![[OK]](Icons/tick.png) Basic Statistics
Basic Statistics
| Measure | Value |
|---|---|
| Filename | LW178_GTTTCG_L002_R1_001.fastq.gz |
| File type | Conventional base calls |
| Encoding | Sanger / Illumina 1.9 |
| Total Sequences | 6143335 |
| Filtered Sequences | 0 |
| Sequence length | 51 |
| %GC | 39 |
![[OK]](Icons/tick.png) Per base sequence quality
Per base sequence quality
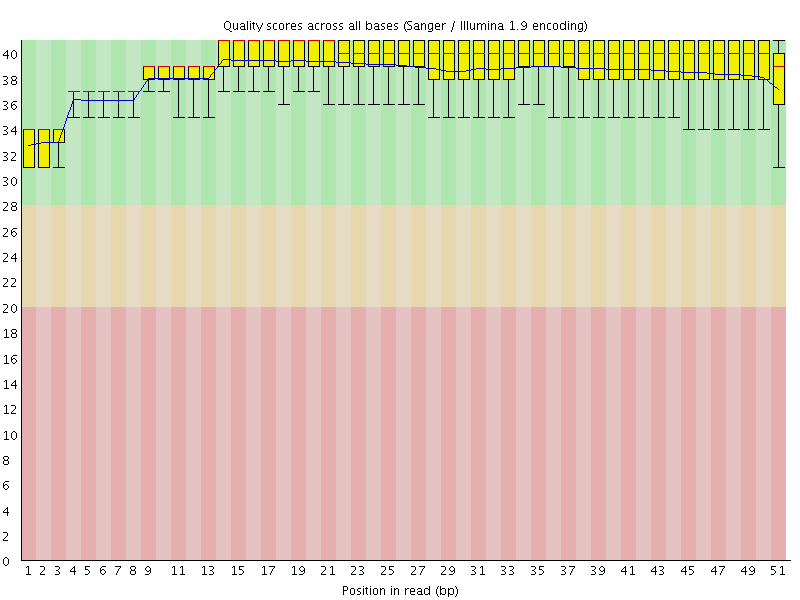
![[OK]](Icons/tick.png) Per sequence quality scores
Per sequence quality scores
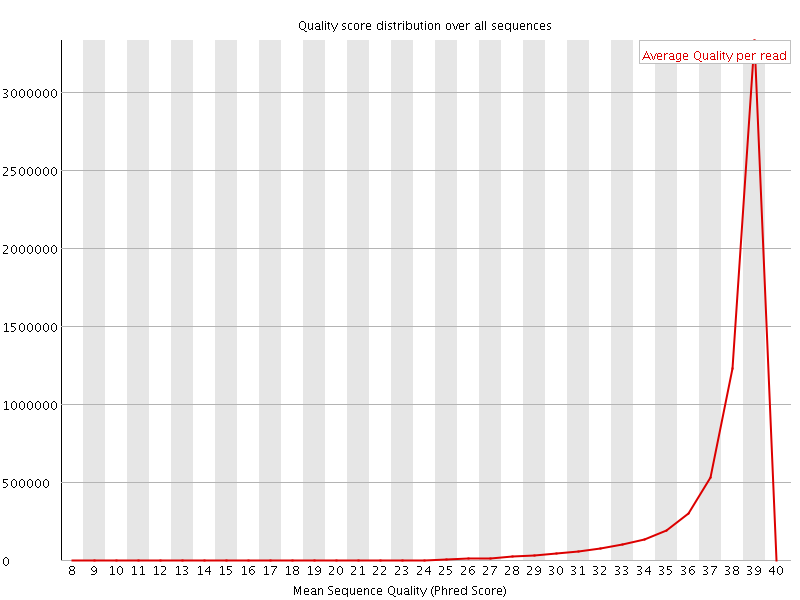
![[WARN]](Icons/warning.png) Per base sequence content
Per base sequence content
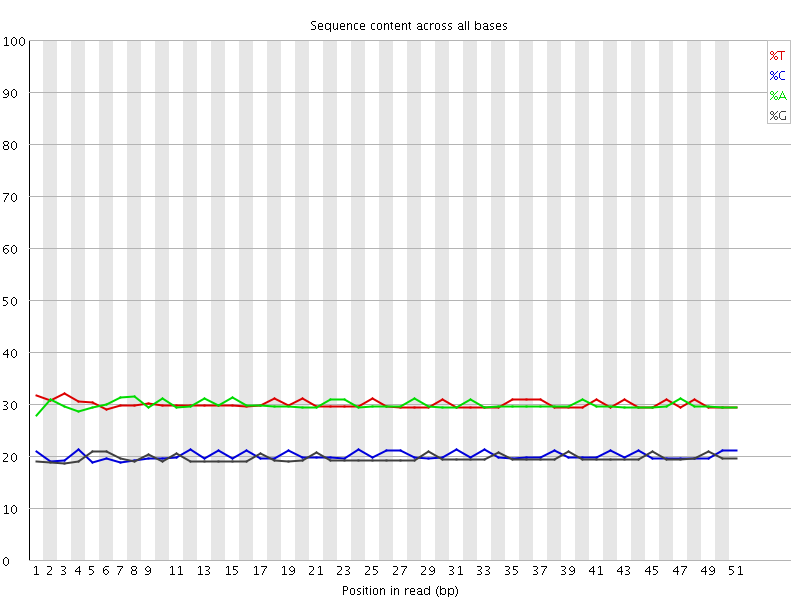
![[OK]](Icons/tick.png) Per base GC content
Per base GC content
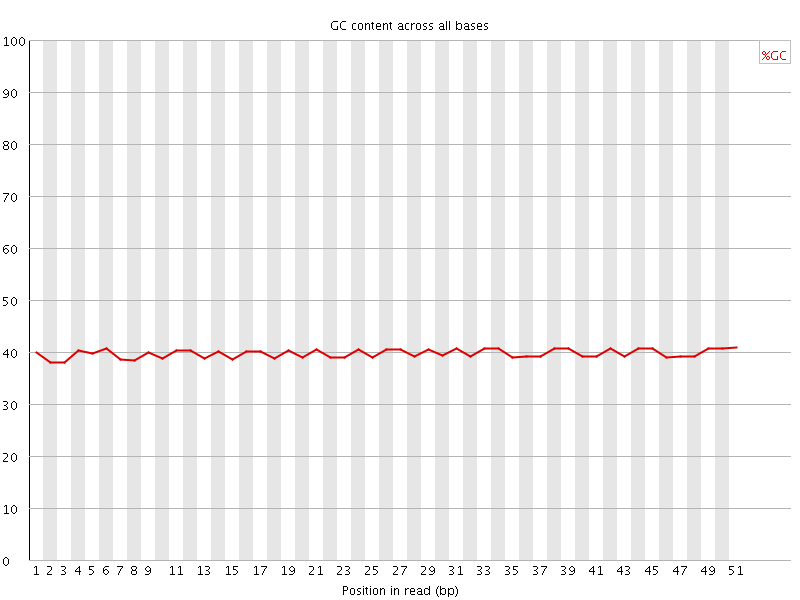
![[OK]](Icons/tick.png) Per sequence GC content
Per sequence GC content
![[OK]](Icons/tick.png) Per base N content
Per base N content
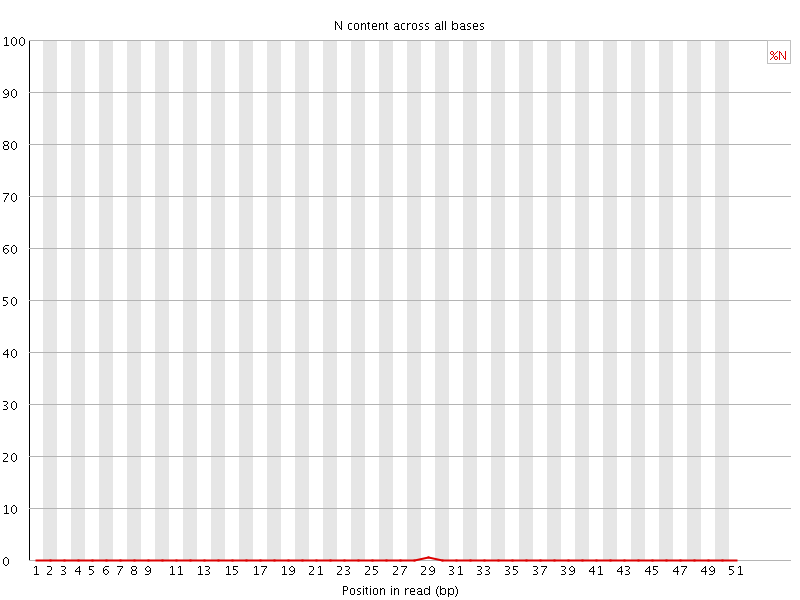
![[OK]](Icons/tick.png) Sequence Length Distribution
Sequence Length Distribution
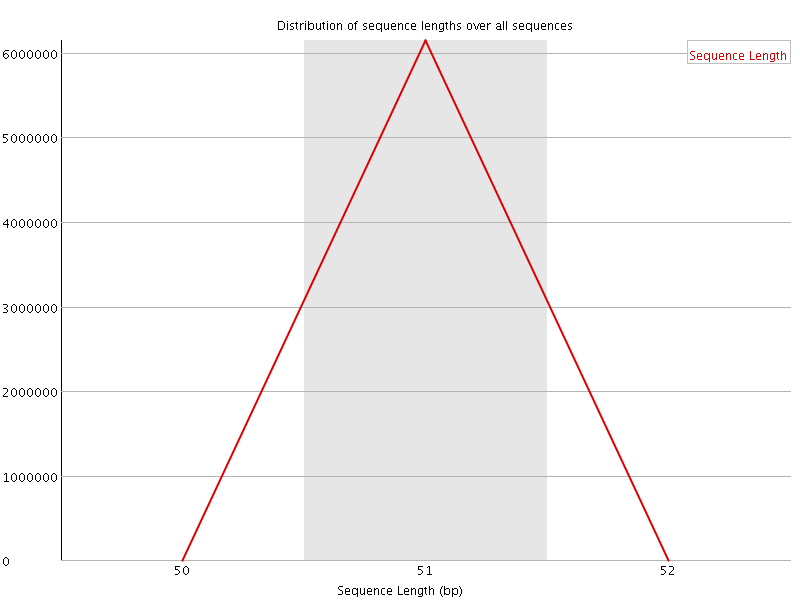
![[OK]](Icons/tick.png) Sequence Duplication Levels
Sequence Duplication Levels
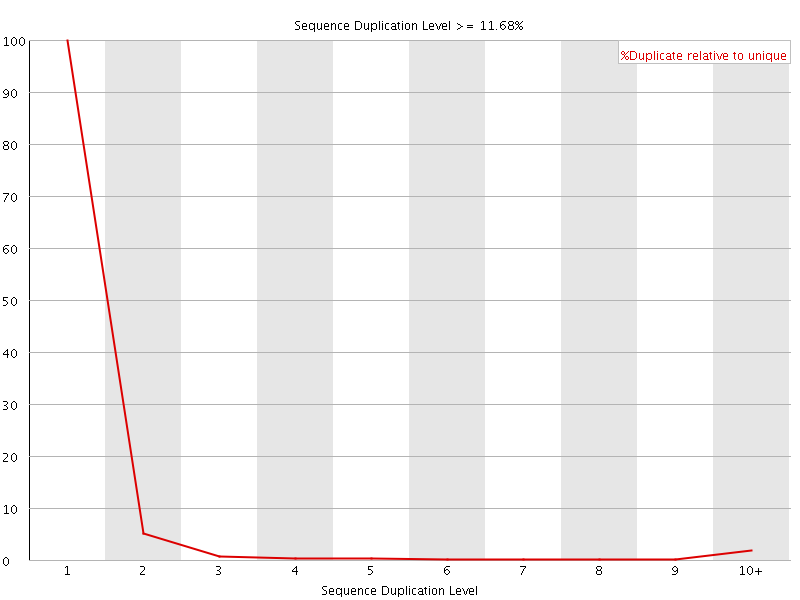
![[FAIL]](Icons/error.png) Overrepresented sequences
Overrepresented sequences
| Sequence | Count | Percentage | Possible Source |
|---|---|---|---|
| GATCGGAAGAGCACACGTCTGAACTCCAGTCACGTTTCGATCTCGTATGCC | 83627 | 1.3612638737754006 | TruSeq Adapter, Index 9 (97% over 36bp) |
![[WARN]](Icons/warning.png) Kmer Content
Kmer Content
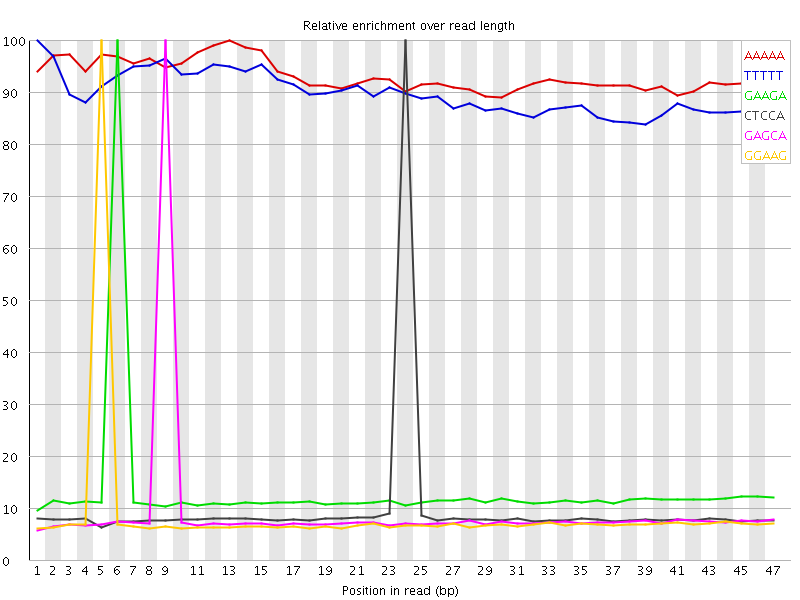
| Sequence | Count | Obs/Exp Overall | Obs/Exp Max | Max Obs/Exp Position |
|---|---|---|---|---|
| AAAAA | 2351005 | 3.3687503 | 3.6155396 | 13 |
| TTTTT | 2350115 | 3.2763827 | 3.648945 | 1 |
| GAAGA | 658080 | 2.1875746 | 16.493671 | 6 |
| CTCCA | 464395 | 2.160263 | 21.786844 | 24 |
| GAGCA | 437460 | 2.1571705 | 23.30012 | 9 |
| GGAAG | 418480 | 2.1188166 | 24.084276 | 5 |
| TCCAG | 416680 | 1.9901888 | 22.17462 | 25 |
| CTGAA | 609060 | 1.9610549 | 15.647946 | 19 |
| CGGAA | 396505 | 1.9552159 | 23.275806 | 4 |
| CACGT | 383445 | 1.8314484 | 21.720924 | 14 |
| TTTCG | 571675 | 1.8206019 | 14.713645 | 35 |
| TCGGA | 349840 | 1.7156695 | 23.037985 | 3 |
| TCTCG | 358355 | 1.7022493 | 21.328506 | 41 |
| GTTTC | 525670 | 1.6740906 | 14.593053 | 34 |
| AGAGC | 334075 | 1.647366 | 22.860254 | 8 |
| TTCGA | 510940 | 1.6361295 | 14.582369 | 36 |
| CACAC | 347070 | 1.6233723 | 21.67724 | 12 |
| AGCAC | 335480 | 1.6111659 | 22.232706 | 10 |
| GCACA | 331015 | 1.5897224 | 22.211287 | 11 |
| TCTGA | 496235 | 1.5890412 | 15.143603 | 18 |
| AAGAG | 452990 | 1.5058191 | 15.777335 | 7 |
| ATCGG | 300790 | 1.4751208 | 22.694925 | 2 |
| CCAGT | 303810 | 1.4510878 | 21.502731 | 26 |
| TGAAC | 439810 | 1.4161028 | 15.088416 | 20 |
| CTCGT | 296225 | 1.407121 | 20.928497 | 42 |
| CGTCT | 290330 | 1.3791186 | 21.514362 | 16 |
| GTCAC | 288325 | 1.3771268 | 21.25543 | 29 |
| CGTTT | 428075 | 1.3632818 | 14.392315 | 33 |
| TCGAT | 421170 | 1.3486685 | 14.323193 | 37 |
| ACTCC | 289780 | 1.3479929 | 21.011105 | 23 |
| GTCTG | 274910 | 1.340827 | 21.961676 | 17 |
| GATCG | 268720 | 1.3178445 | 22.383629 | 1 |
| ATGCC | 272465 | 1.3013747 | 20.980589 | 47 |
| AACTC | 413940 | 1.2980593 | 14.609215 | 22 |
| ACACG | 269420 | 1.2939082 | 21.859552 | 13 |
| ACGTC | 268775 | 1.28375 | 21.586714 | 15 |
| TCACG | 267390 | 1.2771349 | 21.044792 | 30 |
| CAGTC | 263320 | 1.2576954 | 21.288177 | 27 |
| ATCTC | 403055 | 1.2570124 | 14.058207 | 40 |
| GAACT | 389645 | 1.2545812 | 14.938742 | 21 |
| CGATC | 262055 | 1.2516534 | 20.596292 | 38 |
| GATCT | 357920 | 1.1461296 | 14.239628 | 39 |
| AGTCA | 346610 | 1.116017 | 14.560917 | 28 |
| ACGTT | 315295 | 1.009636 | 14.1872 | 32 |
| TCGTA | 287730 | 0.9213675 | 14.118862 | 43 |
| TATGC | 272640 | 0.8730464 | 14.033552 | 46 |
| GTATG | 257560 | 0.8468348 | 14.381962 | 45 |
| CGTAT | 254835 | 0.8160312 | 14.003197 | 44 |