![[OK]](Icons/tick.png) Basic Statistics
Basic Statistics
| Measure | Value |
|---|---|
| Filename | LW189_ACTTGA_L002_R1_001.fastq.gz |
| File type | Conventional base calls |
| Encoding | Sanger / Illumina 1.9 |
| Total Sequences | 4268679 |
| Filtered Sequences | 0 |
| Sequence length | 51 |
| %GC | 42 |
![[OK]](Icons/tick.png) Per base sequence quality
Per base sequence quality
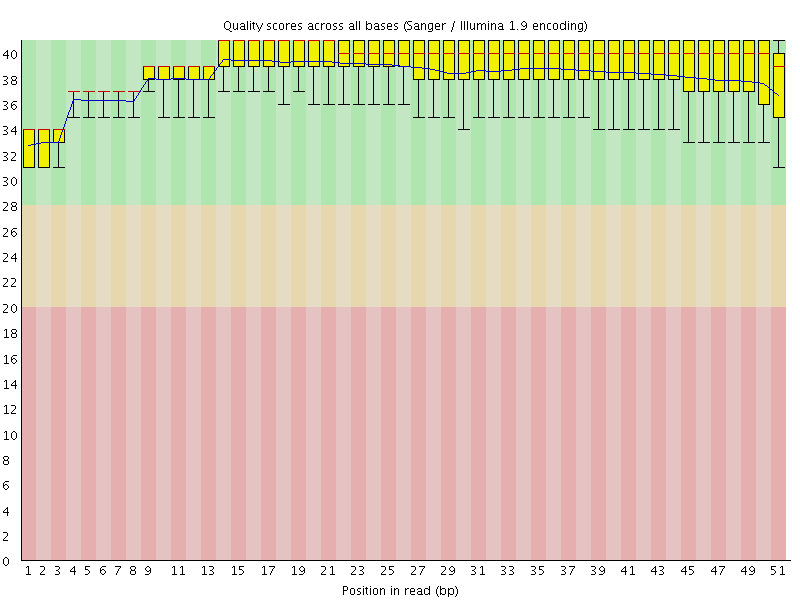
![[OK]](Icons/tick.png) Per sequence quality scores
Per sequence quality scores
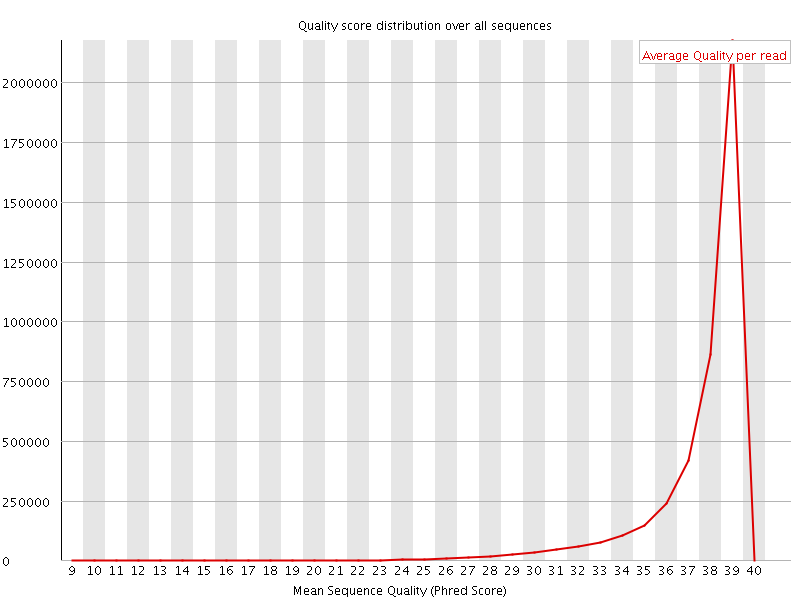
![[WARN]](Icons/warning.png) Per base sequence content
Per base sequence content
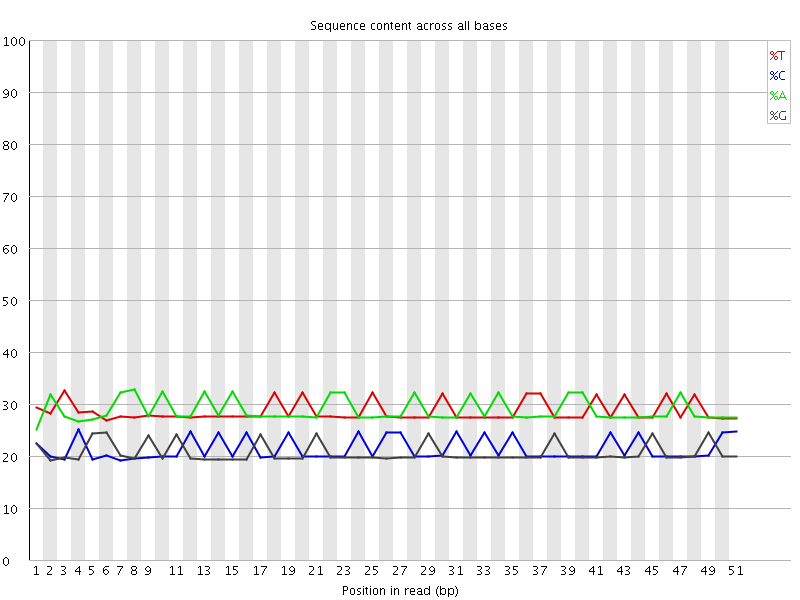
![[OK]](Icons/tick.png) Per base GC content
Per base GC content
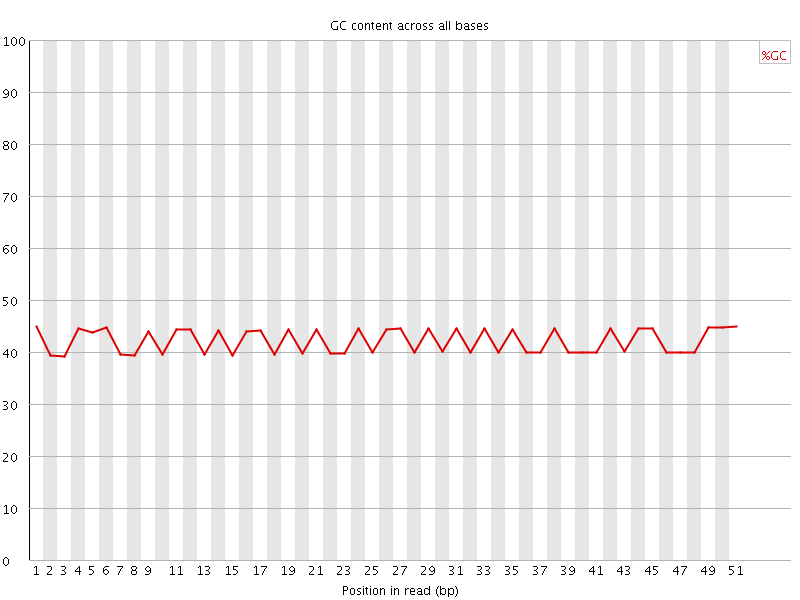
![[FAIL]](Icons/error.png) Per sequence GC content
Per sequence GC content
![[OK]](Icons/tick.png) Per base N content
Per base N content
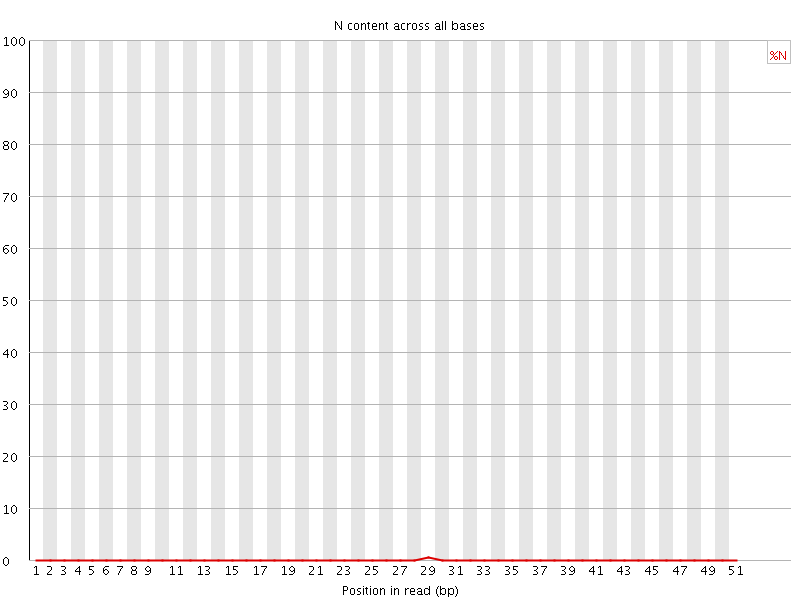
![[OK]](Icons/tick.png) Sequence Length Distribution
Sequence Length Distribution
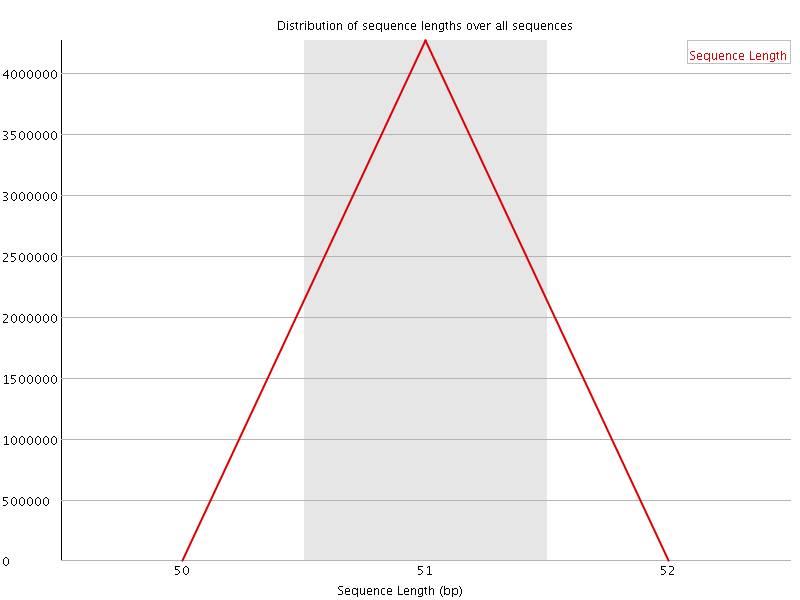
![[WARN]](Icons/warning.png) Sequence Duplication Levels
Sequence Duplication Levels
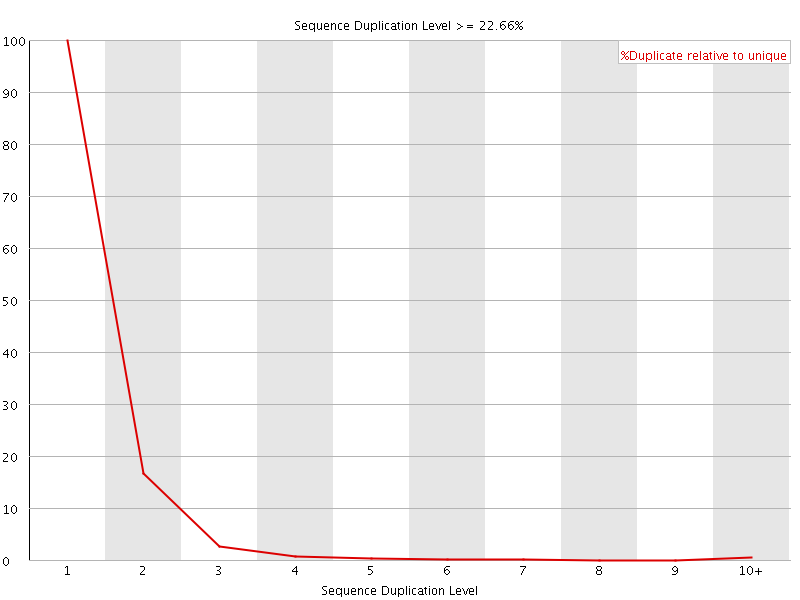
![[FAIL]](Icons/error.png) Overrepresented sequences
Overrepresented sequences
| Sequence | Count | Percentage | Possible Source |
|---|---|---|---|
| GATCGGAAGAGCACACGTCTGAACTCCAGTCACACTTGAATCTCGTATGCC | 180813 | 4.2358069088821155 | TruSeq Adapter, Index 8 (100% over 51bp) |
![[WARN]](Icons/warning.png) Kmer Content
Kmer Content
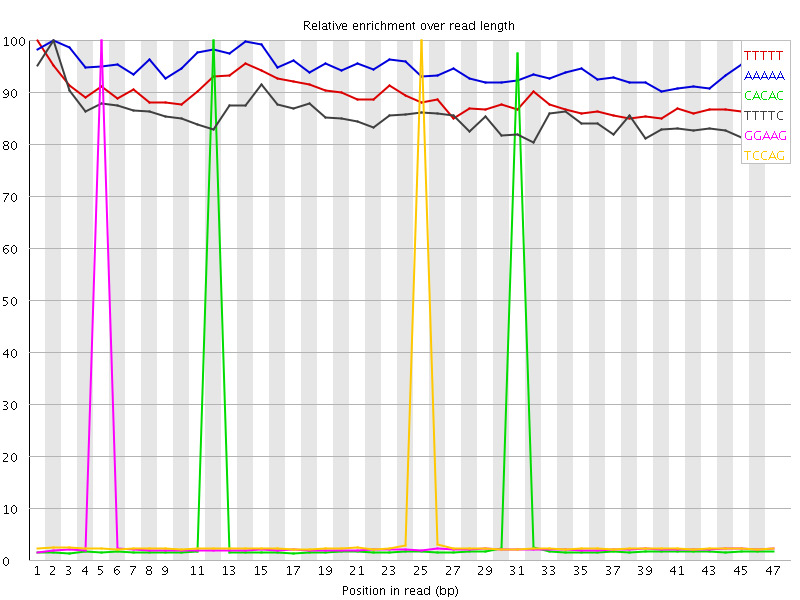
| Sequence | Count | Obs/Exp Overall | Obs/Exp Max | Max Obs/Exp Position |
|---|---|---|---|---|
| TTTTT | 1629970 | 4.1581173 | 4.6633434 | 1 |
| AAAAA | 1643230 | 4.0451837 | 4.2802978 | 2 |
| CACAC | 560270 | 3.3568013 | 57.171627 | 12 |
| TTTTC | 882365 | 3.0063028 | 3.5158784 | 2 |
| GGAAG | 410300 | 2.7046068 | 63.730003 | 5 |
| TCCAG | 423970 | 2.6410925 | 58.998367 | 25 |
| CGGAA | 410585 | 2.6216877 | 61.58487 | 4 |
| GAGCA | 405250 | 2.5876222 | 61.524914 | 9 |
| GAAGA | 538185 | 2.5547397 | 46.496895 | 6 |
| TTGAA | 725075 | 2.5142086 | 33.308796 | 36 |
| CTGAA | 530405 | 2.4563675 | 44.809 | 19 |
| CTCCA | 404995 | 2.4438436 | 57.08413 | 24 |
| TCTCG | 380185 | 2.3852775 | 58.654545 | 41 |
| GATCG | 367065 | 2.3605669 | 61.45989 | 1 |
| ATCGG | 359560 | 2.3123024 | 61.664577 | 2 |
| TCGGA | 350850 | 2.2562895 | 61.68874 | 3 |
| AGAGC | 345690 | 2.2073169 | 61.18916 | 8 |
| CCAGT | 353800 | 2.2039733 | 58.505035 | 26 |
| GCACA | 353570 | 2.1868975 | 59.202316 | 11 |
| AGCAC | 353410 | 2.1859083 | 59.227005 | 10 |
| GTCTG | 335900 | 2.1755984 | 61.310417 | 17 |
| ATGCC | 346695 | 2.159713 | 57.98769 | 47 |
| CGTCT | 336620 | 2.111951 | 59.336235 | 16 |
| GTCAC | 330990 | 2.06188 | 58.164856 | 29 |
| TCTGA | 441835 | 2.0608258 | 44.745148 | 18 |
| CTCGT | 325235 | 2.0405216 | 58.227028 | 42 |
| ACGTC | 321135 | 2.000489 | 58.979168 | 15 |
| CACGT | 319580 | 1.990802 | 59.131535 | 14 |
| AAGAG | 419020 | 1.9890689 | 45.84168 | 7 |
| TGAAC | 422355 | 1.9559753 | 44.252052 | 20 |
| CAGTC | 308335 | 1.9207523 | 58.095642 | 27 |
| ACACG | 310495 | 1.9204706 | 58.7377 | 13 |
| ACTCC | 313390 | 1.8910755 | 56.842754 | 23 |
| CTTGA | 398425 | 1.858351 | 43.39415 | 35 |
| CACTT | 407840 | 1.8426642 | 42.269974 | 33 |
| TGAAT | 524815 | 1.8198041 | 32.666103 | 37 |
| TCACA | 405605 | 1.8195511 | 42.094208 | 30 |
| ATCTC | 399830 | 1.806474 | 42.325783 | 40 |
| GAACT | 387790 | 1.7959007 | 44.08461 | 21 |
| GAATC | 379020 | 1.7552859 | 42.926514 | 38 |
| AACTC | 387705 | 1.7392515 | 42.64823 | 22 |
| ACTTG | 368900 | 1.7206392 | 43.35785 | 34 |
| AGTCA | 351230 | 1.6265872 | 43.2861 | 28 |
| ACACT | 347290 | 1.557949 | 41.72263 | 32 |
| AATCT | 451450 | 1.5163633 | 31.434357 | 39 |
| TATGC | 318085 | 1.4836257 | 43.179348 | 46 |
| GTATG | 305750 | 1.4722191 | 44.59882 | 45 |
| TCGTA | 308640 | 1.4395719 | 43.237053 | 43 |
| CGTAT | 299570 | 1.3972673 | 43.168514 | 44 |