![[OK]](Icons/tick.png) Basic Statistics
Basic Statistics
| Measure | Value |
|---|---|
| Filename | LW188_CAGATC_L002_R1_001.fastq.gz |
| File type | Conventional base calls |
| Encoding | Sanger / Illumina 1.9 |
| Total Sequences | 6425498 |
| Filtered Sequences | 0 |
| Sequence length | 51 |
| %GC | 39 |
![[OK]](Icons/tick.png) Per base sequence quality
Per base sequence quality
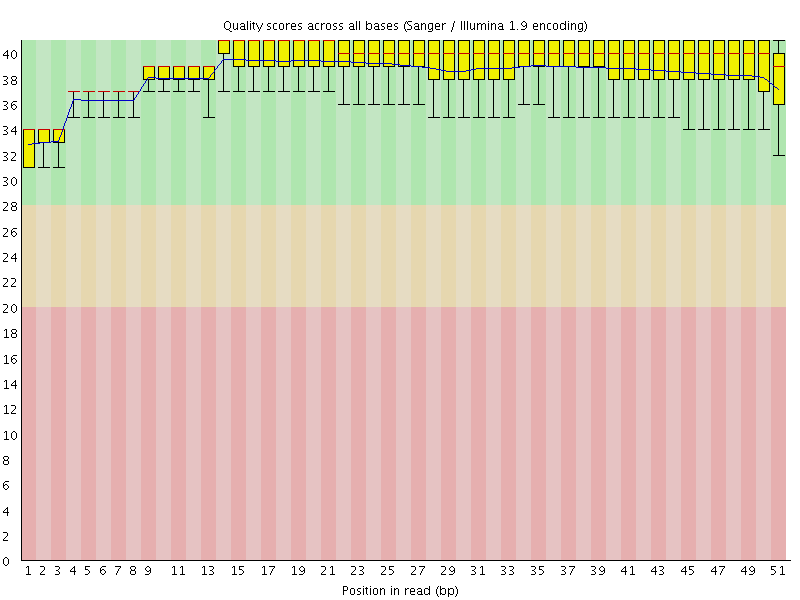
![[OK]](Icons/tick.png) Per sequence quality scores
Per sequence quality scores
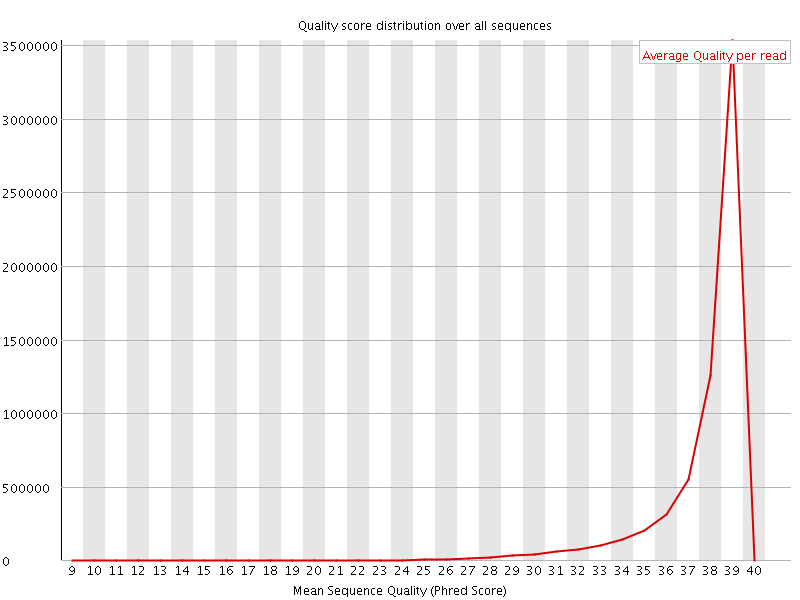
![[WARN]](Icons/warning.png) Per base sequence content
Per base sequence content
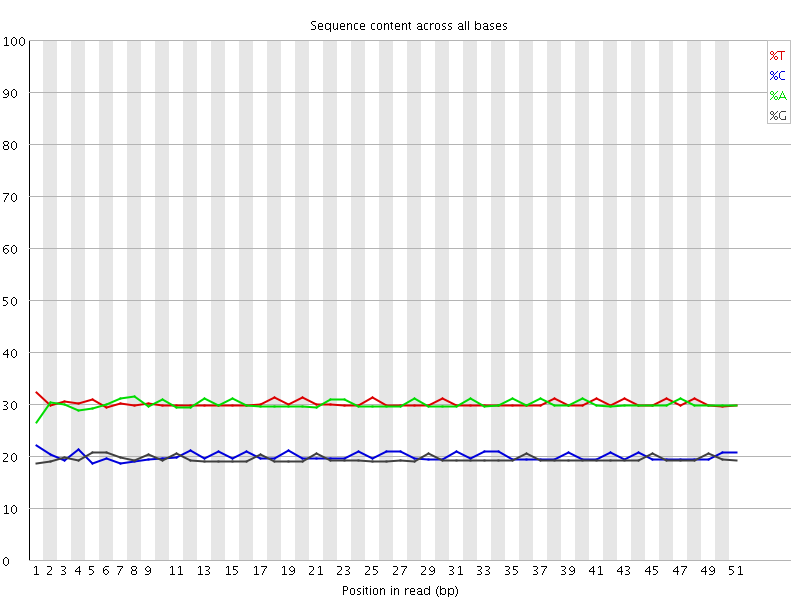
![[OK]](Icons/tick.png) Per base GC content
Per base GC content
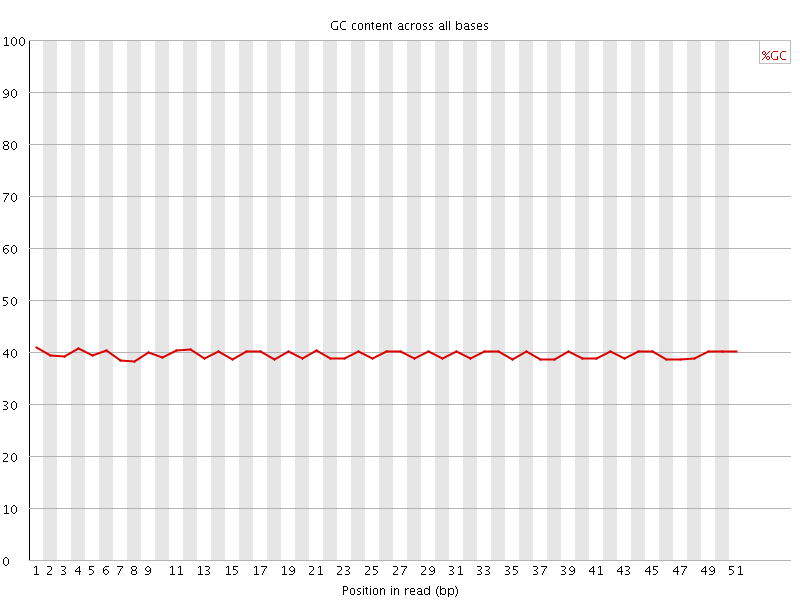
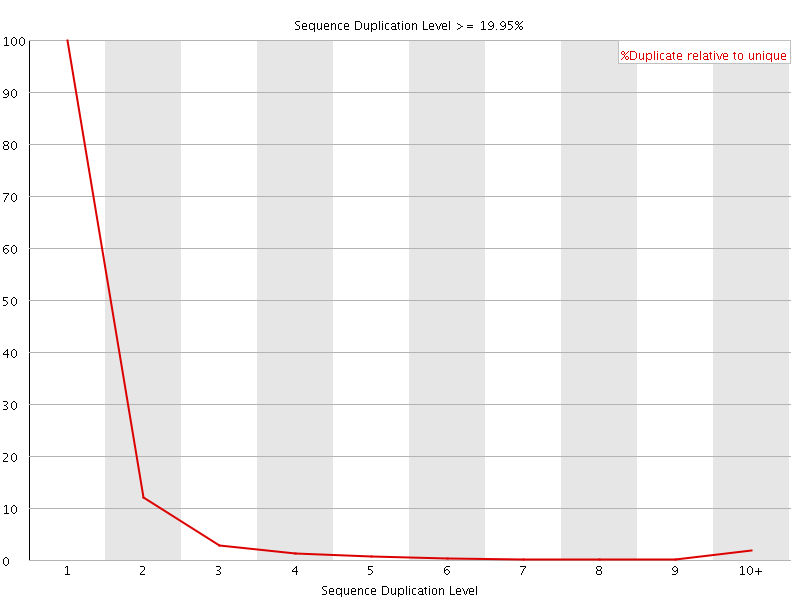
![[OK]](Icons/tick.png) Per sequence GC content
Per sequence GC content
![[OK]](Icons/tick.png) Per base N content
Per base N content
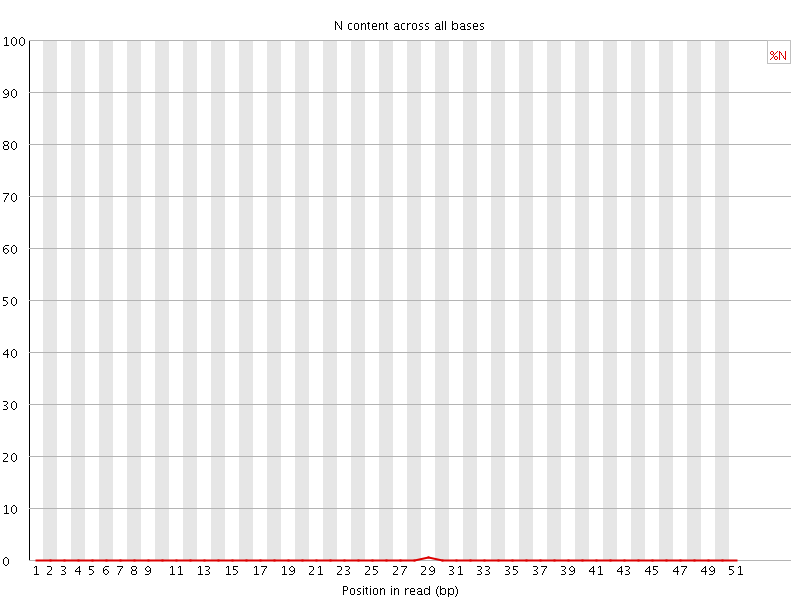
![[OK]](Icons/tick.png) Sequence Length Distribution
Sequence Length Distribution
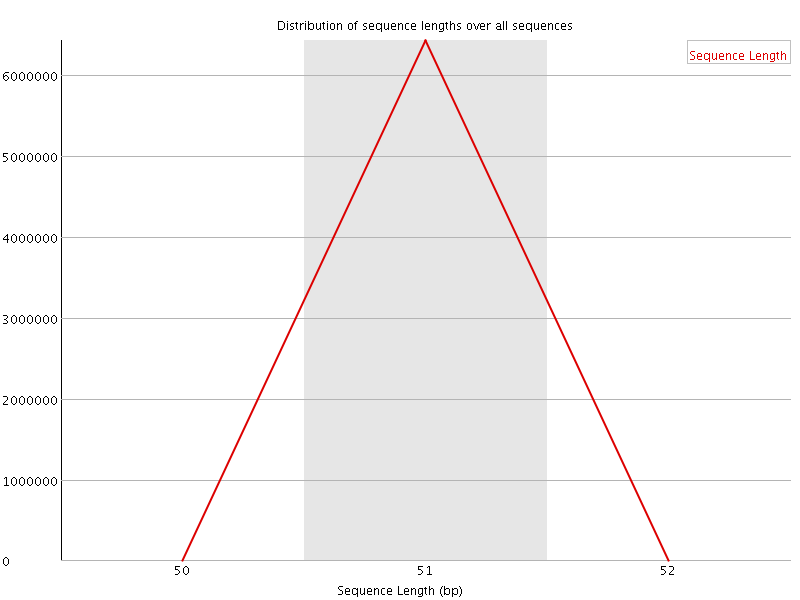
![[OK]](Icons/tick.png) Sequence Duplication Levels
Sequence Duplication Levels
![[FAIL]](Icons/error.png) Overrepresented sequences
Overrepresented sequences
| Sequence | Count | Percentage | Possible Source |
|---|---|---|---|
| GATCGGAAGAGCACACGTCTGAACTCCAGTCACCAGATCATCTCGTATGCC | 80979 | 1.260275857217604 | TruSeq Adapter, Index 7 (100% over 51bp) |
![[WARN]](Icons/warning.png) Kmer Content
Kmer Content
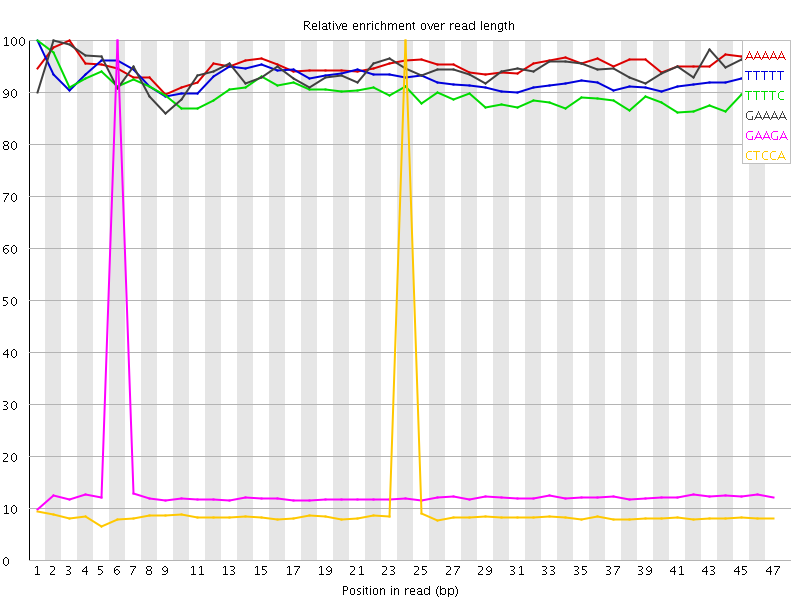
| Sequence | Count | Obs/Exp Overall | Obs/Exp Max | Max Obs/Exp Position |
|---|---|---|---|---|
| AAAAA | 2786050 | 3.7494059 | 3.940054 | 3 |
| TTTTT | 2855715 | 3.7298975 | 4.0271597 | 1 |
| TTTTC | 1557080 | 3.063837 | 3.4103289 | 1 |
| GAAAA | 1477500 | 3.0565476 | 3.2512226 | 2 |
| GAAGA | 695135 | 2.2105634 | 15.839108 | 6 |
| CTCCA | 474755 | 2.132888 | 20.731007 | 24 |
| TCCAG | 437760 | 2.018786 | 21.161692 | 25 |
| GAGCA | 423550 | 2.0170324 | 22.233793 | 9 |
| GGAAG | 409370 | 2.001151 | 23.129204 | 5 |
| CGGAA | 418070 | 1.9909354 | 22.502092 | 4 |
| CACCA | 436820 | 1.9742372 | 20.374052 | 31 |
| CTGAA | 631645 | 1.945149 | 14.9916 | 19 |
| CCAGA | 388785 | 1.8036906 | 20.641766 | 33 |
| TCTCG | 385480 | 1.7670863 | 20.691963 | 41 |
| TCGGA | 356780 | 1.688925 | 22.017828 | 3 |
| TCATC | 540300 | 1.6112418 | 13.688668 | 38 |
| CACAC | 356450 | 1.6109996 | 20.738121 | 12 |
| TCACC | 355220 | 1.5958642 | 19.905846 | 30 |
| AGAGC | 326415 | 1.5544554 | 21.798937 | 8 |
| TCTGA | 502020 | 1.5367473 | 14.454426 | 18 |
| ATCGG | 317650 | 1.5036914 | 21.657793 | 2 |
| GCACA | 323470 | 1.5006747 | 21.230894 | 11 |
| AAGAG | 468195 | 1.4888829 | 15.06574 | 7 |
| GATCG | 307000 | 1.4532765 | 21.379467 | 1 |
| AGCAC | 308645 | 1.4318969 | 21.140501 | 10 |
| CTCGT | 312275 | 1.4315057 | 20.313204 | 42 |
| CGTCT | 308390 | 1.4136964 | 20.715092 | 16 |
| CCAGT | 305605 | 1.4093364 | 20.514053 | 26 |
| TGAAC | 451725 | 1.3910859 | 14.40585 | 20 |
| ACCAG | 293155 | 1.3600342 | 20.262043 | 32 |
| CACGT | 293030 | 1.3513452 | 20.81521 | 14 |
| GTCTG | 287100 | 1.350967 | 21.205126 | 17 |
| CATCT | 441220 | 1.315773 | 13.454437 | 39 |
| ATCTC | 434825 | 1.2967024 | 13.579215 | 40 |
| GTCAC | 280970 | 1.295729 | 20.31451 | 29 |
| ACGTC | 276415 | 1.274723 | 20.654282 | 15 |
| ACACG | 273165 | 1.2672944 | 20.953184 | 13 |
| ACTCC | 280575 | 1.2605134 | 19.97256 | 23 |
| AACTC | 418460 | 1.2553881 | 13.804623 | 22 |
| GATCA | 404860 | 1.246765 | 13.768049 | 36 |
| GAACT | 403010 | 1.241068 | 14.220758 | 21 |
| ATGCC | 267150 | 1.2319963 | 20.149046 | 47 |
| CAGAT | 396790 | 1.2219136 | 13.699462 | 34 |
| CAGTC | 259835 | 1.1982622 | 20.311241 | 27 |
| AGATC | 382770 | 1.1787391 | 13.673578 | 35 |
| ATCAT | 529205 | 1.0538416 | 9.192067 | 37 |
| AGTCA | 329475 | 1.0146172 | 13.738659 | 28 |
| TCGTA | 284555 | 0.8710592 | 13.447335 | 43 |
| GTATG | 260090 | 0.8172611 | 13.744141 | 45 |
| TATGC | 265380 | 0.8123621 | 13.36446 | 46 |
| CGTAT | 242425 | 0.74209386 | 13.323209 | 44 |