![[OK]](Icons/tick.png) Basic Statistics
Basic Statistics
| Measure | Value |
|---|---|
| Filename | LW180_GAGTGG_L002_R1_001.fastq.gz |
| File type | Conventional base calls |
| Encoding | Sanger / Illumina 1.9 |
| Total Sequences | 9174985 |
| Filtered Sequences | 0 |
| Sequence length | 51 |
| %GC | 37 |
![[OK]](Icons/tick.png) Per base sequence quality
Per base sequence quality
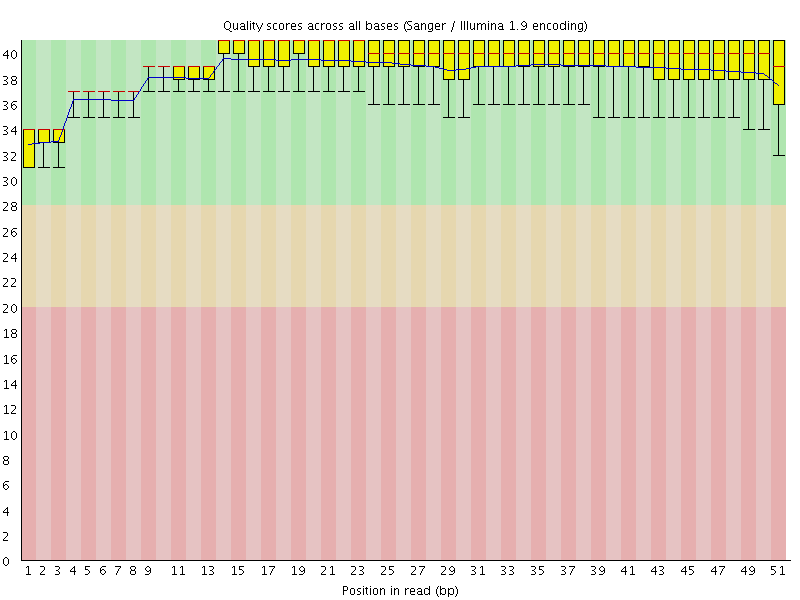
![[OK]](Icons/tick.png) Per sequence quality scores
Per sequence quality scores
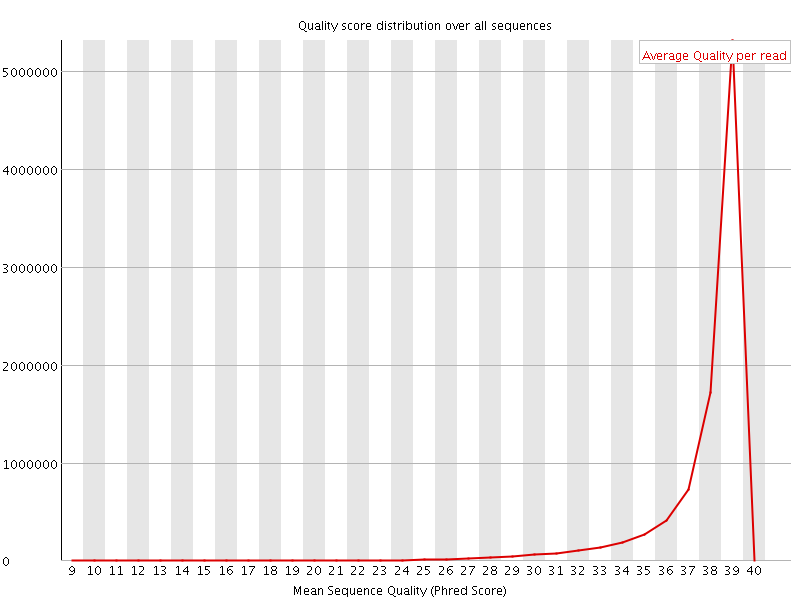
![[WARN]](Icons/warning.png) Per base sequence content
Per base sequence content
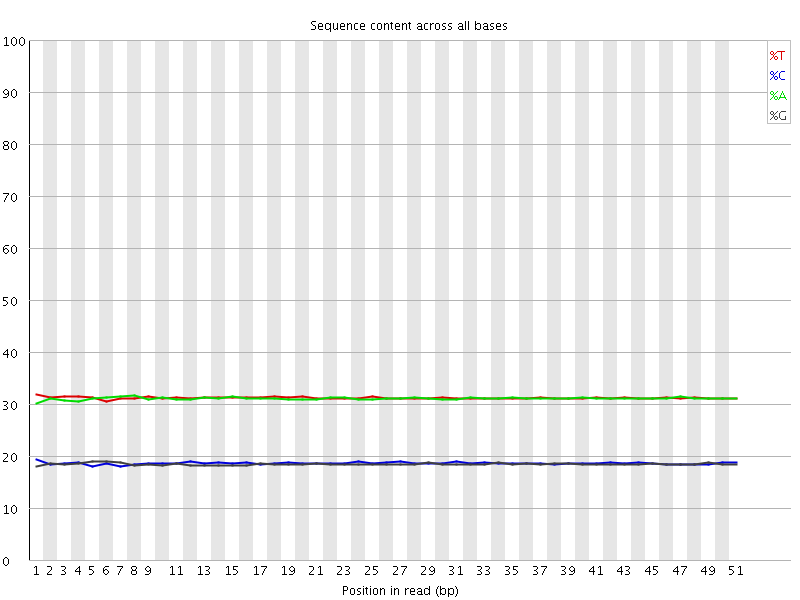
![[OK]](Icons/tick.png) Per base GC content
Per base GC content
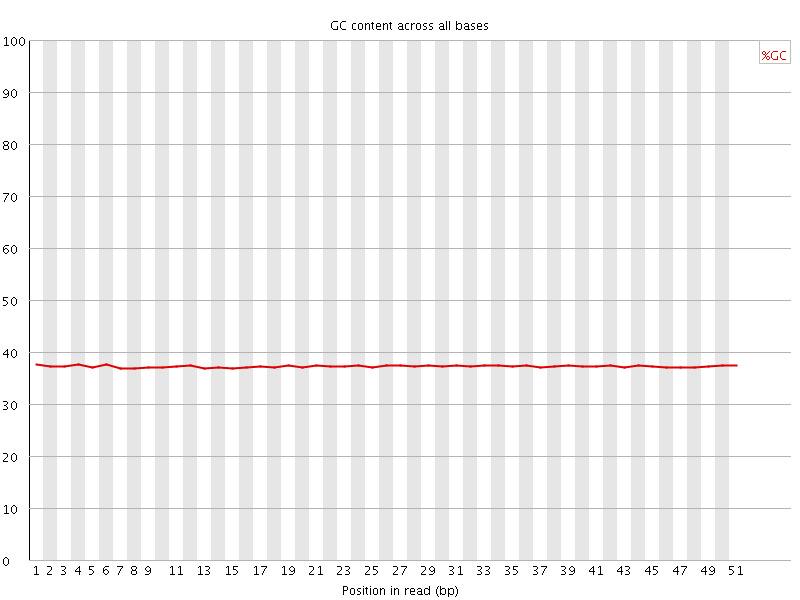
![[OK]](Icons/tick.png) Per sequence GC content
Per sequence GC content
![[OK]](Icons/tick.png) Per base N content
Per base N content
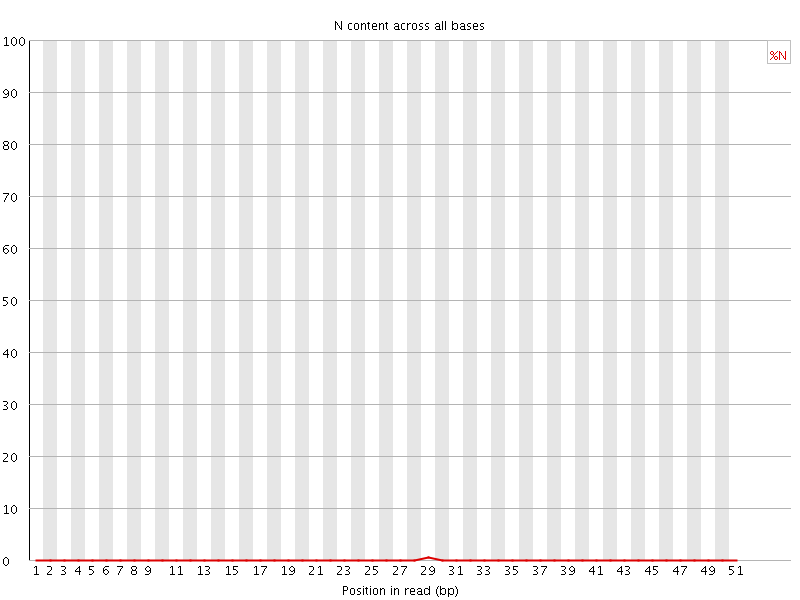
![[OK]](Icons/tick.png) Sequence Length Distribution
Sequence Length Distribution
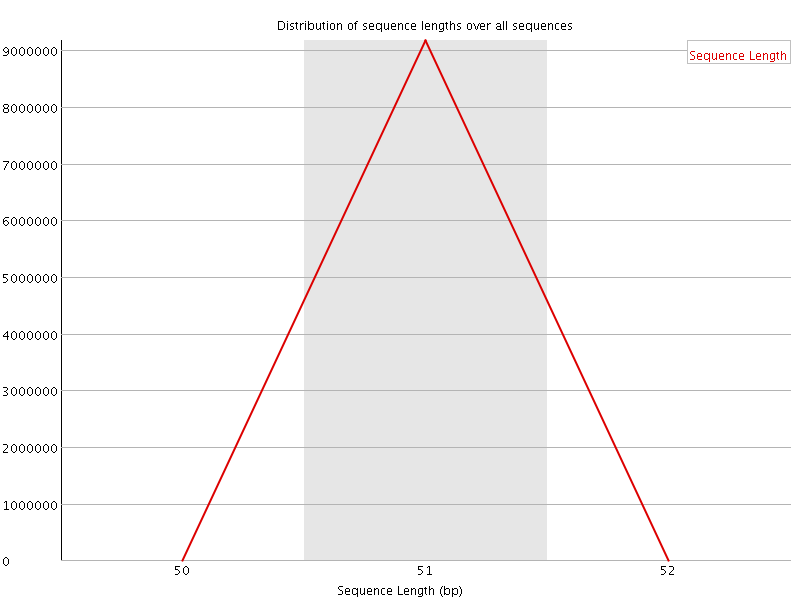
![[OK]](Icons/tick.png) Sequence Duplication Levels
Sequence Duplication Levels
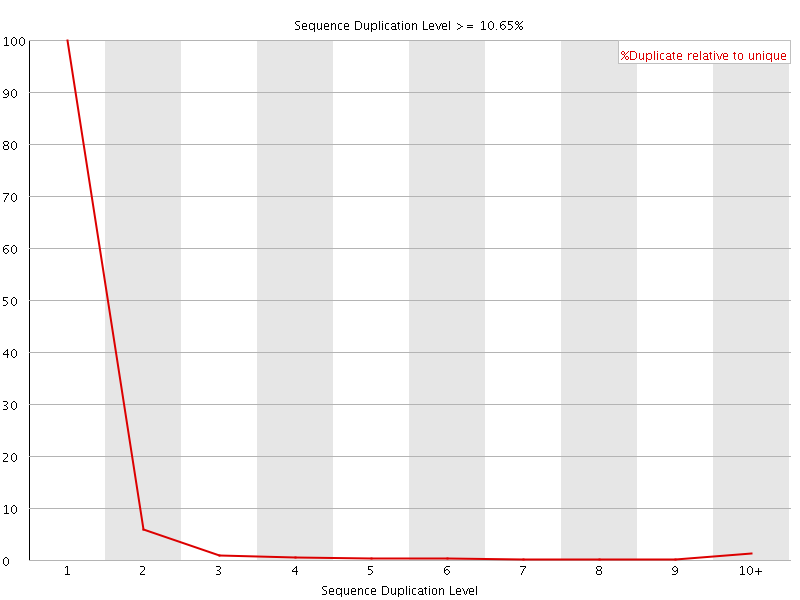
![[WARN]](Icons/warning.png) Overrepresented sequences
Overrepresented sequences
| Sequence | Count | Percentage | Possible Source |
|---|---|---|---|
| GATCGGAAGAGCACACGTCTGAACTCCAGTCACGAGTGGATCTCGTATGCC | 20623 | 0.22477420944012444 | TruSeq Adapter, Index 7 (97% over 36bp) |
![[WARN]](Icons/warning.png) Kmer Content
Kmer Content
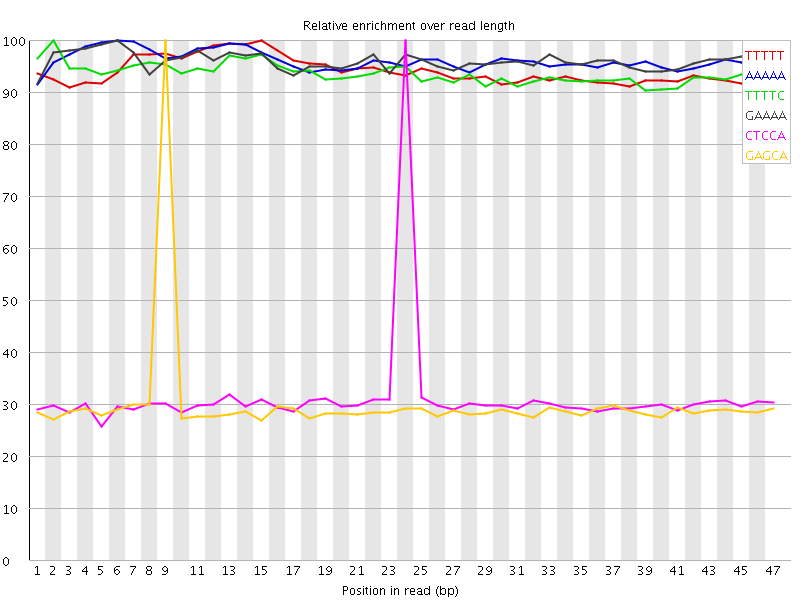
| Sequence | Count | Obs/Exp Overall | Obs/Exp Max | Max Obs/Exp Position |
|---|---|---|---|---|
| TTTTT | 4678900 | 3.5945184 | 3.8157516 | 15 |
| AAAAA | 4612590 | 3.5940979 | 3.7343867 | 6 |
| TTTTC | 2392380 | 3.0661678 | 3.2726314 | 2 |
| GAAAA | 2333720 | 3.0499566 | 3.175481 | 6 |
| CTCCA | 512405 | 1.8329314 | 5.839377 | 24 |
| GAGCA | 486130 | 1.772704 | 5.8831844 | 9 |
| CGGAA | 483690 | 1.7638067 | 5.9516644 | 4 |
| GGAAG | 472520 | 1.7372595 | 5.9091787 | 5 |
| TCCAG | 476605 | 1.7189059 | 5.7029533 | 25 |
| GTGGA | 408825 | 1.49883 | 5.0890155 | 36 |
| TCTCG | 392205 | 1.4105128 | 5.1047626 | 41 |
| TCGGA | 386775 | 1.4064121 | 5.584874 | 3 |
| CACAC | 358935 | 1.2875922 | 5.3799434 | 12 |
| GCACA | 352565 | 1.2751533 | 5.3732934 | 11 |
| AGAGC | 342095 | 1.2474715 | 5.347339 | 8 |
| AGCAC | 325365 | 1.1767765 | 5.272263 | 10 |
| ATCGG | 315585 | 1.1475471 | 5.2340593 | 2 |
| CCAGT | 317835 | 1.1462921 | 5.054038 | 26 |
| CACGT | 285385 | 1.0292591 | 5.0617933 | 14 |
| GTCTG | 280445 | 1.0168861 | 5.054088 | 17 |
| CGTCT | 282715 | 1.0167468 | 5.055146 | 16 |
| ACACG | 264640 | 0.95714694 | 5.0319967 | 13 |
| GATCG | 253775 | 0.9227904 | 5.0075912 | 1 |