | 1.7917273 | 9.331051 | 9 |
| GGAAG | 330265 | 1.7801924 | 9.645492 | 5 |
| TCCAG | 335900 | 1.7316948 | 8.931194 | 25 |
| TCTCG | 287070 | 1.4785604 | 8.453765 | 41 |
| TCGGA | 275280 | 1.450449 | 9.030341 | 3 |
| TCTGA | 449195 | 1.422129 | 5.900352 | 18 |
| CGGCT | 158735 | 1.3606539 | 12.789788 | 33 |
| CACAC | 269085 | 1.3586094 | 8.601146 | 12 |
| GCACA | 256590 | 1.3240709 | 8.691355 | 11 |
| ACGGC | 151920 | 1.303467 | 12.735855 | 32 |
| AGAGC | 246080 | 1.2978196 | 8.863075 | 8 |
| AAGAG | 380465 | 1.2334044 | 5.884054 | 7 |
| TGAAC | 386810 | 1.2257783 | 5.69603 | 20 |
| ATCGG | 232310 | 1.2240403 | 8.734732 | 2 |
| CCAGT | 237200 | 1.2228582 | 8.368121 | 26 |
| AGCAC | 236735 | 1.221614 | 8.622309 | 10 |
| CACGG | 141720 | 1.2159516 | 12.806116 | 31 |
| CTCGT | 219600 | 1.1310548 | 8.103606 | 42 |
| CGTCT | 217470 | 1.120084 | 8.433319 | 16 |
| GTCAC | 215930 | 1.113203 | 8.110952 | 29 |
| CACGT | 213865 | 1.1025571 | 8.4350605 | 14 |
| GTCTG | 208805 | 1.0991541 | 8.518995 | 17 |
| AACTC | 350640 | 1.0871997 | 5.480659 | 22 |
| CATCT | 347305 | 1.0758425 | 5.1863747 | 39 |
| ACTCC | 209760 | 1.058078 | 8.142467 | 23 |
| GAACT | 333785 | 1.0577452 | 5.5442553 | 21 |
| ATCTC | 330955 | 1.0251954 | 5.206821 | 40 |
| ATGCC | 198485 | 1.0232673 | 7.9096622 | 47 |
| ACACG | 197740 | 1.0203898 | 8.370351 | 13 |
| TCACG | 197200 | 1.0166427 | 7.9783688 | 30 |
| GATCG | 190880 | 1.0057459 | 8.354105 | 1 |
| ACGTC | 191485 | 0.9871795 | 8.299691 | 15 |
| CAGTC | 188660 | 0.97261554 | 8.097546 | 27 |
| ACATC | 301285 | 0.9341688 | 5.0251822 | 38 |
| AGTCA | 283700 | 0.8990287 | 5.2044387 | 28 |
| GCTAC | 151045 | 0.7786956 | 7.533114 | 35 |
| GGCTA | 127270 | 0.670585 | 7.6619473 | 34 |