![[OK]](Icons/tick.png) Basic Statistics
Basic Statistics
| Measure | Value |
|---|---|
| Filename | LW186_ACAGTG_L002_R1_001.fastq.gz |
| File type | Conventional base calls |
| Encoding | Sanger / Illumina 1.9 |
| Total Sequences | 10193698 |
| Filtered Sequences | 0 |
| Sequence length | 51 |
| %GC | 39 |
![[OK]](Icons/tick.png) Per base sequence quality
Per base sequence quality
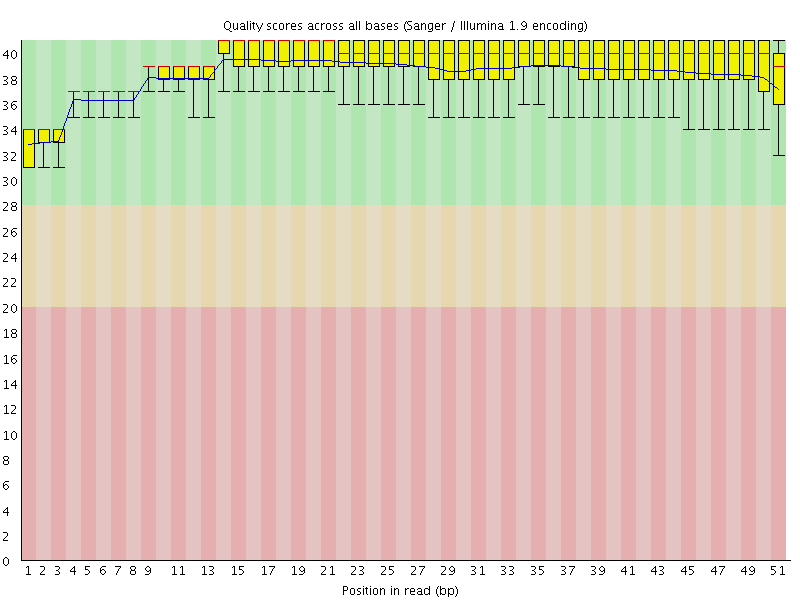
![[OK]](Icons/tick.png) Per sequence quality scores
Per sequence quality scores
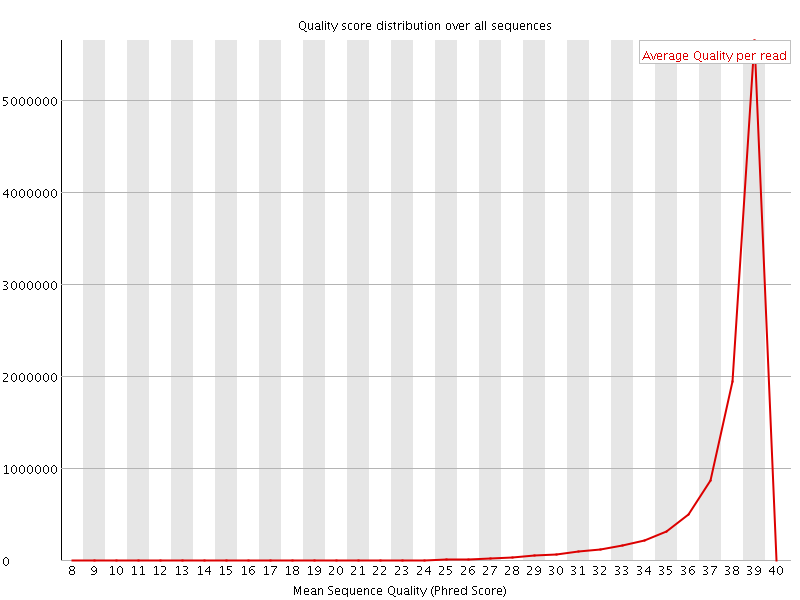
![[WARN]](Icons/warning.png) Per base sequence content
Per base sequence content
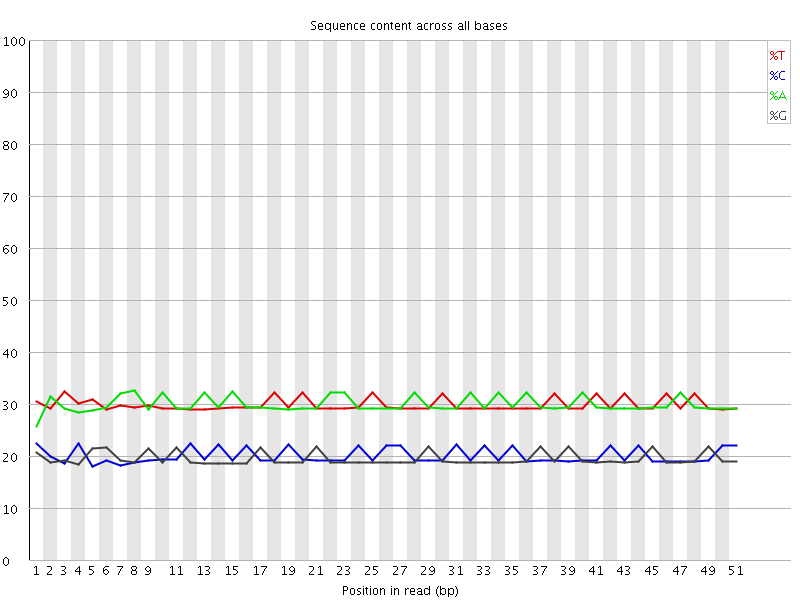
![[OK]](Icons/tick.png) Per base GC content
Per base GC content
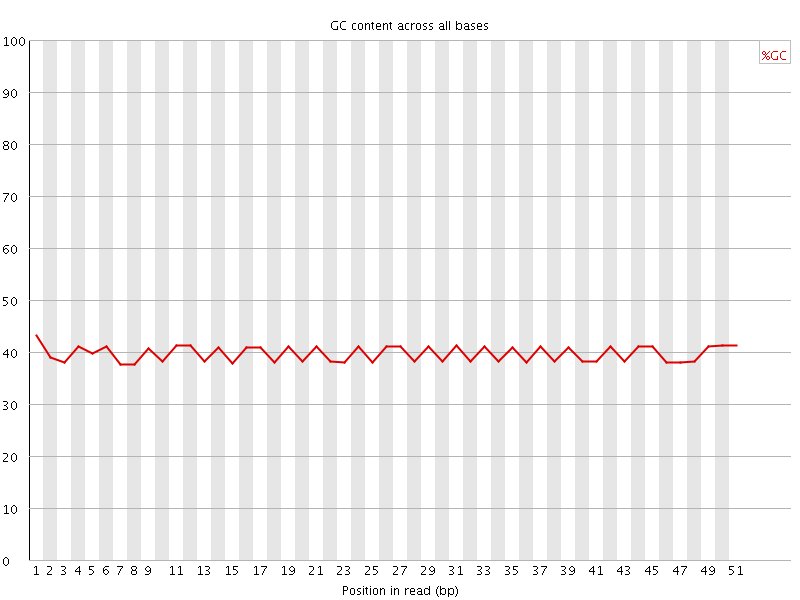
![[OK]](Icons/tick.png) Per sequence GC content
Per sequence GC content
![[OK]](Icons/tick.png) Per base N content
Per base N content
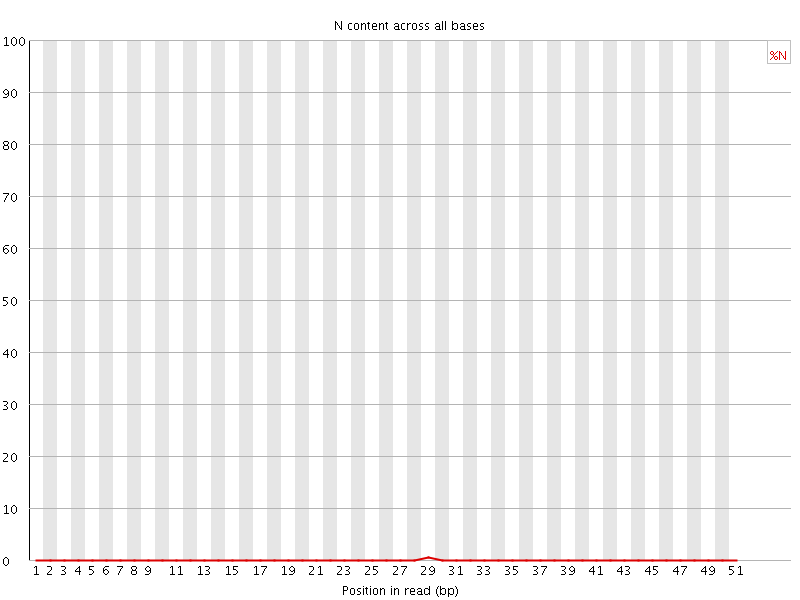
![[OK]](Icons/tick.png) Sequence Length Distribution
Sequence Length Distribution
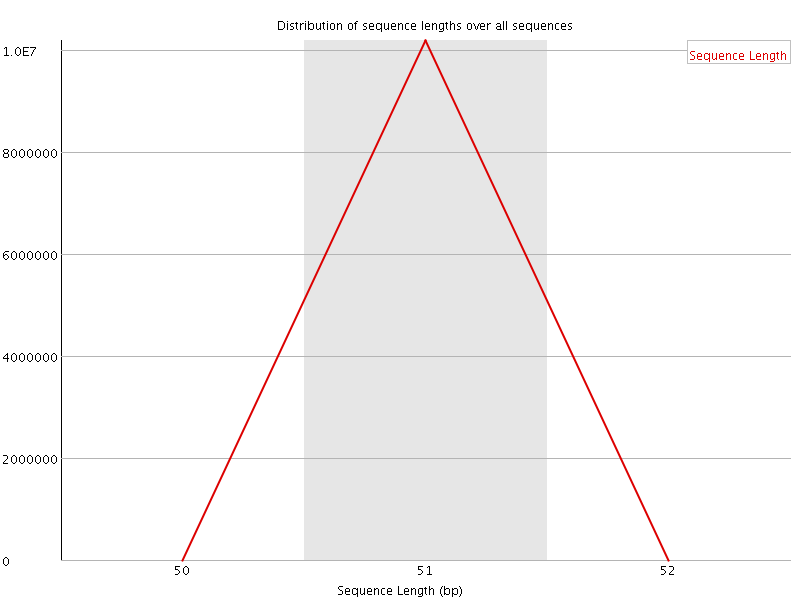
![[WARN]](Icons/warning.png) Sequence Duplication Levels
Sequence Duplication Levels
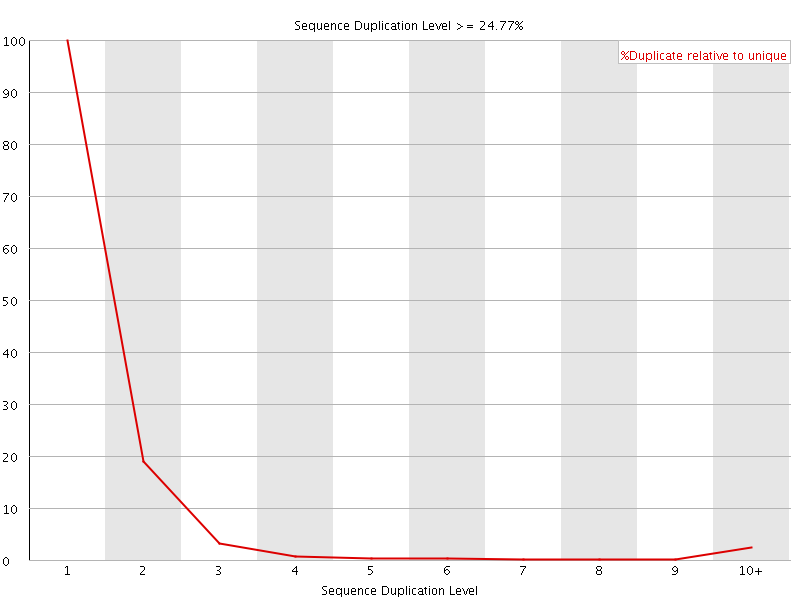
![[FAIL]](Icons/error.png) Overrepresented sequences
Overrepresented sequences
| Sequence | Count | Percentage | Possible Source |
|---|---|---|---|
| GATCGGAAGAGCACACGTCTGAACTCCAGTCACACAGTGATCTCGTATGCC | 272345 | 2.671699710939053 | TruSeq Adapter, Index 5 (100% over 51bp) |
![[WARN]](Icons/warning.png) Kmer Content
Kmer Content
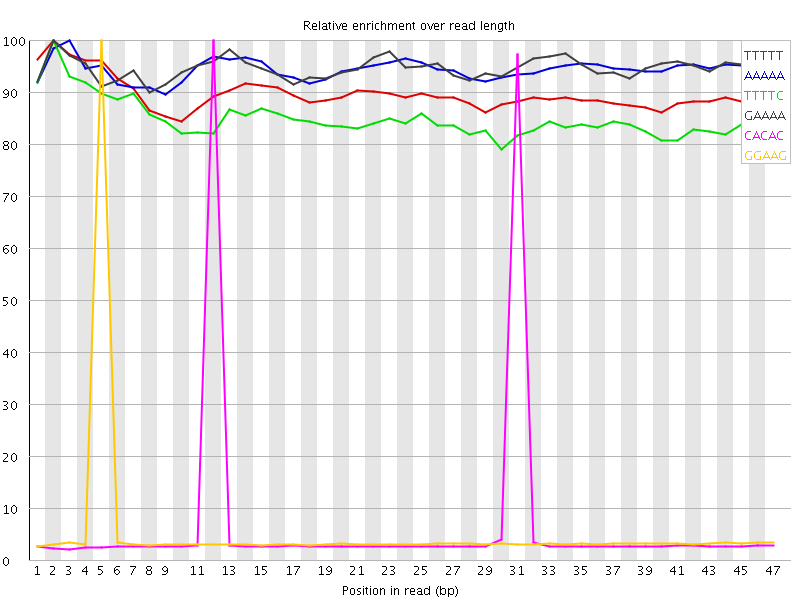
| Sequence | Count | Obs/Exp Overall | Obs/Exp Max | Max Obs/Exp Position |
|---|---|---|---|---|
| TTTTT | 4539640 | 3.8841975 | 4.333546 | 2 |
| AAAAA | 4514465 | 3.8146636 | 4.040089 | 3 |
| TTTTC | 2417320 | 3.0720782 | 3.6177962 | 2 |
| GAAAA | 2322915 | 3.007602 | 3.1773283 | 2 |
| CACAC | 1023415 | 2.855055 | 41.370777 | 12 |
| GGAAG | 806950 | 2.4530573 | 46.027073 | 5 |
| CGGAA | 811635 | 2.3976681 | 44.61785 | 4 |
| GAGCA | 809280 | 2.390711 | 44.306488 | 9 |
| TCCAG | 808325 | 2.3263094 | 42.18255 | 25 |
| CTCCA | 831520 | 2.325527 | 41.059074 | 24 |
| GAAGA | 1162570 | 2.3064451 | 30.617249 | 6 |
| CTGAA | 1150665 | 2.2239559 | 29.34686 | 19 |
| TCTCG | 729110 | 2.1035872 | 42.058266 | 41 |
| TCGGA | 686655 | 2.0335405 | 44.313835 | 3 |
| CAGTG | 669465 | 1.9826323 | 42.587917 | 35 |
| AGAGC | 661450 | 1.9540035 | 43.873768 | 8 |
| GCACA | 680505 | 1.9535602 | 42.640297 | 11 |
| ATCGG | 654675 | 1.9388312 | 44.119873 | 2 |
| AGCAC | 662990 | 1.9032791 | 42.64232 | 10 |
| CACAG | 662205 | 1.9010257 | 41.4759 | 33 |
| CCAGT | 647155 | 1.862472 | 41.637596 | 26 |
| GATCG | 617085 | 1.827508 | 43.810925 | 1 |
| TCTGA | 940075 | 1.8214856 | 29.070969 | 18 |
| GTCTG | 609535 | 1.8096681 | 43.487305 | 17 |
| CGTCT | 617695 | 1.7821387 | 42.199253 | 16 |
| GTCAC | 618745 | 1.7807099 | 41.534164 | 29 |
| CTCGT | 615745 | 1.7765127 | 41.692345 | 42 |
| ATGCC | 606640 | 1.7458724 | 41.613075 | 47 |
| CACGT | 601490 | 1.7310511 | 42.309284 | 14 |
| AAGAG | 863945 | 1.7139972 | 29.993196 | 7 |
| TGAAC | 866825 | 1.675362 | 28.79466 | 20 |
| ACGTC | 576900 | 1.6602826 | 42.120213 | 15 |
| ACTCC | 591600 | 1.6545384 | 40.62506 | 23 |
| ACACG | 574810 | 1.6501364 | 42.22114 | 13 |
| CAGTC | 565910 | 1.628654 | 41.298084 | 27 |
| TCACA | 858865 | 1.61313 | 27.50519 | 30 |
| AGTGA | 803970 | 1.5990052 | 28.912682 | 36 |
| ACACA | 848445 | 1.5895792 | 27.461912 | 32 |
| TGATC | 783275 | 1.5176705 | 28.10063 | 38 |
| GAACT | 775275 | 1.4984182 | 28.57867 | 21 |
| AACTC | 794980 | 1.4931405 | 27.747595 | 22 |
| ATCTC | 790555 | 1.4885468 | 27.470272 | 40 |
| ACAGT | 764360 | 1.4773221 | 28.028334 | 34 |
| GTGAT | 739935 | 1.4753315 | 28.906893 | 37 |
| AGTCA | 704360 | 1.3613567 | 28.003042 | 28 |
| GATCT | 701720 | 1.3596498 | 28.035345 | 39 |
| TCGTA | 593170 | 1.1493238 | 27.993702 | 43 |
| TATGC | 589700 | 1.1426003 | 27.95089 | 46 |
| GTATG | 572350 | 1.1411893 | 28.771114 | 45 |
| CGTAT | 555215 | 1.0757823 | 27.873537 | 44 |