![[OK]](Icons/tick.png) Basic Statistics
Basic Statistics
| Measure | Value |
|---|---|
| Filename | LW175_GTCCGC_L002_R1_001.fastq.gz |
| File type | Conventional base calls |
| Encoding | Sanger / Illumina 1.9 |
| Total Sequences | 4428898 |
| Filtered Sequences | 0 |
| Sequence length | 51 |
| %GC | 40 |
![[OK]](Icons/tick.png) Per base sequence quality
Per base sequence quality
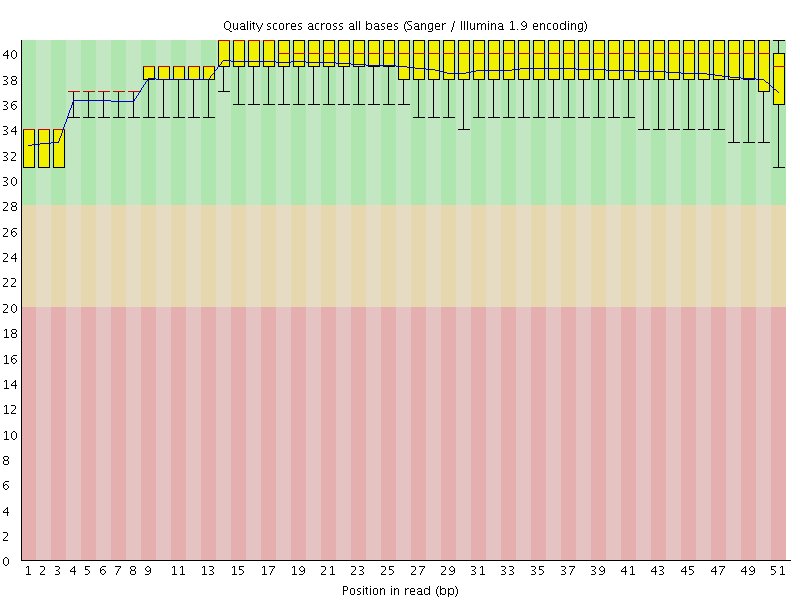
![[OK]](Icons/tick.png) Per sequence quality scores
Per sequence quality scores
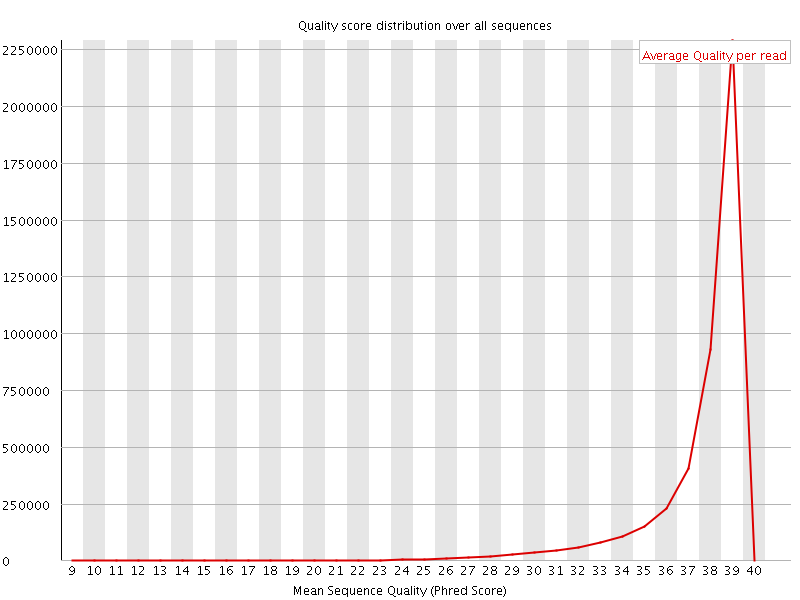
![[WARN]](Icons/warning.png) Per base sequence content
Per base sequence content
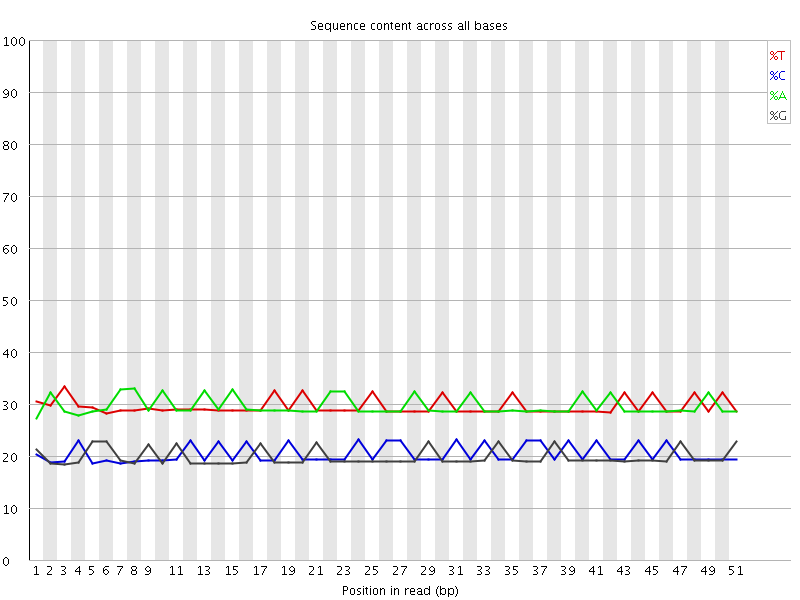
![[OK]](Icons/tick.png) Per base GC content
Per base GC content
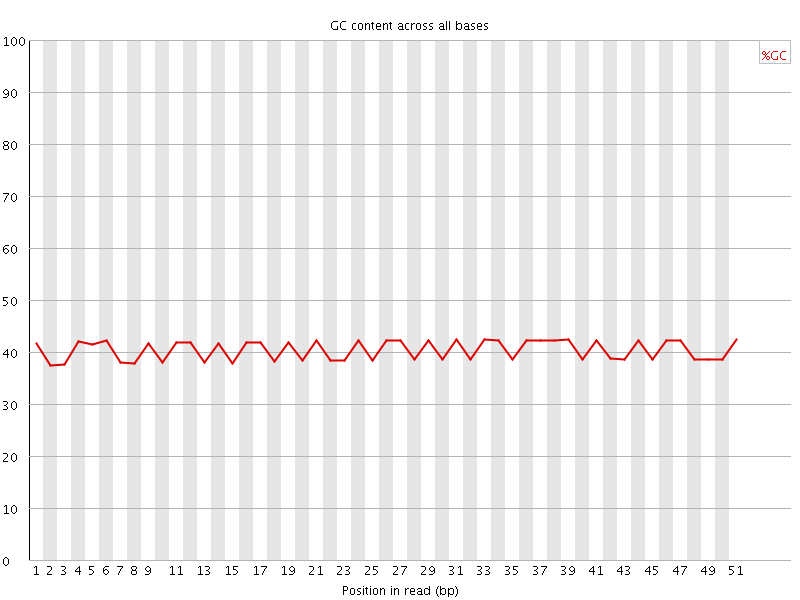
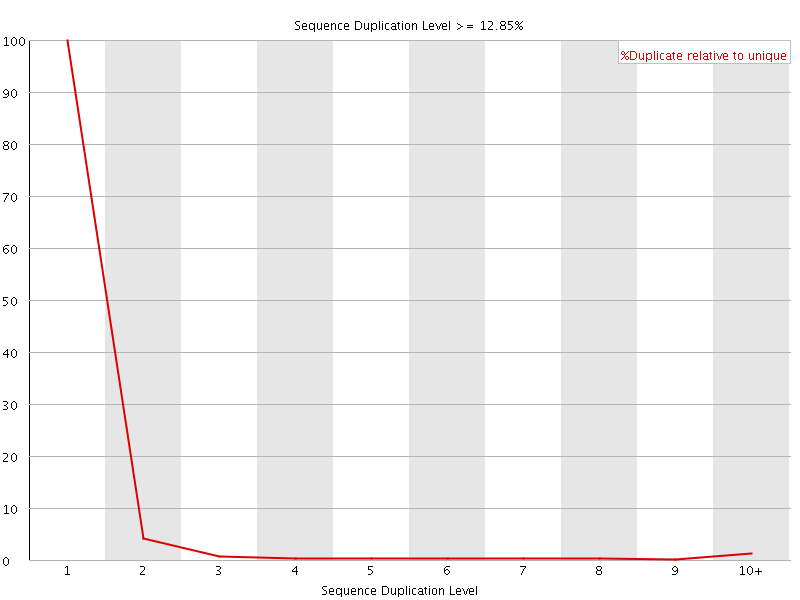
![[OK]](Icons/tick.png) Per sequence GC content
Per sequence GC content
![[OK]](Icons/tick.png) Per base N content
Per base N content
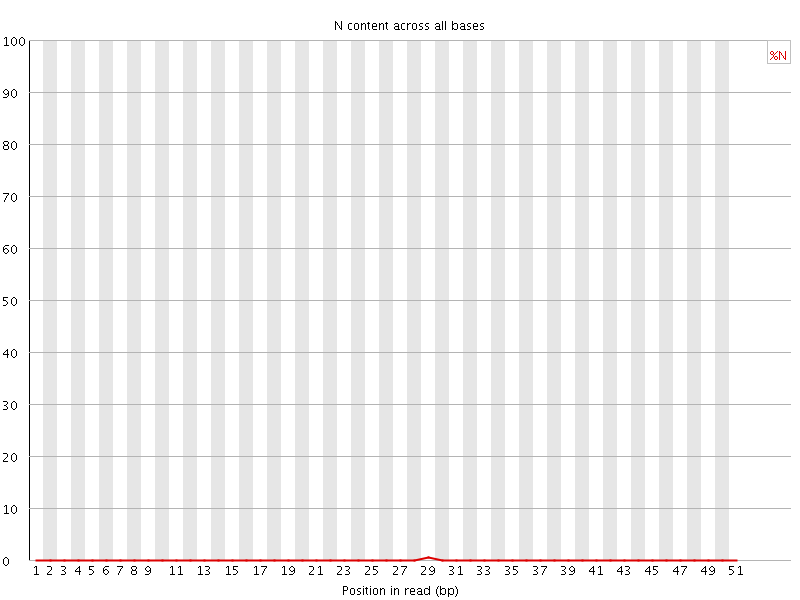
![[OK]](Icons/tick.png) Sequence Length Distribution
Sequence Length Distribution
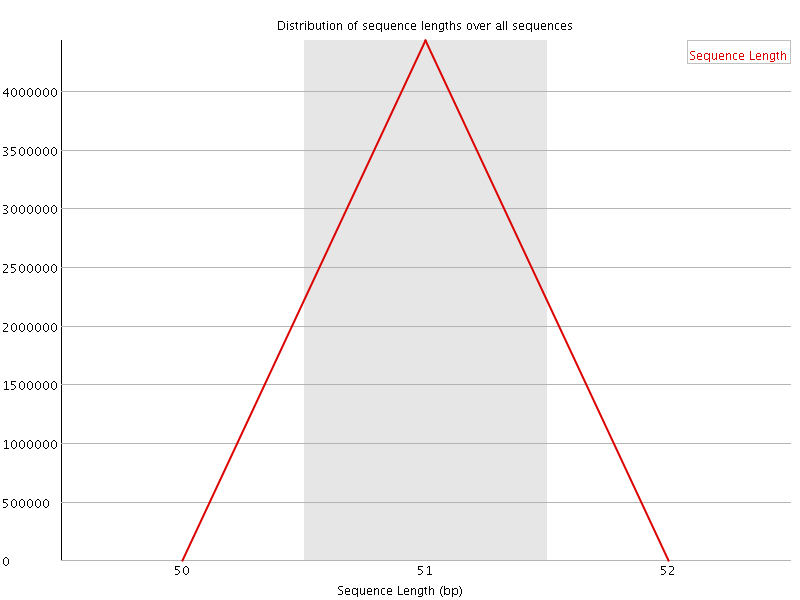
![[OK]](Icons/tick.png) Sequence Duplication Levels
Sequence Duplication Levels
![[FAIL]](Icons/error.png) Overrepresented sequences
Overrepresented sequences
| Sequence | Count | Percentage | Possible Source |
|---|---|---|---|
| GATCGGAAGAGCACACGTCTGAACTCCAGTCACGTCCGCACATCTCGTATG | 142386 | 3.214930666725673 | TruSeq Adapter, Index 1 (97% over 36bp) |
![[WARN]](Icons/warning.png) Kmer Content
Kmer Content
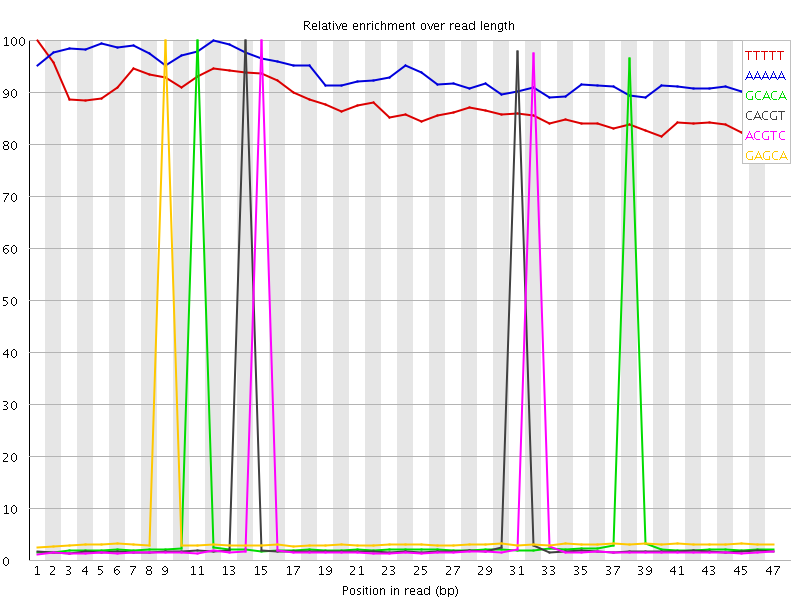
| Sequence | Count | Obs/Exp Overall | Obs/Exp Max | Max Obs/Exp Position |
|---|---|---|---|---|
| TTTTT | 1586135 | 3.317036 | 3.77247 | 1 |
| AAAAA | 1592425 | 3.2664635 | 3.4886794 | 12 |
| GCACA | 499680 | 3.2001812 | 51.01777 | 11 |
| CACGT | 473760 | 3.045925 | 50.863636 | 14 |
| ACGTC | 456850 | 2.9372065 | 50.62336 | 15 |
| GAGCA | 416225 | 2.7540271 | 53.233295 | 9 |
| GGAAG | 400065 | 2.7348156 | 55.708115 | 5 |
| CTCCA | 429940 | 2.6755388 | 48.953407 | 24 |
| GAAGA | 567030 | 2.5950394 | 37.94139 | 6 |
| TCCAG | 397630 | 2.556466 | 50.608356 | 25 |
| TCCGC | 271635 | 2.5249264 | 70.41586 | 35 |
| CGGAA | 379895 | 2.5136428 | 53.654217 | 4 |
| CCGCA | 260335 | 2.4105563 | 70.45051 | 36 |
| CTGAA | 525215 | 2.335586 | 36.139244 | 19 |
| TCGGA | 349045 | 2.3184607 | 53.557907 | 3 |
| GTCCG | 240190 | 2.306616 | 72.71257 | 34 |
| TCTCG | 355890 | 2.2969675 | 50.386574 | 43 |
| CGTCC | 246280 | 2.2892442 | 70.527885 | 33 |
| AGAGC | 341145 | 2.2572467 | 52.91953 | 8 |
| AGCAC | 344090 | 2.2037113 | 51.11399 | 10 |
| CGCAC | 236385 | 2.1887927 | 70.13543 | 37 |
| CACAC | 347155 | 2.1520314 | 49.15009 | 12 |
| ATCGG | 313690 | 2.0836225 | 53.25541 | 2 |
| CTCGT | 315250 | 2.034671 | 49.9994 | 44 |
| CCAGT | 314655 | 2.022998 | 49.968506 | 26 |
| TCTGA | 448290 | 2.0012255 | 35.949013 | 18 |
| CGTCT | 308650 | 1.9920737 | 50.840576 | 16 |
| GTCTG | 297815 | 1.9858347 | 52.619473 | 17 |
| GTCAC | 302910 | 1.9474865 | 50.485188 | 29 |
| AAGAG | 425380 | 1.9467715 | 37.27315 | 7 |
| GATCG | 291365 | 1.9353327 | 52.954742 | 1 |
| ACTCC | 305345 | 1.9001777 | 48.344486 | 23 |
| TCACG | 292505 | 1.8805901 | 50.151455 | 30 |
| ACACG | 292835 | 1.8754505 | 50.641907 | 13 |
| CAGTC | 288200 | 1.8529122 | 50.216183 | 27 |
| TGAAC | 410360 | 1.8248355 | 35.543682 | 20 |
| CATCT | 396530 | 1.7133875 | 33.56382 | 41 |
| AACTC | 390405 | 1.6804154 | 34.041065 | 22 |
| ATCTC | 387190 | 1.6730299 | 33.75738 | 42 |
| GAACT | 374460 | 1.6651913 | 35.34218 | 21 |
| CACAT | 383615 | 1.6511893 | 33.316654 | 39 |
| ACATC | 365060 | 1.5713233 | 33.282623 | 40 |
| AGTCA | 343745 | 1.5286044 | 35.044098 | 28 |
| TCGTA | 314240 | 1.4028087 | 34.613483 | 45 |
| GTATG | 281650 | 1.2989849 | 35.452656 | 47 |
| CGTAT | 285825 | 1.2759604 | 34.401783 | 46 |