![[OK]](Icons/tick.png) Basic Statistics
Basic Statistics
| Measure | Value |
|---|---|
| Filename | LW176_GTGAAA_L002_R1_001.fastq.gz |
| File type | Conventional base calls |
| Encoding | Sanger / Illumina 1.9 |
| Total Sequences | 1447683 |
| Filtered Sequences | 0 |
| Sequence length | 51 |
| %GC | 40 |
![[OK]](Icons/tick.png) Per base sequence quality
Per base sequence quality
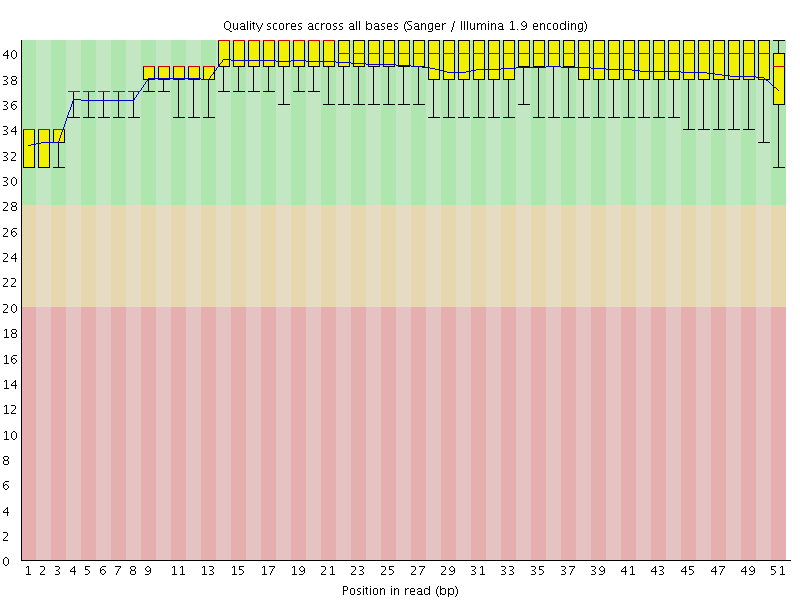
![[OK]](Icons/tick.png) Per sequence quality scores
Per sequence quality scores
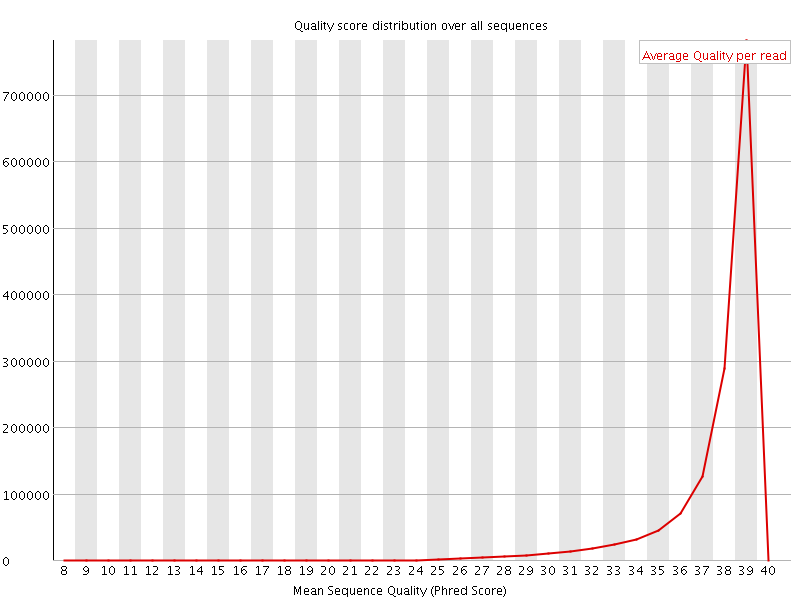
![[WARN]](Icons/warning.png) Per base sequence content
Per base sequence content
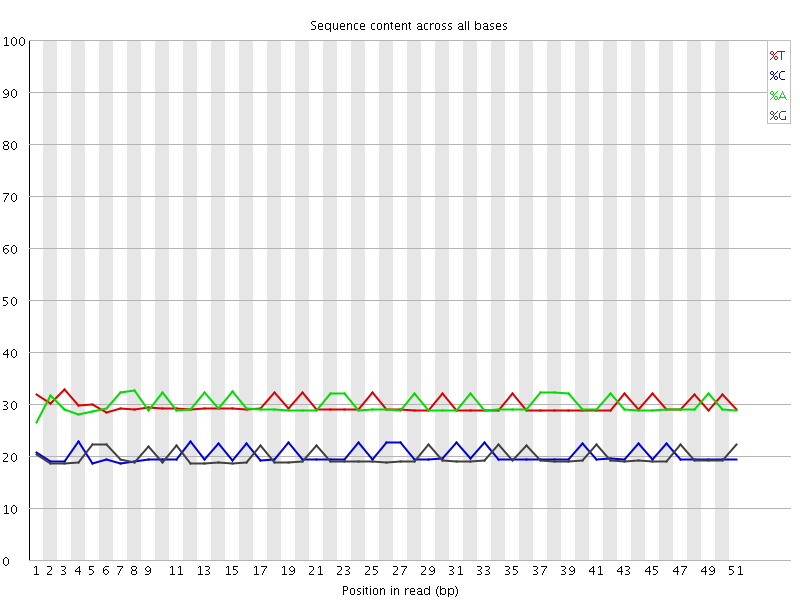
![[OK]](Icons/tick.png) Per base GC content
Per base GC content
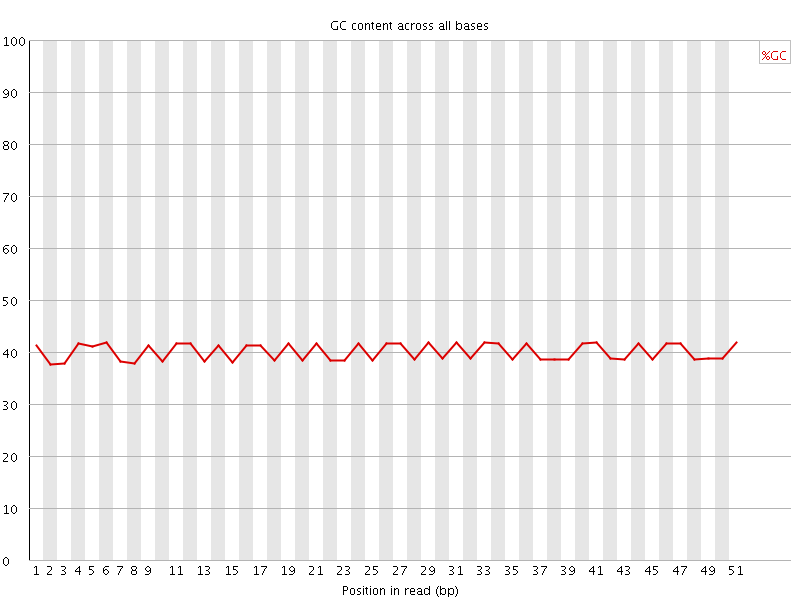
![[OK]](Icons/tick.png) Per sequence GC content
Per sequence GC content
![[OK]](Icons/tick.png) Per base N content
Per base N content
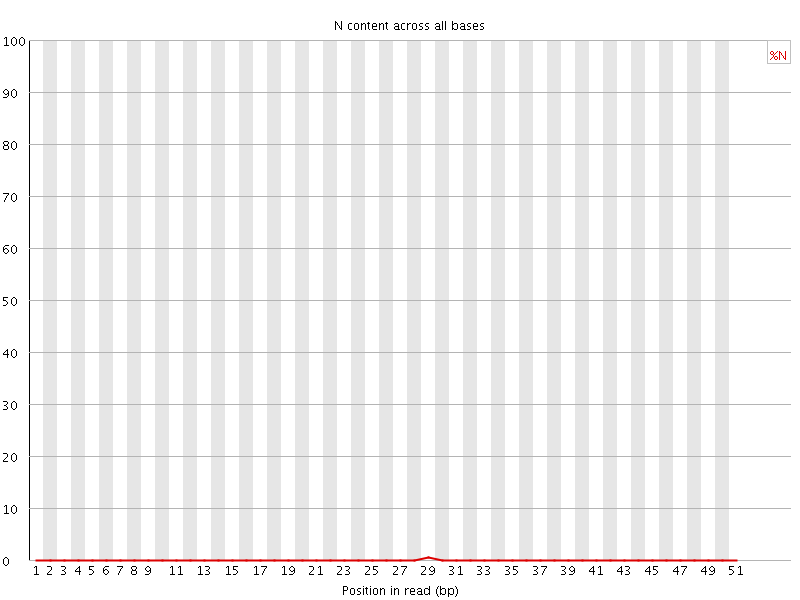
![[OK]](Icons/tick.png) Sequence Length Distribution
Sequence Length Distribution
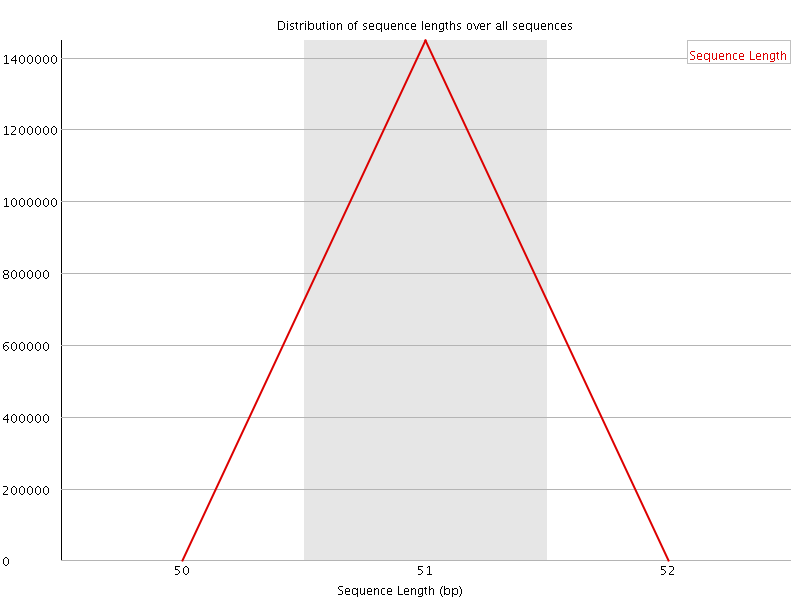
![[OK]](Icons/tick.png) Sequence Duplication Levels
Sequence Duplication Levels
![[FAIL]](Icons/error.png) Overrepresented sequences
Overrepresented sequences
| Sequence | Count | Percentage | Possible Source |
|---|---|---|---|
| GATCGGAAGAGCACACGTCTGAACTCCAGTCACGTGAAACGATCTCGTATG | 39052 | 2.697551881178407 | TruSeq Adapter, Index 1 (97% over 35bp) |
![[WARN]](Icons/warning.png) Kmer Content
Kmer Content
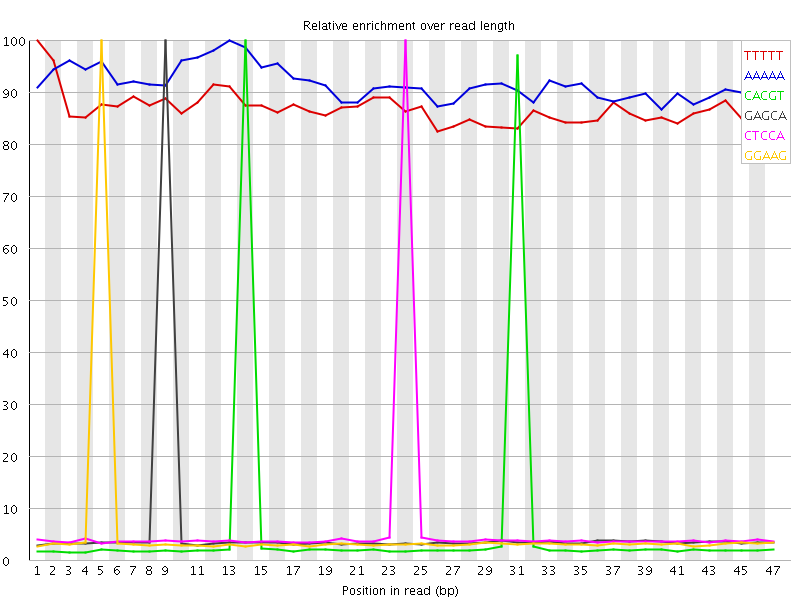
| Sequence | Count | Obs/Exp Overall | Obs/Exp Max | Max Obs/Exp Position |
|---|---|---|---|---|
| TTTTT | 553035 | 3.4309046 | 3.9439714 | 1 |
| AAAAA | 550695 | 3.358978 | 3.6639688 | 13 |
| CACGT | 139925 | 2.8066747 | 45.46053 | 14 |
| GAGCA | 128475 | 2.6251183 | 47.345295 | 9 |
| CTCCA | 132990 | 2.6098187 | 44.26523 | 24 |
| GGAAG | 121490 | 2.5373256 | 48.339043 | 5 |
| TGAAA | 270800 | 2.498024 | 21.643097 | 35 |
| TCCAG | 121375 | 2.4345913 | 45.307503 | 25 |
| GAAGA | 175605 | 2.4332833 | 32.705692 | 6 |
| CGGAA | 117920 | 2.409449 | 47.24937 | 4 |
| CTGAA | 166070 | 2.258986 | 31.57199 | 19 |
| TCGGA | 105300 | 2.1588905 | 47.140278 | 3 |
| TCTCG | 107240 | 2.1583686 | 43.292557 | 43 |
| AGAGC | 103700 | 2.118893 | 46.908825 | 8 |
| AGCAC | 105680 | 2.1126018 | 45.841663 | 10 |
| CACAC | 105350 | 2.0604112 | 44.58754 | 12 |
| GCACA | 103055 | 2.0601265 | 45.719658 | 11 |
| ATCGG | 96130 | 1.9708848 | 46.861145 | 2 |
| GAAAC | 143480 | 1.9450991 | 29.59629 | 36 |
| CCAGT | 95550 | 1.9165825 | 44.60098 | 26 |
| CTCGT | 94815 | 1.9082966 | 43.186436 | 44 |
| TCTGA | 139590 | 1.9052354 | 31.426025 | 18 |
| CGTCT | 92850 | 1.868748 | 45.190456 | 16 |
| ACGTG | 91060 | 1.866938 | 44.671593 | 32 |
| GTGAA | 134245 | 1.8664911 | 30.658655 | 34 |
| GTCAC | 91740 | 1.8401598 | 44.496662 | 29 |
| GTCTG | 89110 | 1.8331614 | 46.19264 | 17 |
| GATCG | 88425 | 1.8129146 | 46.58945 | 1 |
| ACTCC | 92060 | 1.8066015 | 43.506233 | 23 |
| AAGAG | 129760 | 1.7980288 | 32.16575 | 7 |
| ACACG | 88920 | 1.77756 | 45.23632 | 13 |
| ACGTC | 88470 | 1.7745686 | 45.028126 | 15 |
| CGTGA | 86455 | 1.772525 | 44.432144 | 33 |
| TCACG | 87600 | 1.757118 | 44.211277 | 30 |
| CAGTC | 86320 | 1.731443 | 44.387596 | 27 |
| TGAAC | 126640 | 1.722635 | 31.06101 | 20 |
| AAACG | 124170 | 1.6833215 | 29.266706 | 37 |
| CGATC | 83240 | 1.6696632 | 41.58654 | 40 |
| AACTC | 119440 | 1.5895226 | 30.113577 | 22 |
| GAACT | 115165 | 1.566545 | 30.728876 | 21 |
| ATCTC | 116960 | 1.561803 | 28.707554 | 42 |
| AACGA | 113560 | 1.539486 | 28.974148 | 38 |
| ACGAT | 106445 | 1.4479301 | 28.90722 | 39 |
| GATCT | 105570 | 1.4409035 | 29.068768 | 41 |
| AGTCA | 105690 | 1.4376605 | 30.435863 | 28 |
| TCGTA | 95560 | 1.3042791 | 29.562862 | 45 |
| GTATG | 85635 | 1.1946787 | 29.982744 | 47 |
| CGTAT | 86440 | 1.1798021 | 29.47229 | 46 |