![[OK]](Icons/tick.png) Basic Statistics
Basic Statistics
| Measure | Value |
|---|---|
| Filename | LW185_TGACCA_L002_R1_001.fastq.gz |
| File type | Conventional base calls |
| Encoding | Sanger / Illumina 1.9 |
| Total Sequences | 11701716 |
| Filtered Sequences | 0 |
| Sequence length | 51 |
| %GC | 43 |
![[OK]](Icons/tick.png) Per base sequence quality
Per base sequence quality
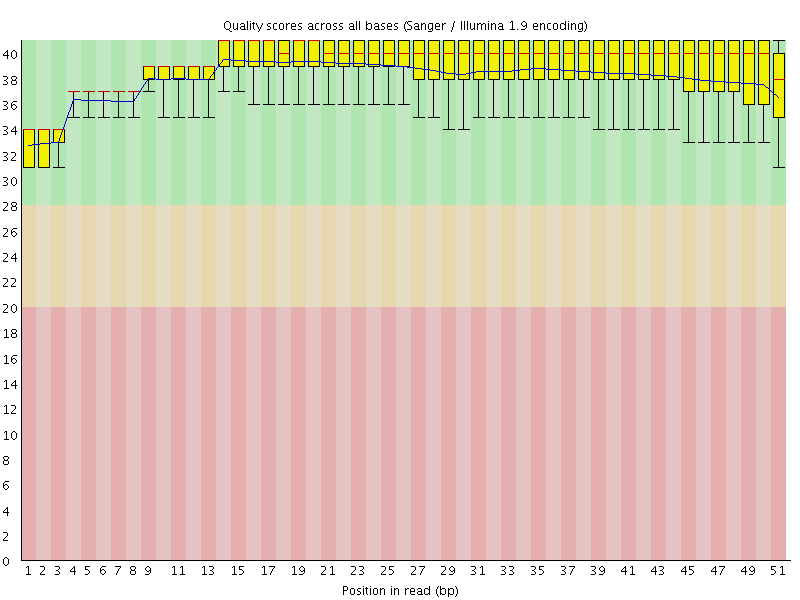
![[OK]](Icons/tick.png) Per sequence quality scores
Per sequence quality scores
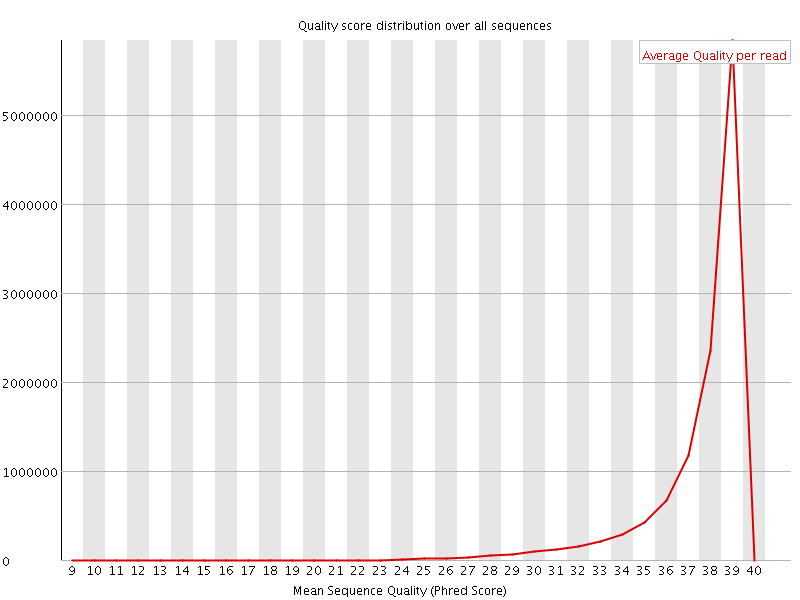
![[WARN]](Icons/warning.png) Per base sequence content
Per base sequence content
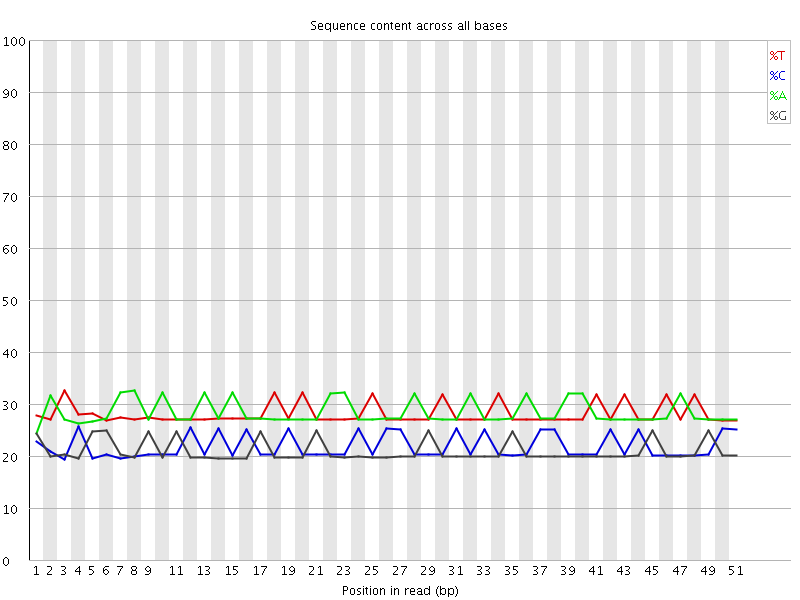
![[OK]](Icons/tick.png) Per base GC content
Per base GC content
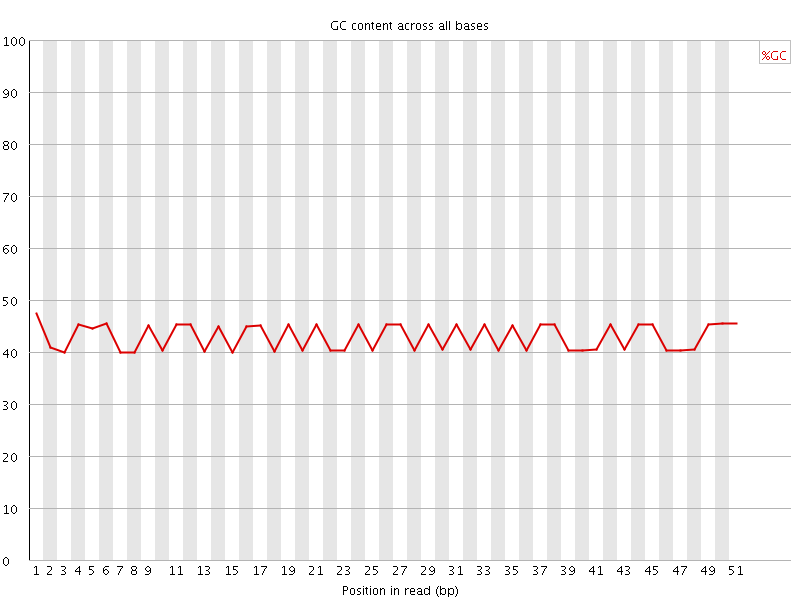
![[FAIL]](Icons/error.png) Per sequence GC content
Per sequence GC content
![[OK]](Icons/tick.png) Per base N content
Per base N content
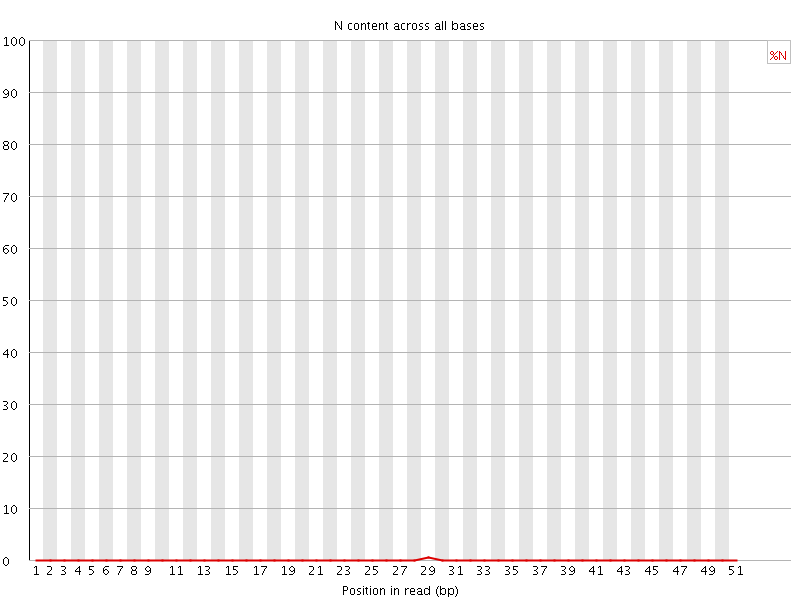
![[OK]](Icons/tick.png) Sequence Length Distribution
Sequence Length Distribution
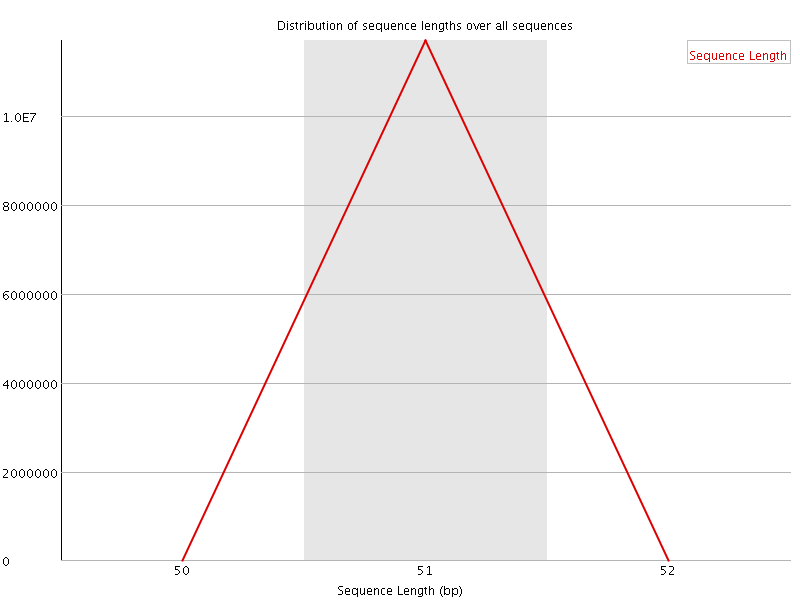
![[WARN]](Icons/warning.png) Sequence Duplication Levels
Sequence Duplication Levels
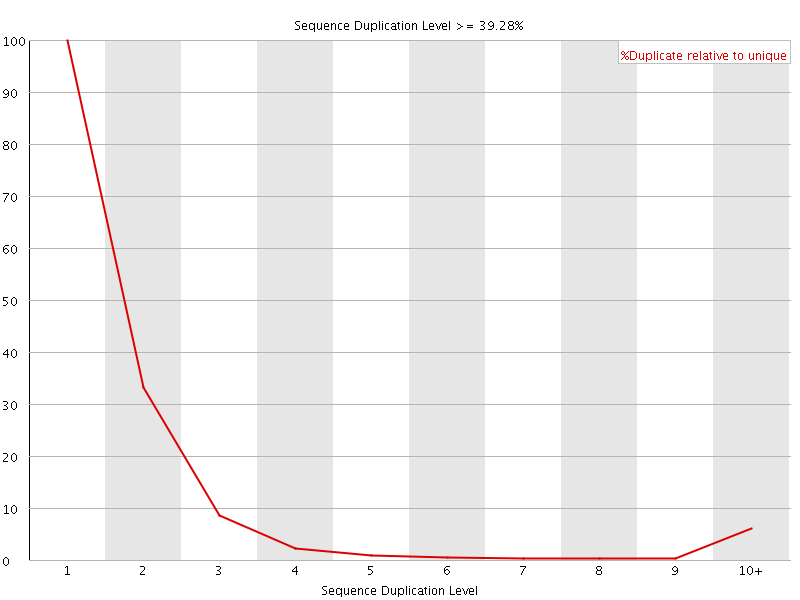
![[FAIL]](Icons/error.png) Overrepresented sequences
Overrepresented sequences
| Sequence | Count | Percentage | Possible Source |
|---|---|---|---|
| GATCGGAAGAGCACACGTCTGAACTCCAGTCACTGACCAATCTCGTATGCC | 515057 | 4.401551020380259 | TruSeq Adapter, Index 4 (100% over 51bp) |
![[WARN]](Icons/warning.png) Kmer Content
Kmer Content
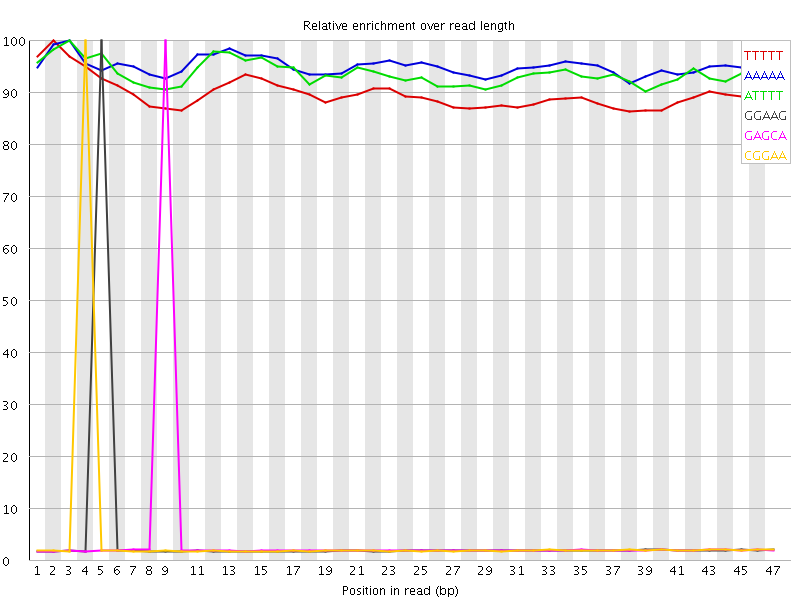
| Sequence | Count | Obs/Exp Overall | Obs/Exp Max | Max Obs/Exp Position |
|---|---|---|---|---|
| TTTTT | 4354955 | 4.378118 | 4.8787317 | 2 |
| AAAAA | 4329750 | 4.1300645 | 4.3429365 | 3 |
| ATTTT | 3034805 | 3.019066 | 3.2243395 | 3 |
| GGAAG | 1157490 | 2.734701 | 67.2688 | 5 |
| GAGCA | 1161825 | 2.6354713 | 64.34748 | 9 |
| CGGAA | 1159220 | 2.6295621 | 64.47049 | 4 |
| GAAGA | 1497275 | 2.614536 | 50.250942 | 6 |
| TCCAG | 1175355 | 2.5868635 | 61.040836 | 25 |
| GATCG | 1116435 | 2.5592513 | 64.76696 | 1 |
| CTGAA | 1463180 | 2.479007 | 47.941856 | 19 |
| TCTCG | 1088665 | 2.421368 | 61.21956 | 41 |
| ATCGG | 1054230 | 2.4166563 | 64.80741 | 2 |
| CTCCA | 1135850 | 2.4002168 | 58.338295 | 24 |
| TCGGA | 1007350 | 2.3091915 | 64.82679 | 3 |
| AGAGC | 987350 | 2.239694 | 64.067924 | 8 |
| CACTG | 1014935 | 2.2337923 | 60.65457 | 31 |
| CCAGT | 1010545 | 2.2241302 | 60.532 | 26 |
| GCACA | 1020365 | 2.2222767 | 61.290928 | 11 |
| AGCAC | 1015320 | 2.211289 | 61.36352 | 10 |
| ATGCC | 1003050 | 2.207634 | 60.233738 | 47 |
| GTCTG | 950340 | 2.201509 | 64.386086 | 17 |
| CTGAC | 982270 | 2.1618989 | 60.228035 | 33 |
| CACAC | 1031915 | 2.1578014 | 58.497074 | 12 |
| GACCA | 980345 | 2.135116 | 58.92142 | 35 |
| CGTCT | 954095 | 2.122062 | 61.596962 | 16 |
| CTCGT | 946845 | 2.1059368 | 60.629433 | 42 |
| TGACC | 949380 | 2.0895107 | 59.605076 | 34 |
| TCTGA | 1219345 | 2.0877023 | 48.046944 | 18 |
| GTCAC | 946575 | 2.083337 | 60.688576 | 29 |
| AAGAG | 1189315 | 2.0767775 | 49.711494 | 7 |
| ACGTC | 933245 | 2.053999 | 61.01377 | 15 |
| TGAAC | 1195605 | 2.0256653 | 47.46556 | 20 |
| CACGT | 919510 | 2.0237694 | 61.258736 | 14 |
| CAGTC | 892035 | 1.9632989 | 60.315018 | 27 |
| ACACG | 895420 | 1.950156 | 60.662075 | 13 |
| ACCAA | 1188080 | 1.912444 | 43.70382 | 36 |
| CCAAT | 1165905 | 1.8965667 | 44.152287 | 37 |
| ACTCC | 894700 | 1.8906317 | 58.11729 | 23 |
| ATCTC | 1143145 | 1.8791795 | 45.24989 | 40 |
| ACTGA | 1107400 | 1.8762232 | 46.879757 | 32 |
| GAACT | 1099415 | 1.8626945 | 47.282185 | 21 |
| CAATC | 1129760 | 1.83777 | 44.149437 | 38 |
| TCACT | 1106160 | 1.8183812 | 45.46879 | 30 |
| AACTC | 1098850 | 1.787489 | 45.30146 | 22 |
| AGTCA | 995005 | 1.6857969 | 46.73383 | 28 |
| AATCT | 1277035 | 1.6160144 | 34.825565 | 39 |
| GTATG | 875830 | 1.5618409 | 48.529034 | 45 |
| TATGC | 902725 | 1.545601 | 46.573563 | 46 |
| TCGTA | 888470 | 1.5211943 | 46.622417 | 43 |
| CGTAT | 870450 | 1.4903413 | 46.56971 | 44 |