![[OK]](Icons/tick.png) Basic Statistics
Basic Statistics
| Measure | Value |
|---|---|
| Filename | LW183_CGATGT_L002_R1_001.fastq.gz |
| File type | Conventional base calls |
| Encoding | Sanger / Illumina 1.9 |
| Total Sequences | 5270968 |
| Filtered Sequences | 0 |
| Sequence length | 51 |
| %GC | 37 |
![[OK]](Icons/tick.png) Per base sequence quality
Per base sequence quality
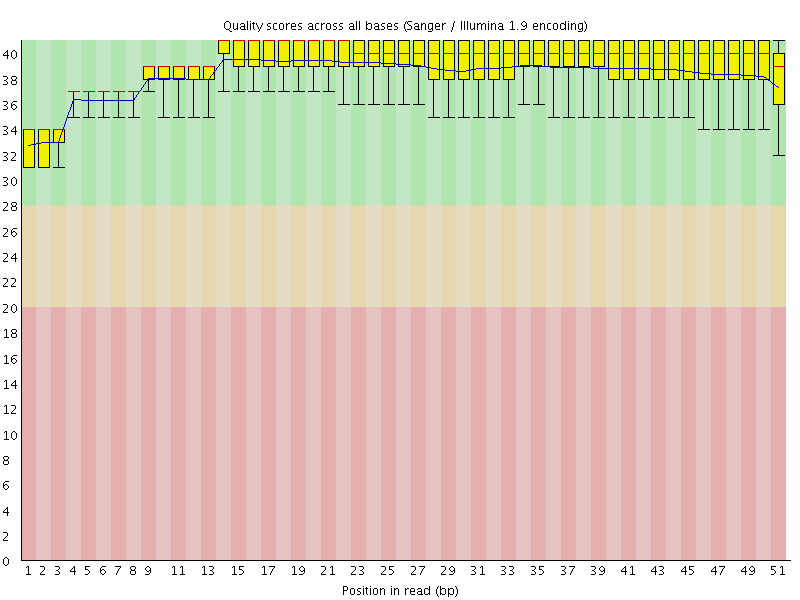
![[OK]](Icons/tick.png) Per sequence quality scores
Per sequence quality scores
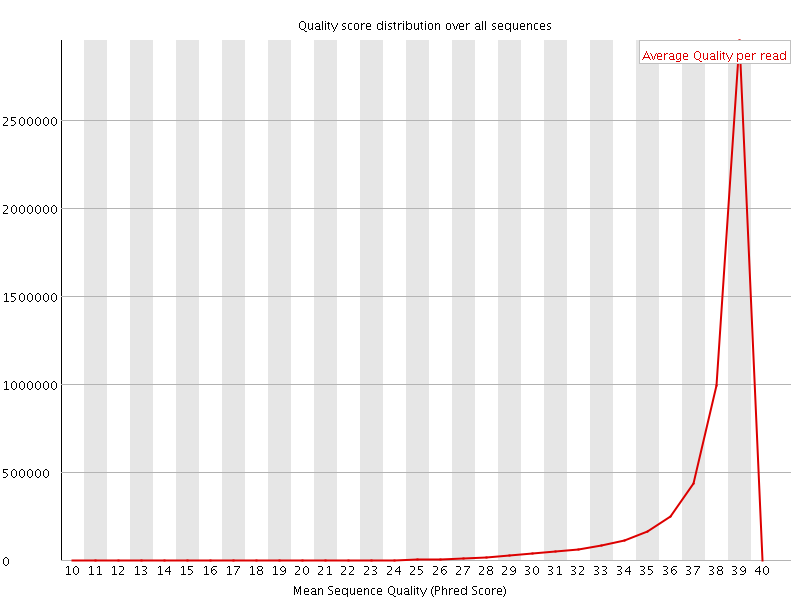
![[WARN]](Icons/warning.png) Per base sequence content
Per base sequence content
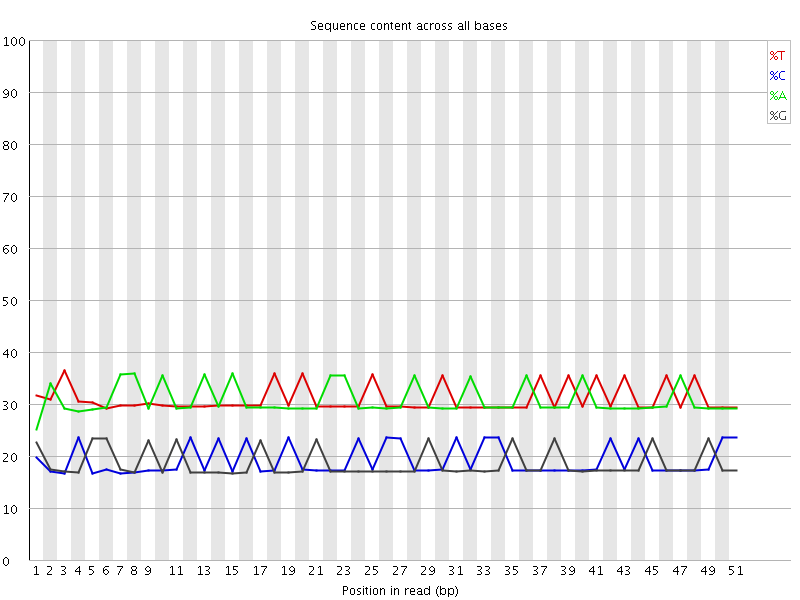
![[OK]](Icons/tick.png) Per base GC content
Per base GC content
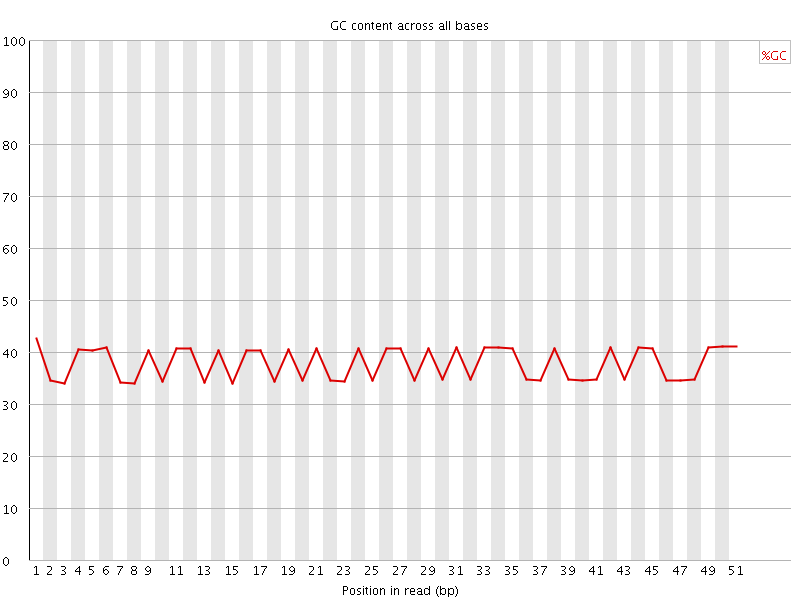
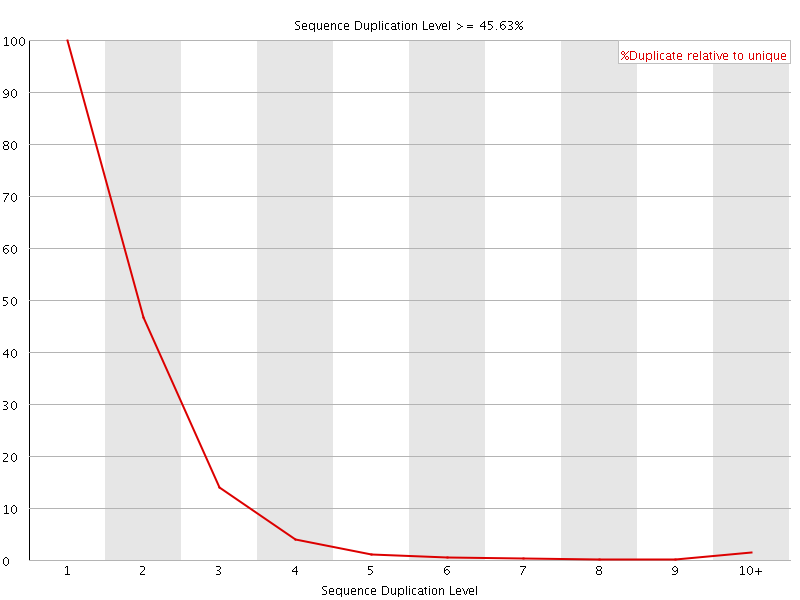
![[FAIL]](Icons/error.png) Per sequence GC content
Per sequence GC content
![[OK]](Icons/tick.png) Per base N content
Per base N content
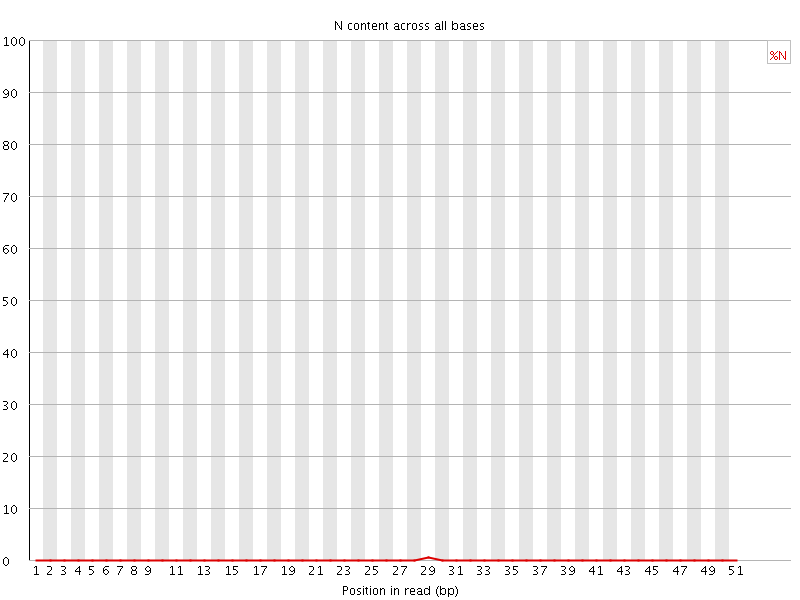
![[OK]](Icons/tick.png) Sequence Length Distribution
Sequence Length Distribution
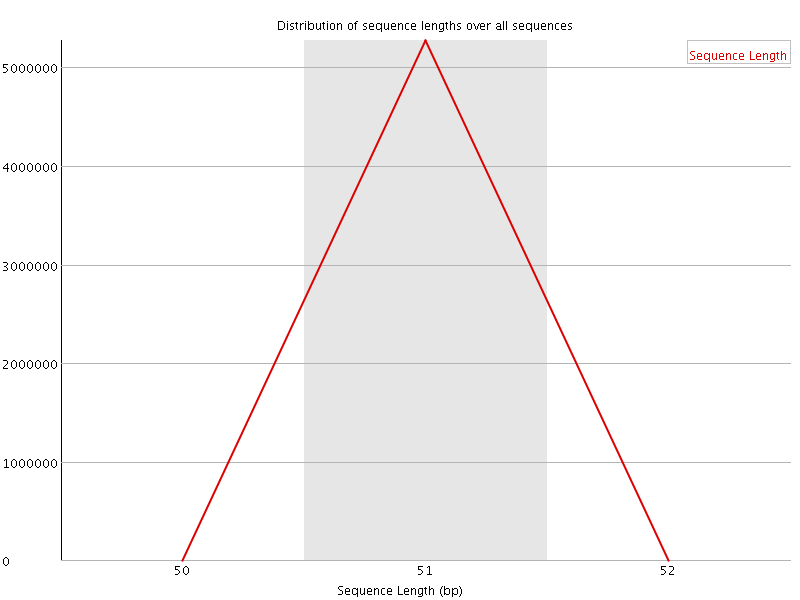
![[WARN]](Icons/warning.png) Sequence Duplication Levels
Sequence Duplication Levels
![[FAIL]](Icons/error.png) Overrepresented sequences
Overrepresented sequences
| Sequence | Count | Percentage | Possible Source |
|---|---|---|---|
| GATCGGAAGAGCACACGTCTGAACTCCAGTCACCGATGTATCTCGTATGCC | 298394 | 5.661085402150041 | TruSeq Adapter, Index 2 (100% over 51bp) |
![[FAIL]](Icons/error.png) Kmer Content
Kmer Content
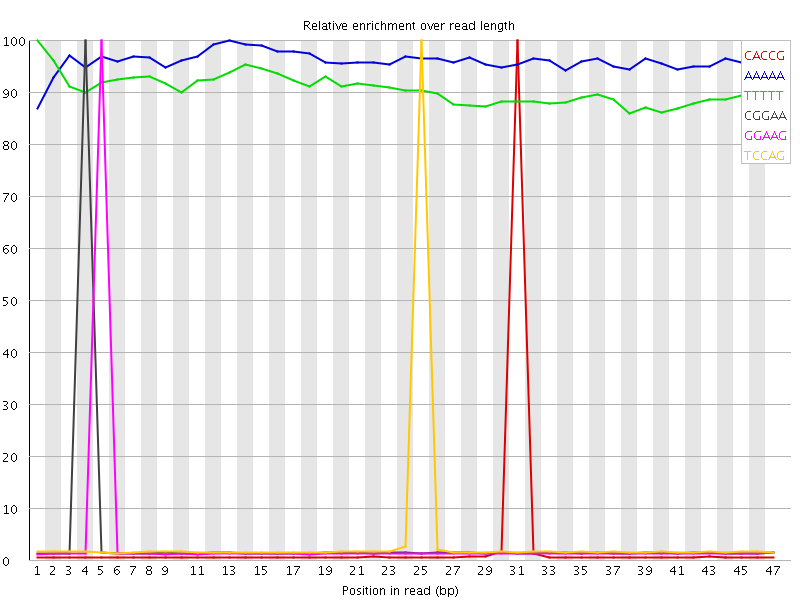
| Sequence | Count | Obs/Exp Overall | Obs/Exp Max | Max Obs/Exp Position |
|---|---|---|---|---|
| CACCG | 437750 | 4.2715435 | 148.14137 | 31 |
| AAAAA | 2780015 | 3.9772005 | 4.139274 | 13 |
| TTTTT | 2858770 | 3.9589434 | 4.371302 | 1 |
| CGGAA | 572505 | 3.5959837 | 98.88306 | 4 |
| GGAAG | 554625 | 3.5954013 | 102.026924 | 5 |
| TCCAG | 589480 | 3.5642815 | 93.1568 | 25 |
| GAGCA | 554705 | 3.4841797 | 98.28294 | 9 |
| CTCCA | 572045 | 3.3513792 | 90.00893 | 24 |
| TCTCG | 534390 | 3.2102225 | 92.00898 | 41 |
| GAAAA | 1355405 | 3.2087703 | 3.4645214 | 2 |
| TCGGA | 507205 | 3.165162 | 97.8696 | 3 |
| TTTTC | 1412875 | 3.1576178 | 3.6917932 | 1 |
| AGAGC | 489320 | 3.0734875 | 97.96002 | 8 |
| TCACC | 521270 | 3.053909 | 89.56901 | 30 |
| GCACA | 500285 | 3.0447133 | 94.70454 | 11 |
| CACAC | 511820 | 3.0181215 | 91.624626 | 12 |
| CGATG | 481515 | 3.0048459 | 94.820244 | 34 |
| ATCGG | 481205 | 3.0029113 | 97.515076 | 2 |
| GTCTG | 469690 | 2.9120421 | 95.59857 | 17 |
| CTGAA | 769660 | 2.9024992 | 59.321648 | 19 |
| CCGAT | 479745 | 2.9007702 | 91.91176 | 33 |
| AGCAC | 476060 | 2.897281 | 94.55452 | 10 |
| CCAGT | 478490 | 2.8931818 | 92.33638 | 26 |
| GAAGA | 733790 | 2.8746204 | 62.212414 | 6 |
| ACCGA | 463830 | 2.8228498 | 92.54333 | 32 |
| CTCGT | 466175 | 2.800437 | 91.33479 | 42 |
| CGTCT | 465470 | 2.7962017 | 92.6134 | 16 |
| GTCAC | 462260 | 2.7950478 | 92.310486 | 29 |
| CACGT | 458930 | 2.7749126 | 93.50831 | 14 |
| GATCG | 442810 | 2.7633114 | 97.15075 | 1 |
| ATGCC | 454595 | 2.748701 | 91.931496 | 47 |
| ACACG | 449835 | 2.7376769 | 94.1302 | 13 |
| ACTCC | 458990 | 2.6890361 | 89.82978 | 23 |
| ACGTC | 442095 | 2.6731203 | 93.24217 | 15 |
| CAGTC | 441990 | 2.6724854 | 91.98318 | 27 |
| TCTGA | 660520 | 2.47476 | 58.471535 | 18 |
| AAGAG | 592030 | 2.319276 | 61.65328 | 7 |
| TGAAC | 596320 | 2.248809 | 58.60191 | 20 |
| GAACT | 571050 | 2.153512 | 58.42748 | 21 |
| AACTC | 569145 | 2.0796325 | 56.552402 | 22 |
| ATCTC | 556145 | 2.0189505 | 55.657295 | 40 |
| GATGT | 520765 | 2.013717 | 58.850807 | 35 |
| AGTCA | 514165 | 1.9389907 | 57.70794 | 28 |
| GTATG | 454205 | 1.7563399 | 58.79101 | 45 |
| TATGC | 466405 | 1.7474723 | 56.903652 | 46 |
| TCGTA | 465540 | 1.7442316 | 56.99725 | 43 |
| GTATC | 451095 | 1.6901107 | 56.69265 | 38 |
| CGTAT | 444830 | 1.6666377 | 56.91826 | 44 |
| TGTAT | 571225 | 1.3261681 | 35.506054 | 37 |
| ATGTA | 554415 | 1.2955446 | 35.66546 | 36 |
| TATCT | 539030 | 1.2125365 | 34.24937 | 39 |