![[OK]](Icons/tick.png) Basic Statistics
Basic Statistics
| Measure | Value |
|---|---|
| Filename | LW190_GATCAG_L002_R1_001.fastq.gz |
| File type | Conventional base calls |
| Encoding | Sanger / Illumina 1.9 |
| Total Sequences | 6042676 |
| Filtered Sequences | 0 |
| Sequence length | 51 |
| %GC | 37 |
![[OK]](Icons/tick.png) Per base sequence quality
Per base sequence quality
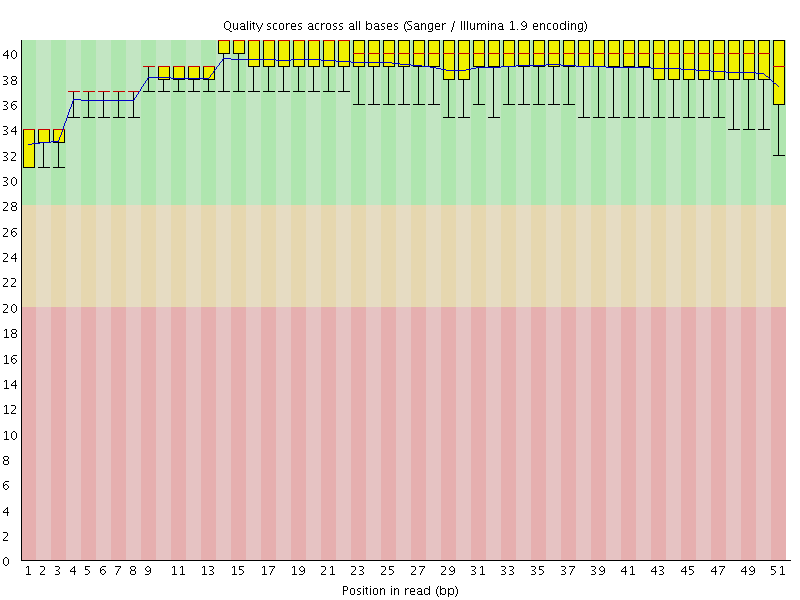
![[OK]](Icons/tick.png) Per sequence quality scores
Per sequence quality scores
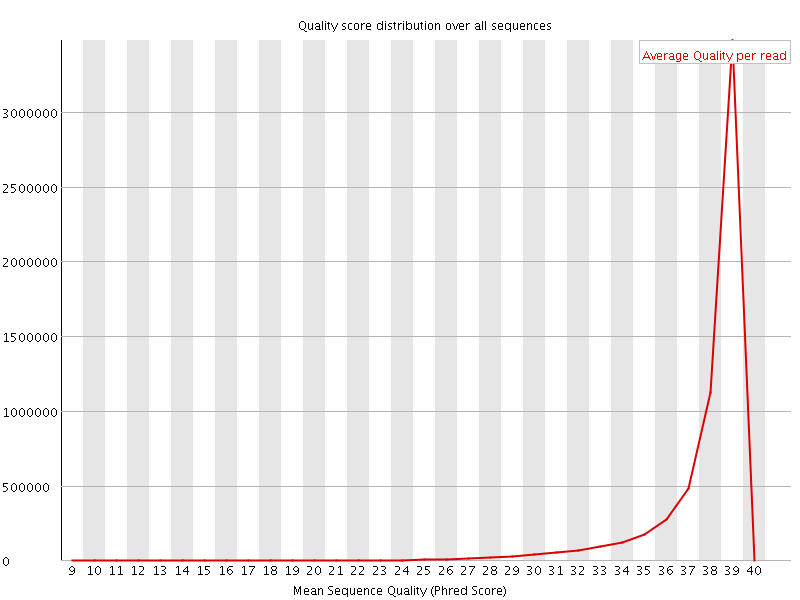
![[WARN]](Icons/warning.png) Per base sequence content
Per base sequence content
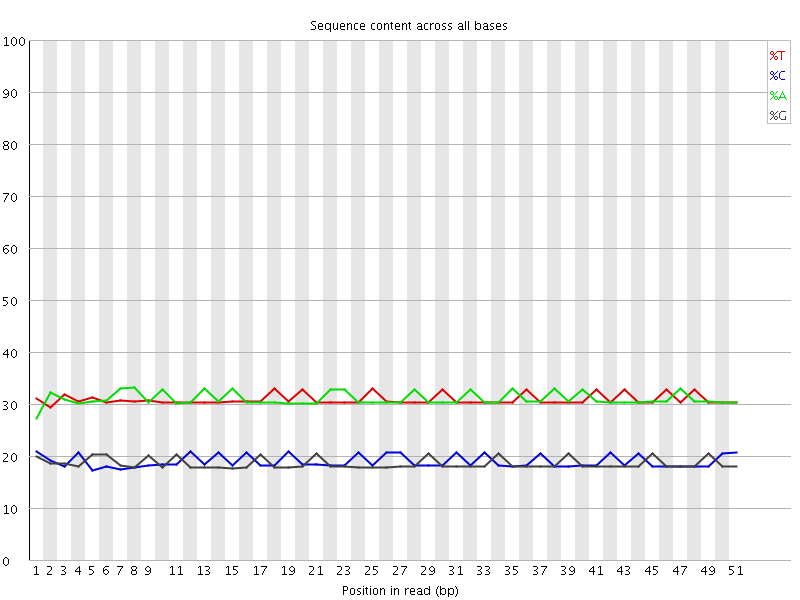
![[OK]](Icons/tick.png) Per base GC content
Per base GC content
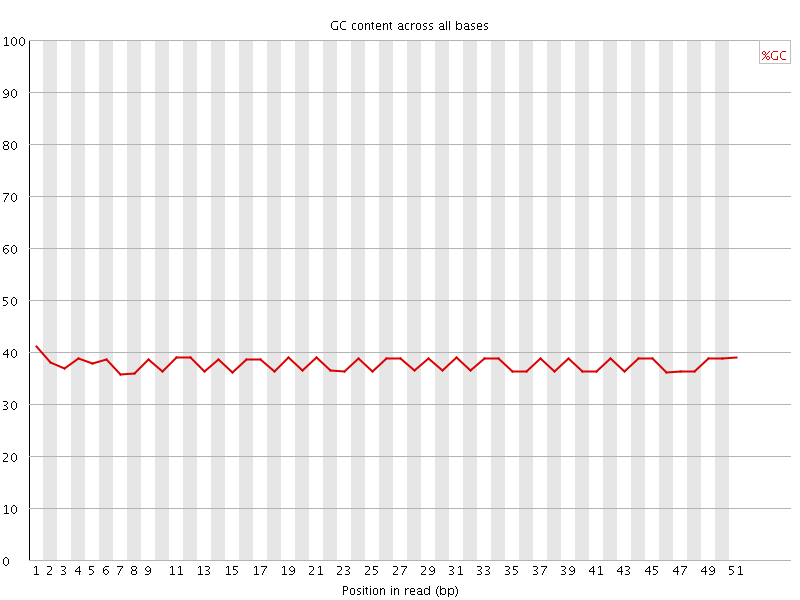
![[OK]](Icons/tick.png) Per sequence GC content
Per sequence GC content
![[OK]](Icons/tick.png) Per base N content
Per base N content
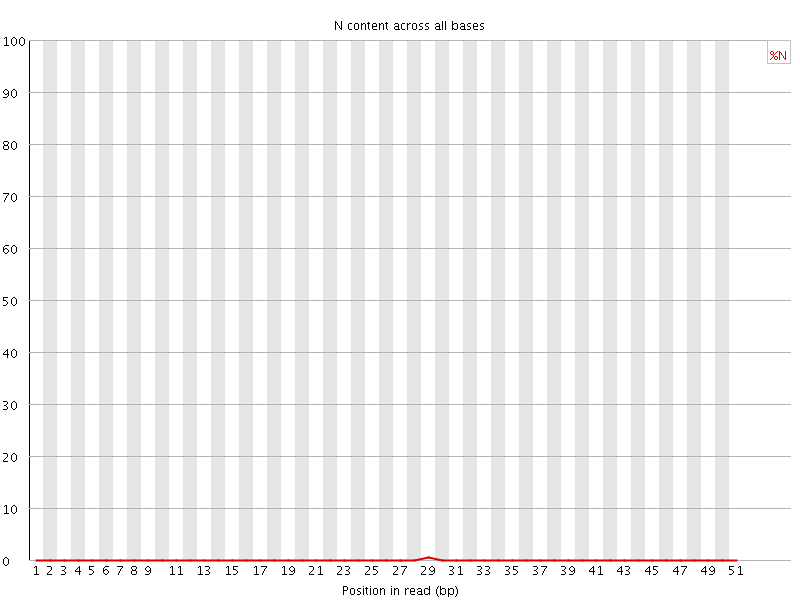
![[OK]](Icons/tick.png) Sequence Length Distribution
Sequence Length Distribution
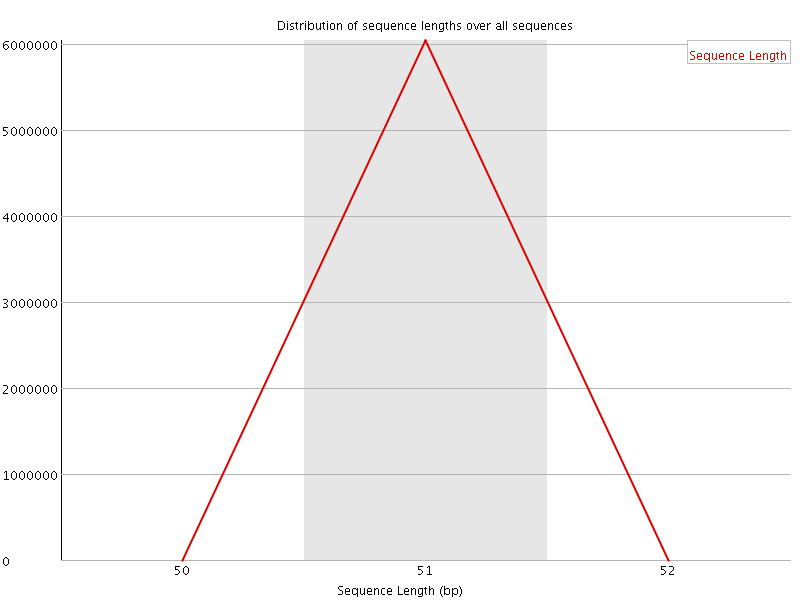
![[OK]](Icons/tick.png) Sequence Duplication Levels
Sequence Duplication Levels
![[FAIL]](Icons/error.png) Overrepresented sequences
Overrepresented sequences
| Sequence | Count | Percentage | Possible Source |
|---|---|---|---|
| GATCGGAAGAGCACACGTCTGAACTCCAGTCACGATCAGATCTCGTATGCC | 138092 | 2.285278906232934 | TruSeq Adapter, Index 9 (100% over 51bp) |
![[WARN]](Icons/warning.png) Kmer Content
Kmer Content
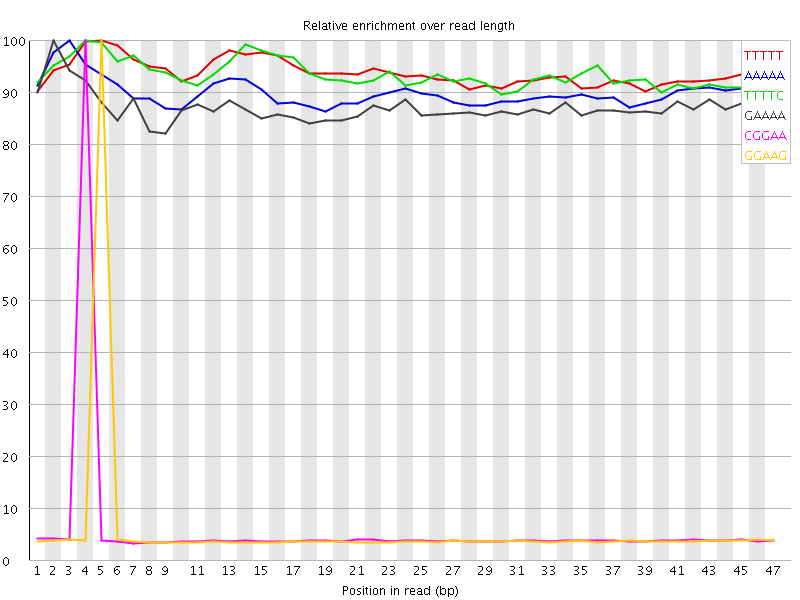
| Sequence | Count | Obs/Exp Overall | Obs/Exp Max | Max Obs/Exp Position |
|---|---|---|---|---|
| TTTTT | 3013165 | 3.70001 | 3.943509 | 5 |
| AAAAA | 3006825 | 3.5952463 | 3.9979515 | 3 |
| TTTTC | 1541620 | 3.0727687 | 3.2868805 | 4 |
| GAAAA | 1504890 | 3.001638 | 3.4497545 | 2 |
| CGGAA | 445240 | 2.4174929 | 40.705227 | 4 |
| GGAAG | 434680 | 2.4126217 | 41.486324 | 5 |
| GAGCA | 440430 | 2.391376 | 40.478355 | 9 |
| CTCCA | 456695 | 2.385679 | 38.340652 | 24 |
| TCCAG | 435710 | 2.326654 | 39.02992 | 25 |
| GAAGA | 671510 | 2.2342875 | 25.566467 | 6 |
| CTGAA | 681925 | 2.2314465 | 24.911064 | 19 |
| TCTCG | 390035 | 2.0938704 | 38.677826 | 41 |
| TCGGA | 378550 | 2.066361 | 40.50349 | 3 |
| GCACA | 365160 | 1.9395708 | 39.073177 | 11 |
| CACAC | 370845 | 1.9269316 | 38.128338 | 12 |
| AGAGC | 352700 | 1.9150339 | 39.96599 | 8 |
| ATCGG | 338125 | 1.8456963 | 40.07424 | 2 |
| AGCAC | 345445 | 1.8348533 | 39.019566 | 10 |
| TCAGA | 559515 | 1.8308873 | 23.845114 | 36 |
| TCTGA | 553840 | 1.8219903 | 24.683342 | 18 |
| CCAGT | 340460 | 1.8180274 | 38.418896 | 26 |
| CACGA | 333800 | 1.7730001 | 37.82265 | 31 |
| GTCTG | 318925 | 1.7501825 | 39.8547 | 17 |
| GTCAC | 327535 | 1.7490088 | 38.33936 | 29 |
| CTCGT | 325525 | 1.7475538 | 38.23951 | 42 |
| CGTCT | 322980 | 1.7338911 | 38.95899 | 16 |
| CACGT | 319525 | 1.7062362 | 38.785355 | 14 |
| ACTCC | 317270 | 1.6573522 | 37.70424 | 23 |
| GATCG | 301815 | 1.6474934 | 39.840305 | 1 |
| ATGCC | 307530 | 1.6421841 | 38.05601 | 47 |
| ACACG | 308755 | 1.6399722 | 38.596893 | 13 |
| TGAAC | 498810 | 1.6322438 | 24.335428 | 20 |
| TCACG | 305190 | 1.6296884 | 38.08363 | 30 |
| ACGTC | 299745 | 1.6006128 | 38.681957 | 15 |
| CGATC | 298985 | 1.5965544 | 37.63794 | 33 |
| CAGTC | 298070 | 1.5916684 | 38.18534 | 27 |
| AAGAG | 477980 | 1.590363 | 24.92211 | 7 |
| AACTC | 453815 | 1.4527142 | 23.580727 | 22 |
| GAACT | 440450 | 1.4412738 | 24.071922 | 21 |
| ATCAG | 435385 | 1.4246999 | 23.379322 | 35 |
| CAGAT | 428635 | 1.4026119 | 23.364056 | 37 |
| GATCA | 428205 | 1.4012047 | 23.389214 | 34 |
| ATCTC | 435285 | 1.4008349 | 23.292152 | 40 |
| AGTCA | 397595 | 1.3010404 | 23.731688 | 28 |
| AGATC | 388020 | 1.2697084 | 23.362928 | 38 |
| GATCT | 383465 | 1.2615006 | 23.498968 | 39 |
| ACGAT | 379260 | 1.2410432 | 23.451145 | 32 |
| TCGTA | 328425 | 1.0804332 | 23.507912 | 43 |
| TATGC | 318950 | 1.0492631 | 23.471275 | 46 |
| GTATG | 311505 | 1.0475513 | 23.982693 | 45 |
| CGTAT | 300075 | 0.9871691 | 23.417772 | 44 |