![[OK]](Icons/tick.png) Basic Statistics
Basic Statistics
| Measure | Value |
|---|---|
| Filename | LW187_GCCAAT_L002_R1_001.fastq.gz |
| File type | Conventional base calls |
| Encoding | Sanger / Illumina 1.9 |
| Total Sequences | 5486215 |
| Filtered Sequences | 0 |
| Sequence length | 51 |
| %GC | 43 |
![[OK]](Icons/tick.png) Per base sequence quality
Per base sequence quality
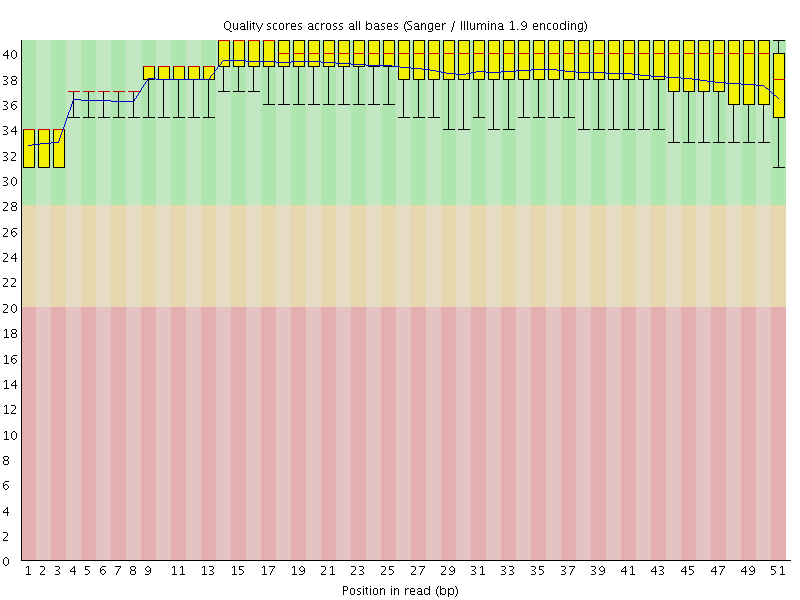
![[OK]](Icons/tick.png) Per sequence quality scores
Per sequence quality scores
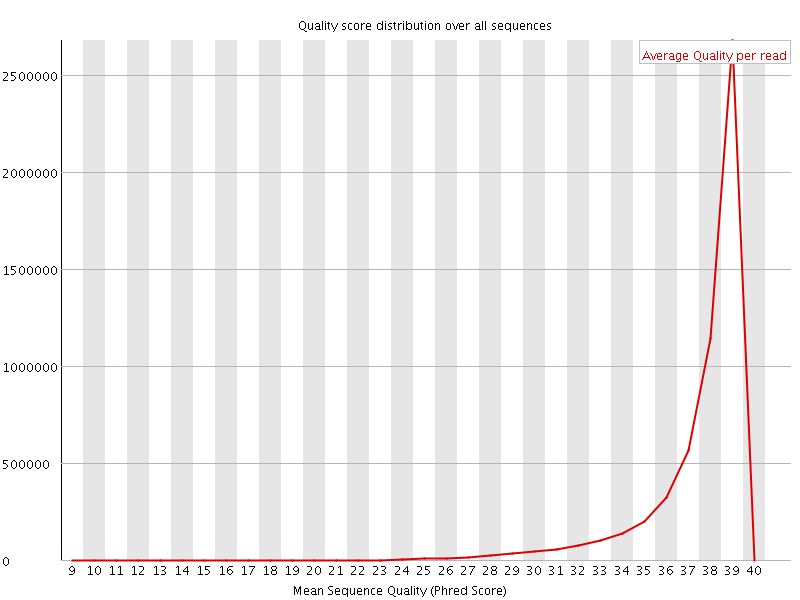
![[WARN]](Icons/warning.png) Per base sequence content
Per base sequence content
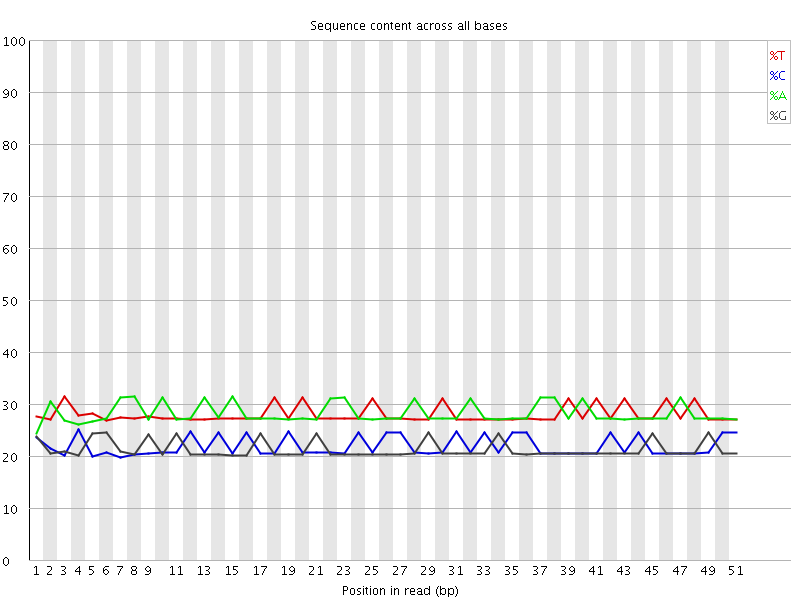
![[OK]](Icons/tick.png) Per base GC content
Per base GC content
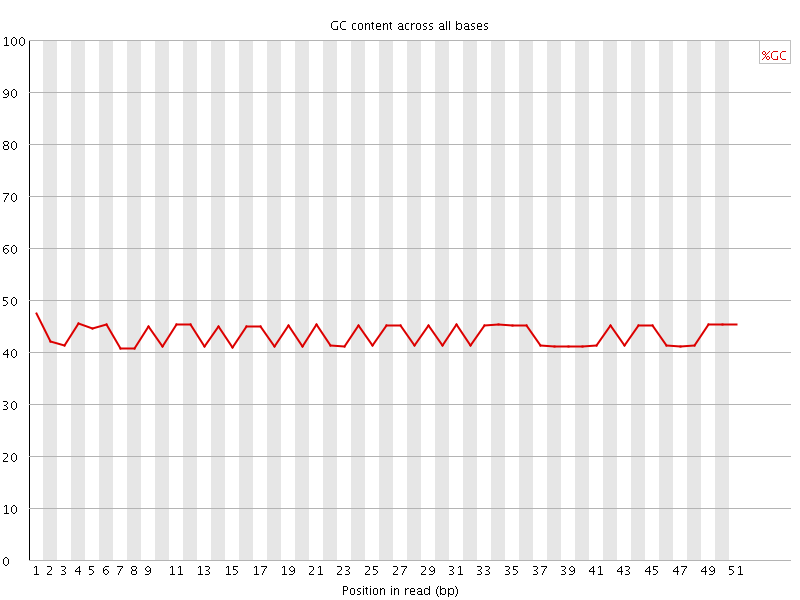
![[FAIL]](Icons/error.png) Per sequence GC content
Per sequence GC content
![[OK]](Icons/tick.png) Per base N content
Per base N content
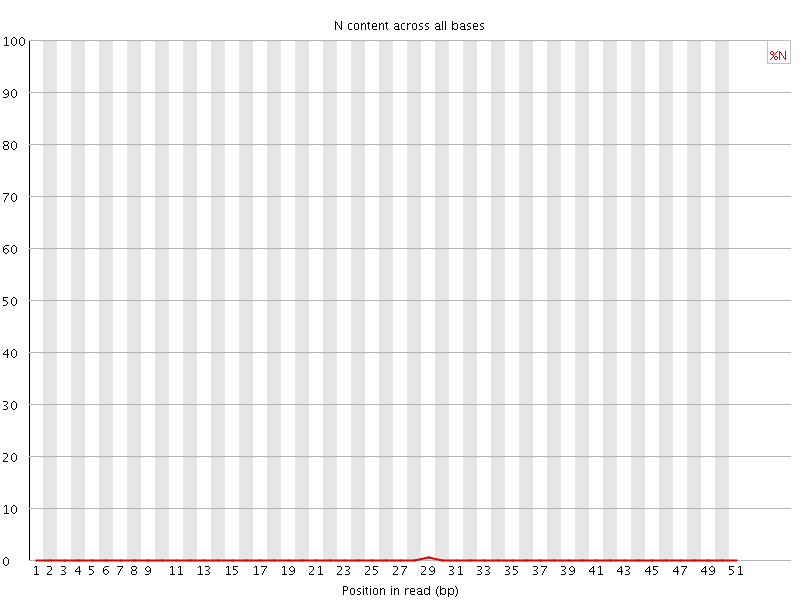
![[OK]](Icons/tick.png) Sequence Length Distribution
Sequence Length Distribution
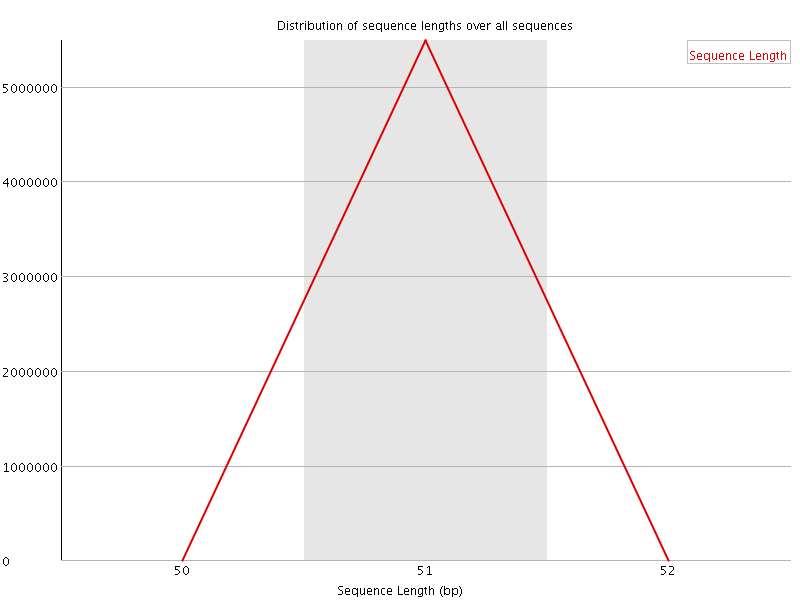
![[WARN]](Icons/warning.png) Sequence Duplication Levels
Sequence Duplication Levels
![[FAIL]](Icons/error.png) Overrepresented sequences
Overrepresented sequences
| Sequence | Count | Percentage | Possible Source |
|---|---|---|---|
| GATCGGAAGAGCACACGTCTGAACTCCAGTCACGCCAATATCTCGTATGCC | 196508 | 3.581850146230142 | TruSeq Adapter, Index 6 (100% over 51bp) |
![[WARN]](Icons/warning.png) Kmer Content
Kmer Content
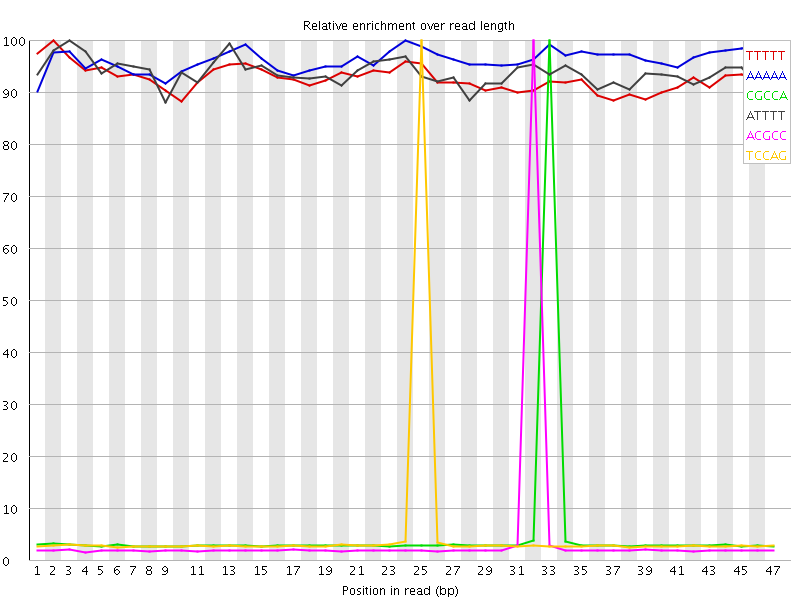
| Sequence | Count | Obs/Exp Overall | Obs/Exp Max | Max Obs/Exp Position |
|---|---|---|---|---|
| TTTTT | 1969210 | 4.324124 | 4.663838 | 2 |
| AAAAA | 1946790 | 4.1334367 | 4.292755 | 24 |
| CGCCA | 528315 | 3.1442087 | 61.943253 | 33 |
| ATTTT | 1388275 | 3.0280204 | 3.2255862 | 3 |
| ACGCC | 426640 | 2.5391011 | 61.5065 | 32 |
| TCCAG | 524930 | 2.4477513 | 49.24925 | 25 |
| GGAAG | 495370 | 2.4261787 | 53.094627 | 5 |
| CACGC | 398990 | 2.374545 | 61.732967 | 31 |
| CGGAA | 495180 | 2.3584745 | 51.62496 | 4 |
| GAAGA | 636215 | 2.3582869 | 40.690136 | 6 |
| CTGAA | 649565 | 2.3572907 | 39.27357 | 19 |
| GAGCA | 483810 | 2.3043208 | 51.4785 | 9 |
| GCCAA | 481485 | 2.2301078 | 47.946358 | 34 |
| GATCG | 461545 | 2.2131195 | 51.433792 | 1 |
| CTCCA | 468325 | 2.1236758 | 47.448715 | 24 |
| ATCGG | 439480 | 2.1073174 | 51.52446 | 2 |
| CCAGT | 443905 | 2.0699313 | 48.743286 | 26 |
| TCTCG | 437050 | 2.051728 | 48.94464 | 41 |
| ATGCC | 436105 | 2.0335598 | 48.718445 | 47 |
| AGCAC | 425135 | 1.9691098 | 49.651264 | 10 |
| GTCTG | 407000 | 1.9647534 | 51.08636 | 17 |
| AGAGC | 407925 | 1.9428909 | 51.072655 | 8 |
| GCACA | 418185 | 1.9369193 | 49.622997 | 11 |
| TCGGA | 400385 | 1.9198558 | 51.4648 | 3 |
| TCTGA | 523335 | 1.9120232 | 39.10732 | 18 |
| TCACG | 400465 | 1.8673702 | 48.65694 | 30 |
| TGAAC | 513295 | 1.8627628 | 38.681454 | 20 |
| GTCAC | 396355 | 1.8482053 | 48.770798 | 29 |
| CACAC | 406475 | 1.8308467 | 47.904663 | 12 |
| CGTCT | 388250 | 1.8226367 | 49.500305 | 16 |
| AAGAG | 483190 | 1.7910624 | 40.054085 | 7 |
| ACGTC | 381775 | 1.7802187 | 49.26899 | 15 |
| CTCGT | 374875 | 1.7598479 | 48.53626 | 42 |
| CACGT | 373185 | 1.7401636 | 49.280533 | 14 |
| CAGTC | 370215 | 1.7263144 | 48.572384 | 27 |
| CCAAT | 486395 | 1.716543 | 36.67847 | 35 |
| GAACT | 465915 | 1.6908194 | 38.49316 | 21 |
| ATCTC | 473110 | 1.6809332 | 37.11656 | 40 |
| ACTCC | 364005 | 1.6506244 | 47.12669 | 23 |
| ACACG | 356235 | 1.6499835 | 49.04458 | 13 |
| AATAT | 758240 | 1.6317153 | 23.169407 | 37 |
| AACTC | 454480 | 1.6039112 | 37.18652 | 22 |
| AGTCA | 413875 | 1.5019647 | 38.035286 | 28 |
| CAATA | 525585 | 1.4435545 | 28.708775 | 36 |
| TATGC | 382935 | 1.3990667 | 37.922802 | 46 |
| ATATC | 494545 | 1.3674732 | 28.90554 | 38 |
| GTATG | 363500 | 1.3656604 | 38.95145 | 45 |
| TCGTA | 362875 | 1.3257766 | 37.862198 | 43 |
| CGTAT | 354765 | 1.2961464 | 37.79142 | 44 |
| TATCT | 440405 | 1.2259933 | 28.975912 | 39 |