![[OK]](Icons/tick.png) Basic Statistics
Basic Statistics
| Measure | Value |
|---|---|
| Filename | LW177_GTGGCC_L002_R1_001.fastq.gz |
| File type | Conventional base calls |
| Encoding | Sanger / Illumina 1.9 |
| Total Sequences | 5187616 |
| Filtered Sequences | 0 |
| Sequence length | 51 |
| %GC | 39 |
![[OK]](Icons/tick.png) Per base sequence quality
Per base sequence quality
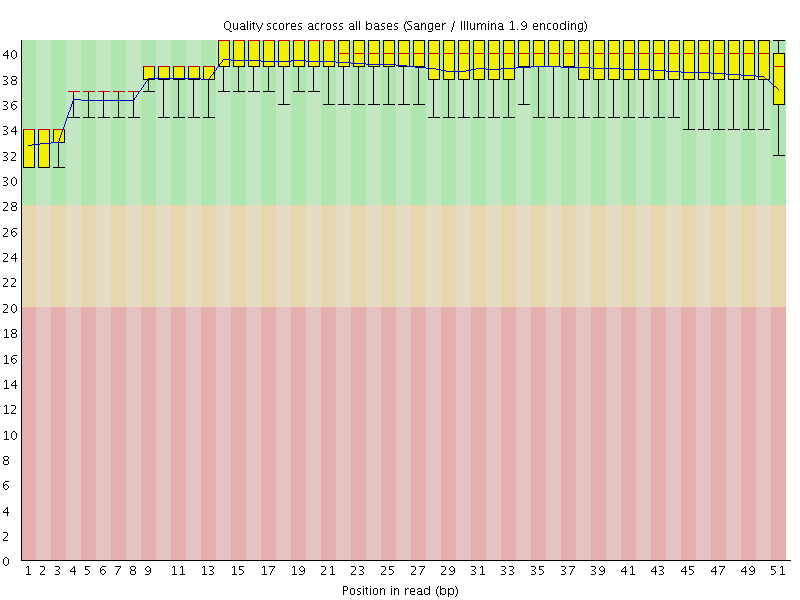
![[OK]](Icons/tick.png) Per sequence quality scores
Per sequence quality scores
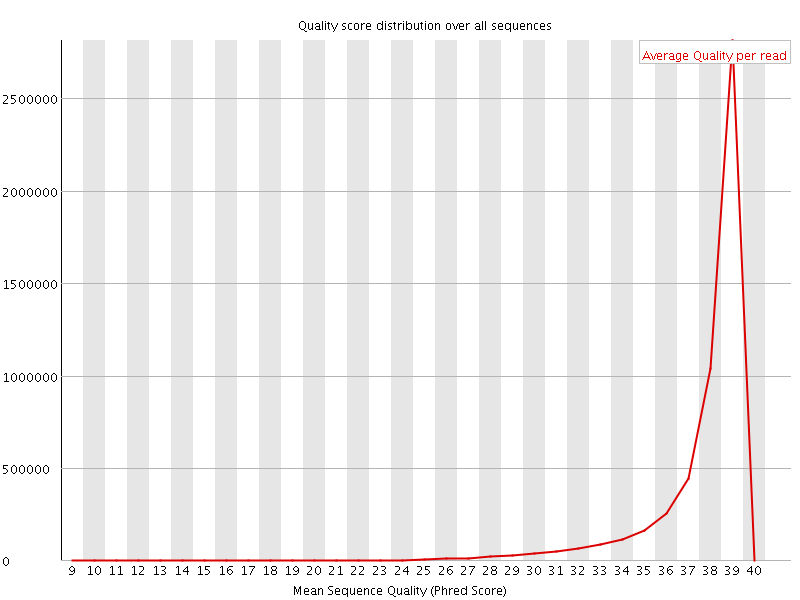
![[WARN]](Icons/warning.png) Per base sequence content
Per base sequence content
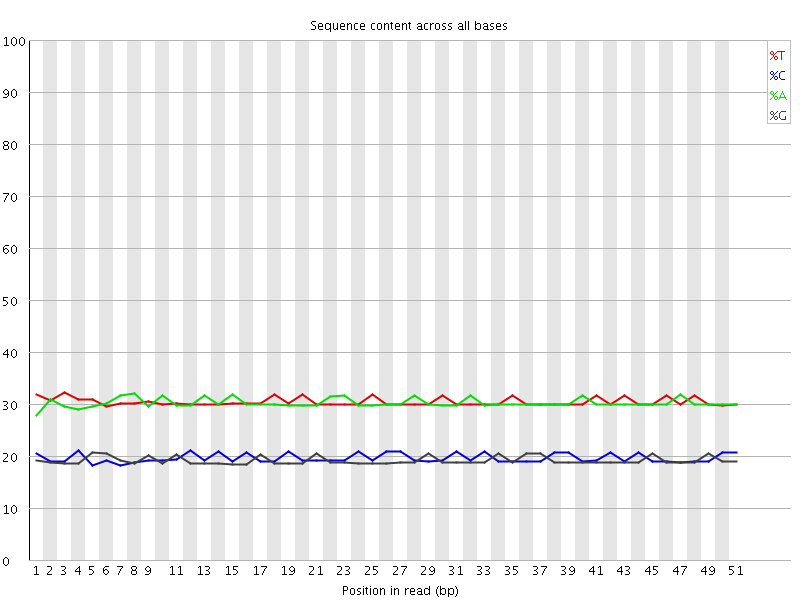
![[OK]](Icons/tick.png) Per base GC content
Per base GC content
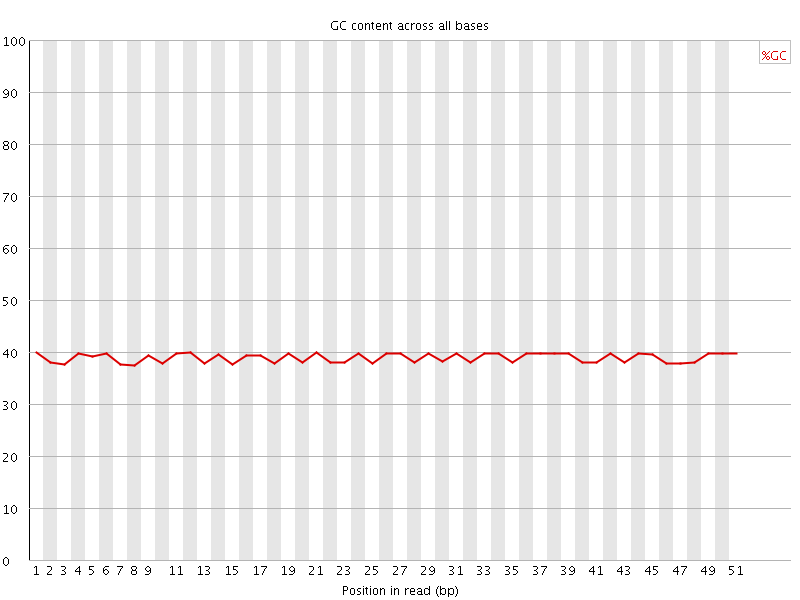
![[OK]](Icons/tick.png) Per sequence GC content
Per sequence GC content
![[OK]](Icons/tick.png) Per base N content
Per base N content
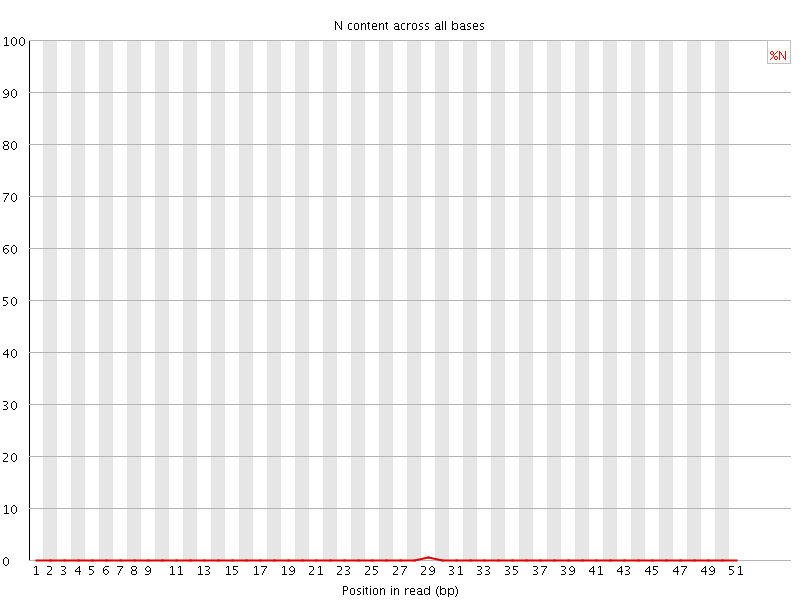
![[OK]](Icons/tick.png) Sequence Length Distribution
Sequence Length Distribution
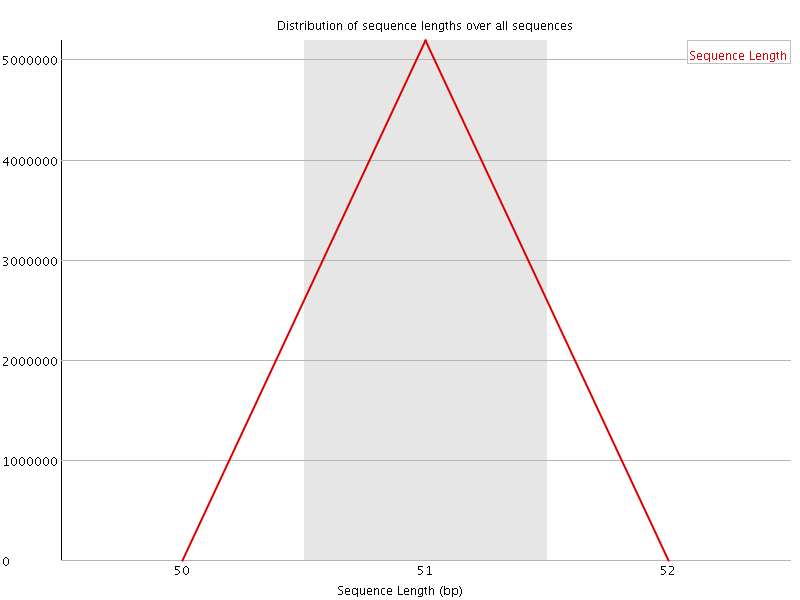
![[OK]](Icons/tick.png) Sequence Duplication Levels
Sequence Duplication Levels
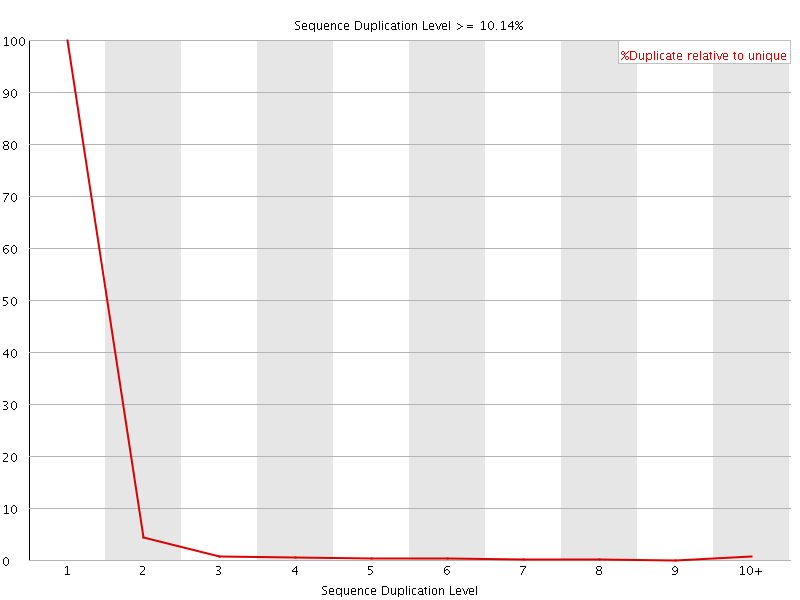
![[FAIL]](Icons/error.png) Overrepresented sequences
Overrepresented sequences
| Sequence | Count | Percentage | Possible Source |
|---|---|---|---|
| GATCGGAAGAGCACACGTCTGAACTCCAGTCACGTGGCCATCTCGTATGCC | 84964 | 1.6378236168598446 | TruSeq Adapter, Index 1 (97% over 35bp) |
![[WARN]](Icons/warning.png) Kmer Content
Kmer Content
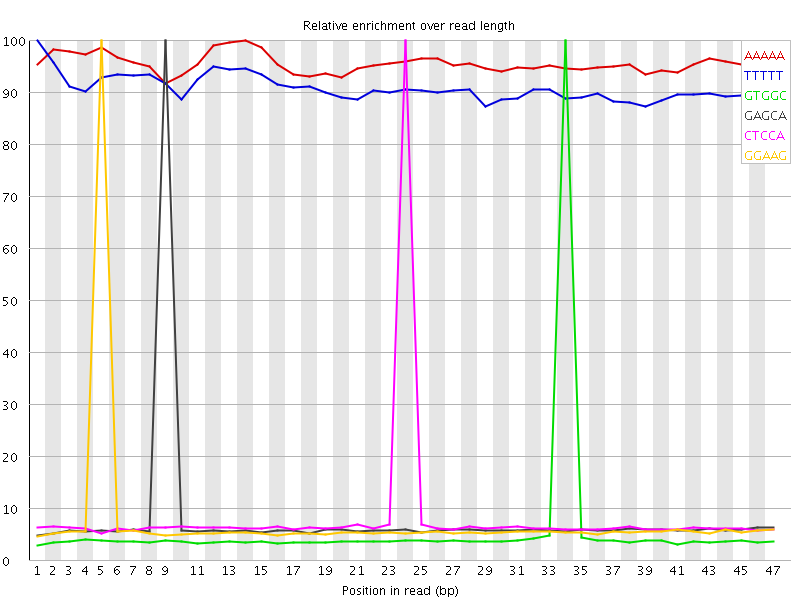
| Sequence | Count | Obs/Exp Overall | Obs/Exp Max | Max Obs/Exp Position |
|---|---|---|---|---|
| AAAAA | 2258855 | 3.579368 | 3.7442493 | 14 |
| TTTTT | 2292765 | 3.538884 | 3.894554 | 1 |
| GTGGC | 256660 | 2.4262426 | 41.62693 | 34 |
| GAGCA | 369335 | 2.2299025 | 28.28038 | 9 |
| CTCCA | 384755 | 2.2065077 | 26.540287 | 24 |
| GGAAG | 351890 | 2.1742194 | 29.099272 | 5 |
| CGGAA | 355225 | 2.1447117 | 28.468878 | 4 |
| CGTGG | 224665 | 2.1237893 | 41.50968 | 33 |
| GAAGA | 540365 | 2.121245 | 19.073122 | 6 |
| TCCAG | 349660 | 2.052098 | 27.06831 | 25 |
| CACGT | 346865 | 2.0356948 | 26.62879 | 14 |
| CTGAA | 532665 | 2.0325654 | 18.320847 | 19 |
| GGCCA | 218750 | 2.0313058 | 40.34952 | 36 |
| TGGCC | 218725 | 2.0204284 | 39.943413 | 35 |
| TCGGA | 301330 | 1.8097789 | 27.995398 | 3 |
| TCTCG | 306175 | 1.7874738 | 26.085165 | 41 |
| CCATC | 303685 | 1.7415845 | 25.384886 | 38 |
| AGAGC | 284835 | 1.719724 | 27.784332 | 8 |
| CACAC | 296925 | 1.7117889 | 26.470497 | 12 |
| AGCAC | 289555 | 1.7083055 | 27.20811 | 10 |
| GCACA | 288885 | 1.7043526 | 27.138865 | 11 |
| TCTGA | 430450 | 1.6339202 | 17.8728 | 18 |
| ATCGG | 265135 | 1.592393 | 27.631659 | 2 |
| CCAGT | 262330 | 1.5395726 | 26.463404 | 26 |
| ACGTG | 249690 | 1.4996307 | 26.53461 | 32 |
| AAGAG | 379390 | 1.4893252 | 18.50364 | 7 |
| GTCAC | 252845 | 1.4839065 | 26.188694 | 29 |
| CTCGT | 254090 | 1.4833974 | 25.687796 | 42 |
| TGAAC | 384320 | 1.4665043 | 17.806591 | 20 |
| CGTCT | 249805 | 1.4583812 | 26.38273 | 16 |
| GTCTG | 242810 | 1.4506663 | 27.001848 | 17 |
| GCCAT | 245180 | 1.4389218 | 25.69921 | 37 |
| ACTCC | 247595 | 1.4199173 | 25.856136 | 23 |
| ACACG | 240440 | 1.4185387 | 26.742779 | 13 |
| ATGCC | 237975 | 1.396637 | 25.794592 | 47 |
| GATCG | 229505 | 1.3784003 | 27.281582 | 1 |
| TCACG | 232815 | 1.3663538 | 26.041458 | 30 |
| ACGTC | 232375 | 1.3637713 | 26.458366 | 15 |
| CAGTC | 227365 | 1.3343685 | 26.215055 | 27 |
| AACTC | 357380 | 1.3325685 | 17.269484 | 22 |
| CATCT | 350745 | 1.300974 | 16.70005 | 39 |
| GAACT | 337640 | 1.2883809 | 17.552158 | 21 |
| ATCTC | 342980 | 1.2721722 | 16.71239 | 40 |
| AGTCA | 304710 | 1.1627252 | 17.285538 | 28 |
| TCGTA | 260300 | 0.9880576 | 16.689371 | 43 |
| TATGC | 241735 | 0.9175879 | 16.646488 | 46 |
| GTATG | 233190 | 0.905835 | 17.065222 | 45 |
| CGTAT | 232695 | 0.8832735 | 16.653181 | 44 |