![[OK]](Icons/tick.png) Basic Statistics
Basic Statistics
| Measure | Value |
|---|---|
| Filename | LW182_ATCACG_L002_R1_001.fastq.gz |
| File type | Conventional base calls |
| Encoding | Sanger / Illumina 1.9 |
| Total Sequences | 10329059 |
| Filtered Sequences | 0 |
| Sequence length | 51 |
| %GC | 39 |
![[OK]](Icons/tick.png) Per base sequence quality
Per base sequence quality
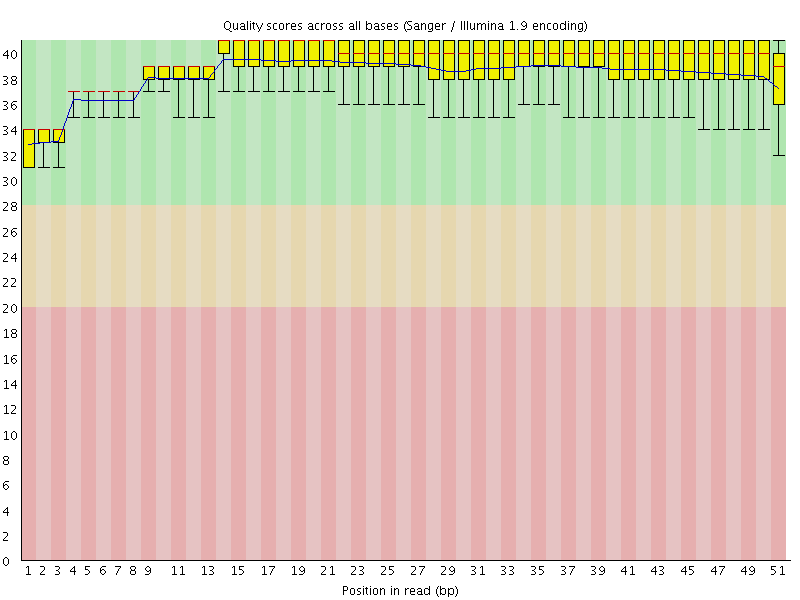
![[OK]](Icons/tick.png) Per sequence quality scores
Per sequence quality scores
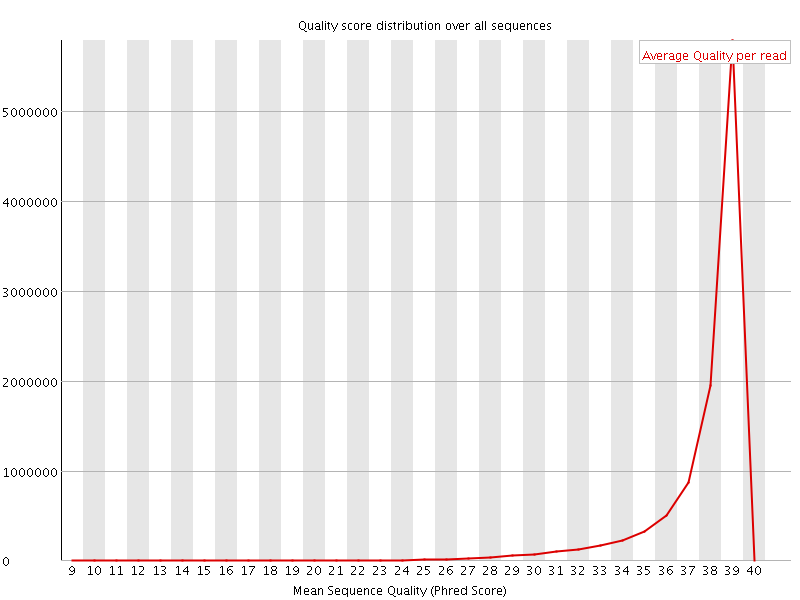
![[WARN]](Icons/warning.png) Per base sequence content
Per base sequence content
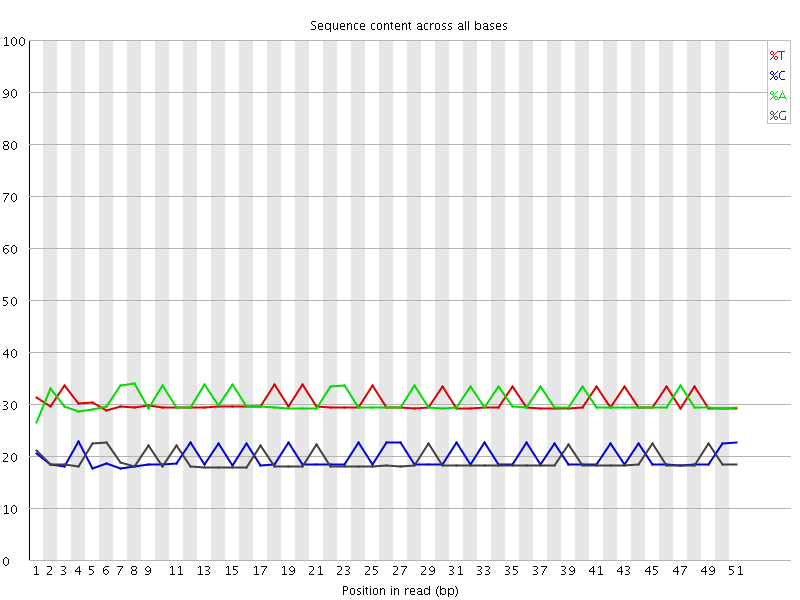
![[OK]](Icons/tick.png) Per base GC content
Per base GC content
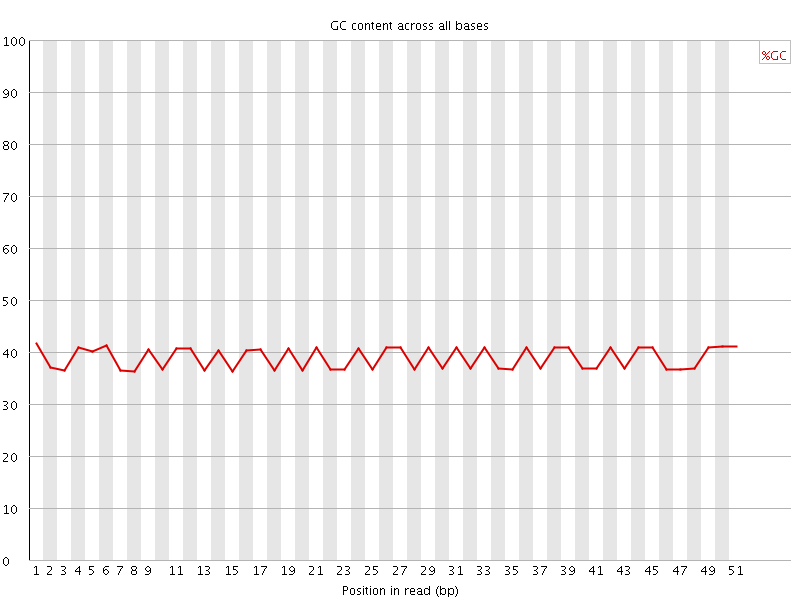
![[OK]](Icons/tick.png) Per sequence GC content
Per sequence GC content
![[OK]](Icons/tick.png) Per base N content
Per base N content
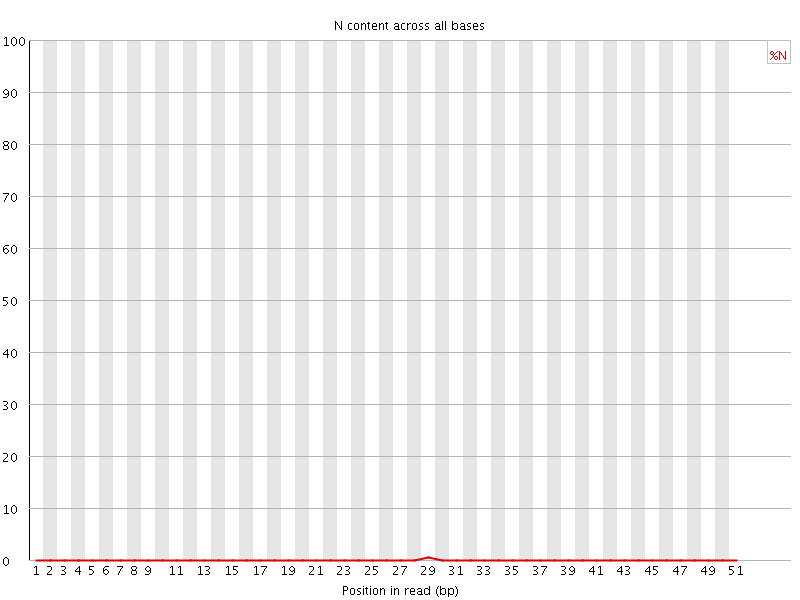
![[OK]](Icons/tick.png) Sequence Length Distribution
Sequence Length Distribution
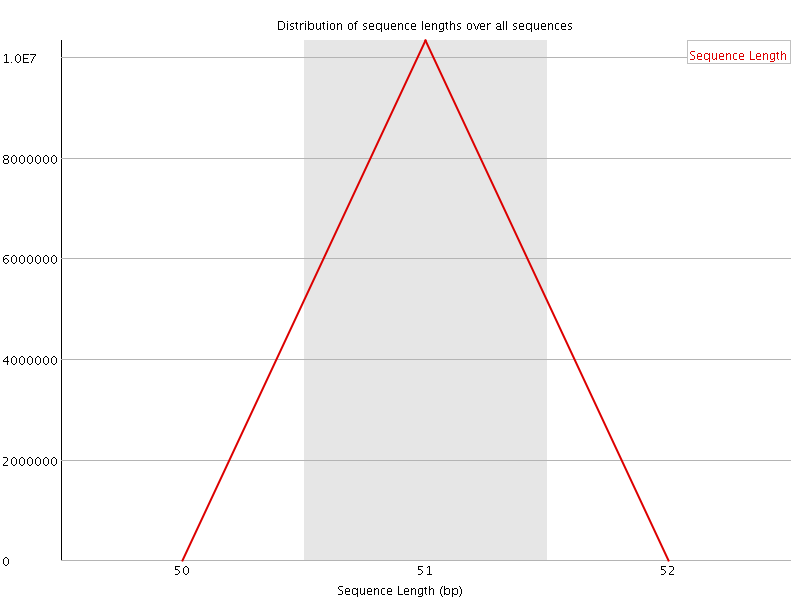
![[WARN]](Icons/warning.png) Sequence Duplication Levels
Sequence Duplication Levels
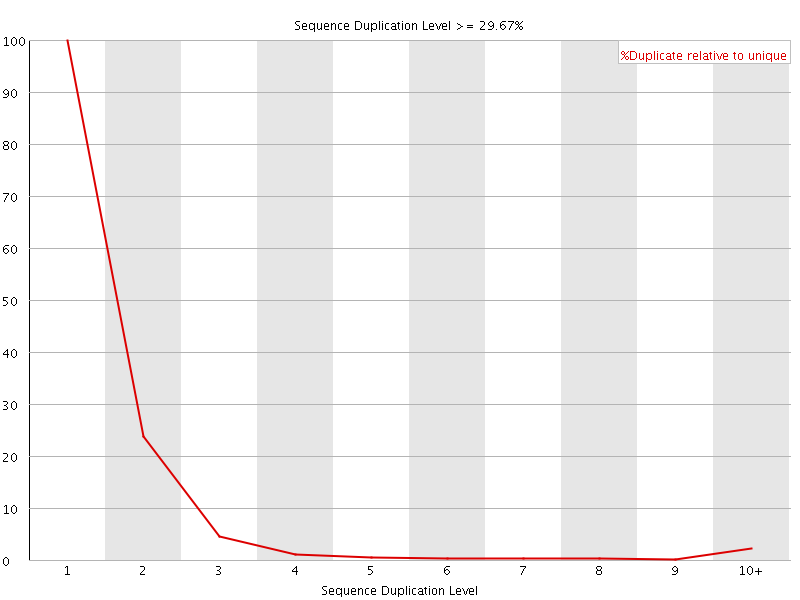
![[FAIL]](Icons/error.png) Overrepresented sequences
Overrepresented sequences
| Sequence | Count | Percentage | Possible Source |
|---|---|---|---|
| GATCGGAAGAGCACACGTCTGAACTCCAGTCACATCACGATCTCGTATGCC | 394187 | 3.816291493736264 | TruSeq Adapter, Index 1 (100% over 51bp) |
![[WARN]](Icons/warning.png) Kmer Content
Kmer Content
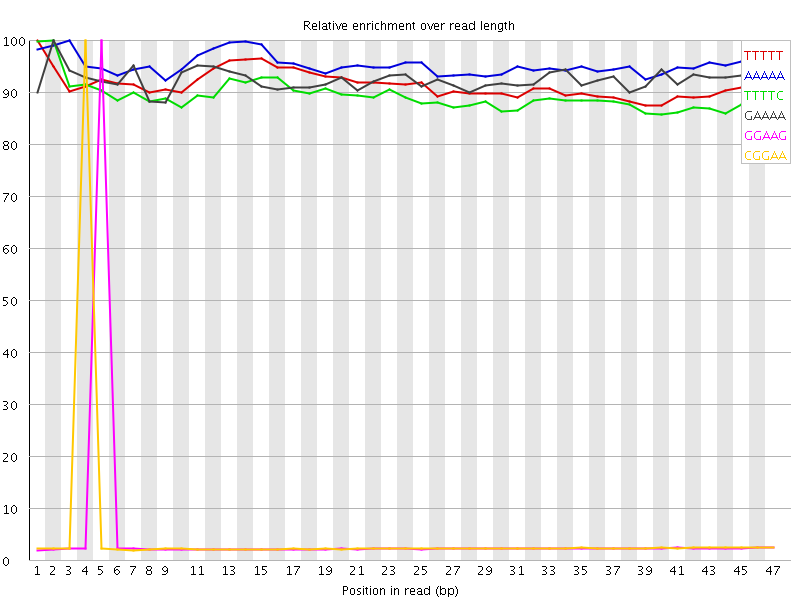
| Sequence | Count | Obs/Exp Overall | Obs/Exp Max | Max Obs/Exp Position |
|---|---|---|---|---|
| TTTTT | 4909240 | 3.9012132 | 4.262089 | 1 |
| AAAAA | 4907455 | 3.784567 | 3.9694939 | 3 |
| TTTTC | 2504095 | 3.0471182 | 3.4120917 | 2 |
| GAAAA | 2442435 | 3.0041175 | 3.2499301 | 2 |
| GGAAG | 930930 | 2.9125872 | 65.70619 | 5 |
| CGGAA | 936170 | 2.8290513 | 63.458767 | 4 |
| GAGCA | 929085 | 2.8076406 | 63.270065 | 9 |
| CTCCA | 954380 | 2.706829 | 58.329662 | 24 |
| TCCAG | 910625 | 2.67396 | 60.284466 | 25 |
| GAAGA | 1270905 | 2.4931076 | 41.78845 | 6 |
| TCGGA | 813220 | 2.4722893 | 63.445213 | 3 |
| TCTCG | 830710 | 2.4539738 | 60.146343 | 41 |
| CTGAA | 1284240 | 2.4479547 | 40.214348 | 19 |
| AGAGC | 783640 | 2.3681145 | 62.905445 | 8 |
| GCACA | 803980 | 2.3466887 | 60.6599 | 11 |
| ATCGG | 765710 | 2.3278527 | 63.109077 | 2 |
| CACAC | 825290 | 2.3267033 | 58.503616 | 12 |
| AGCAC | 779735 | 2.2759216 | 60.660587 | 10 |
| CCAGT | 754170 | 2.2145455 | 59.646564 | 26 |
| GTCTG | 714485 | 2.1851912 | 62.732986 | 17 |
| CACGA | 739595 | 2.1587594 | 58.58095 | 36 |
| GTCAC | 732740 | 2.1516182 | 59.53131 | 29 |
| GATCG | 705510 | 2.1448371 | 62.80938 | 1 |
| CTCGT | 725615 | 2.143516 | 59.649963 | 42 |
| CGTCT | 721725 | 2.1320248 | 60.58786 | 16 |
| CACGT | 721200 | 2.1177323 | 60.40724 | 14 |
| ATGCC | 712210 | 2.091334 | 59.24627 | 47 |
| TCTGA | 1074380 | 2.0602512 | 40.047943 | 18 |
| TCACG | 697680 | 2.0486681 | 58.95838 | 35 |
| ACACG | 696770 | 2.0337598 | 60.130936 | 13 |
| ACTCC | 714675 | 2.0269735 | 57.91584 | 23 |
| CGATC | 687365 | 2.0183792 | 58.821728 | 38 |
| ACGTC | 686450 | 2.0156922 | 60.235157 | 15 |
| CAGTC | 678170 | 1.9913789 | 59.32793 | 27 |
| AAGAG | 976480 | 1.9155402 | 41.235855 | 7 |
| CATCA | 1028405 | 1.8934137 | 37.810307 | 33 |
| TGAAC | 987935 | 1.8831528 | 39.6218 | 20 |
| TCACA | 989755 | 1.8222545 | 37.78985 | 30 |
| GAACT | 902190 | 1.71971 | 39.33896 | 21 |
| AACTC | 924745 | 1.7025635 | 38.034824 | 22 |
| CACAT | 911315 | 1.6778374 | 37.647266 | 31 |
| ATCTC | 894020 | 1.6558986 | 37.715374 | 40 |
| AGTCA | 831270 | 1.5845257 | 38.791805 | 28 |
| ATCAC | 851125 | 1.5670205 | 37.34142 | 34 |
| ACATC | 846675 | 1.5588275 | 37.4765 | 32 |
| GATCT | 811090 | 1.5553613 | 38.7652 | 39 |
| ACGAT | 797600 | 1.5203457 | 38.438923 | 37 |
| GTATG | 693495 | 1.3768336 | 40.02827 | 45 |
| TCGTA | 715980 | 1.3729768 | 38.694447 | 43 |
| TATGC | 715775 | 1.3725837 | 38.682713 | 46 |
| CGTAT | 678505 | 1.3011141 | 38.62583 | 44 |