![[OK]](Icons/tick.png) Basic Statistics
Basic Statistics
| Measure | Value |
|---|---|
| Filename | lane2_Undetermined_L002_R1_001.fastq.gz |
| File type | Conventional base calls |
| Encoding | Sanger / Illumina 1.9 |
| Total Sequences | 5345021 |
| Filtered Sequences | 0 |
| Sequence length | 51 |
| %GC | 44 |
![[OK]](Icons/tick.png) Per base sequence quality
Per base sequence quality
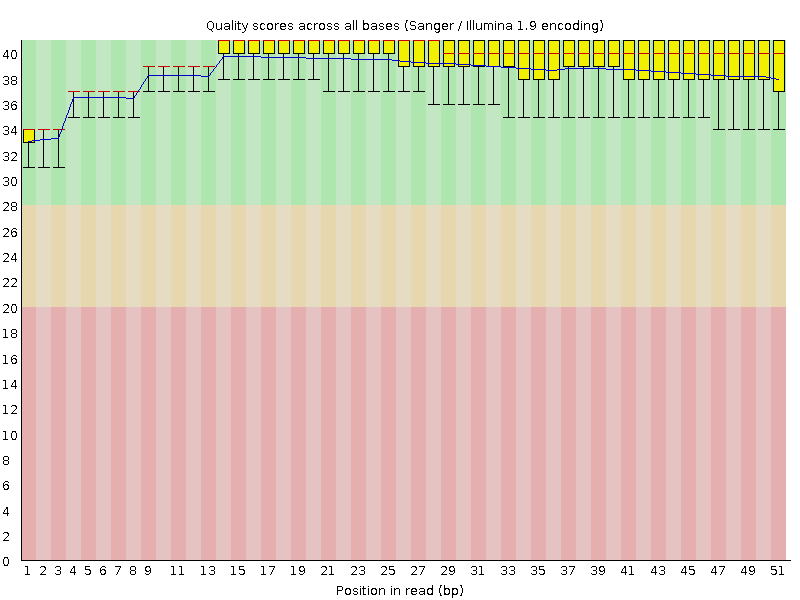
![[OK]](Icons/tick.png) Per sequence quality scores
Per sequence quality scores
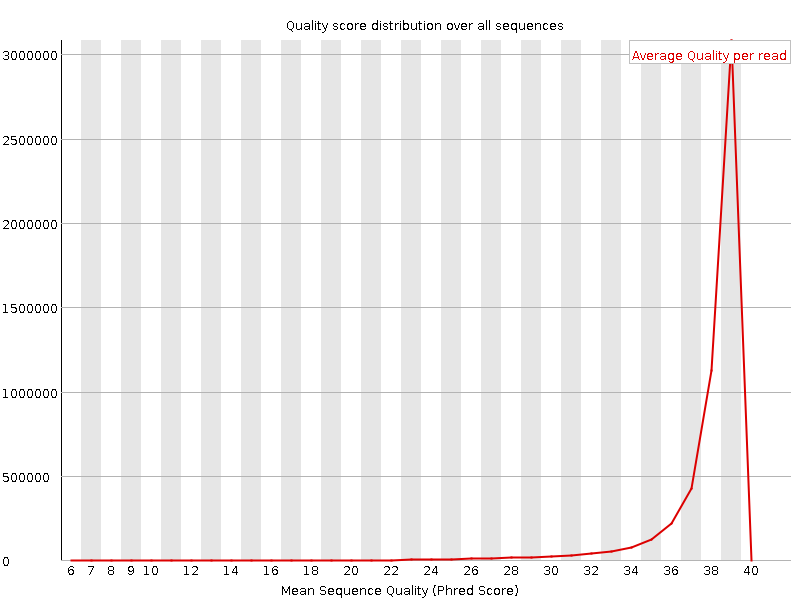
![[OK]](Icons/tick.png) Per base sequence content
Per base sequence content
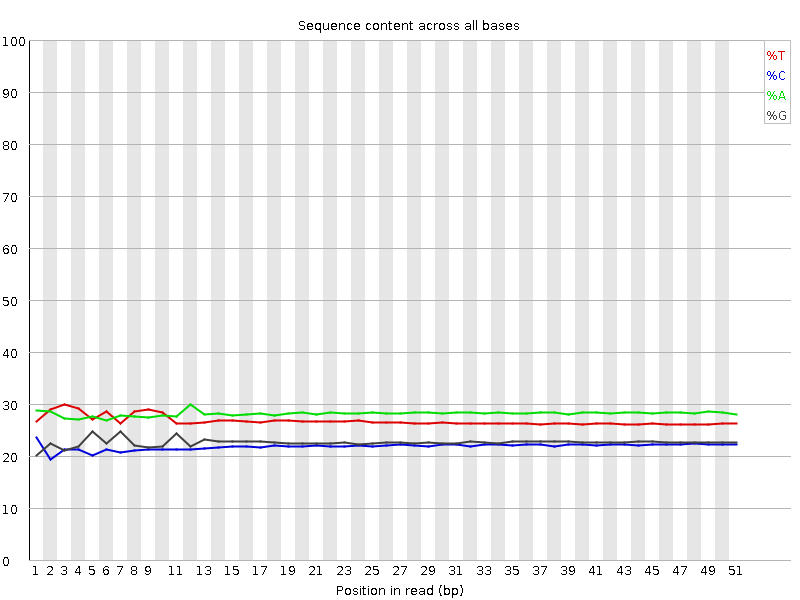
![[OK]](Icons/tick.png) Per base GC content
Per base GC content
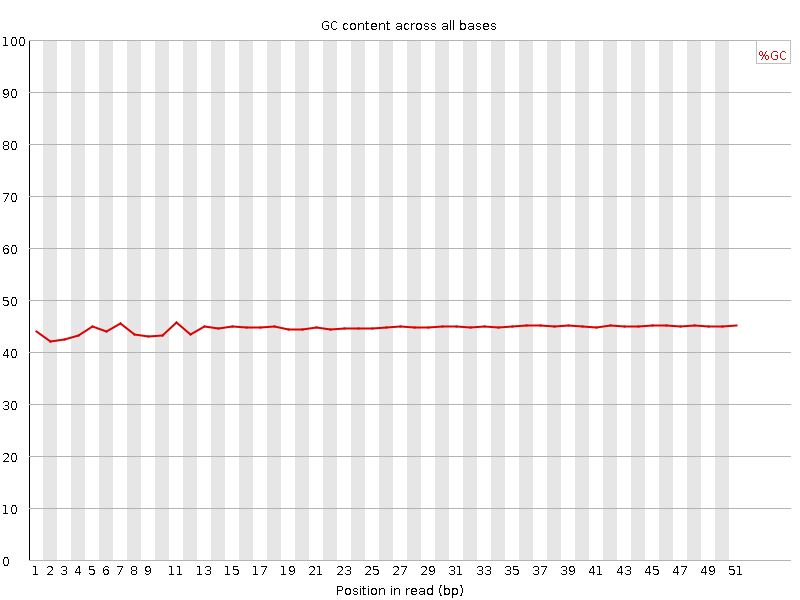
![[WARN]](Icons/warning.png) Per sequence GC content
Per sequence GC content
![[OK]](Icons/tick.png) Per base N content
Per base N content
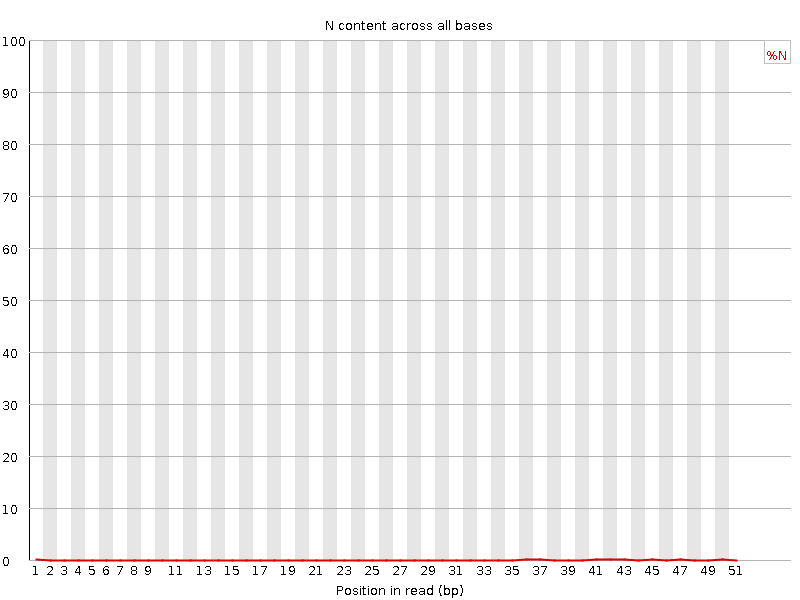
![[OK]](Icons/tick.png) Sequence Length Distribution
Sequence Length Distribution
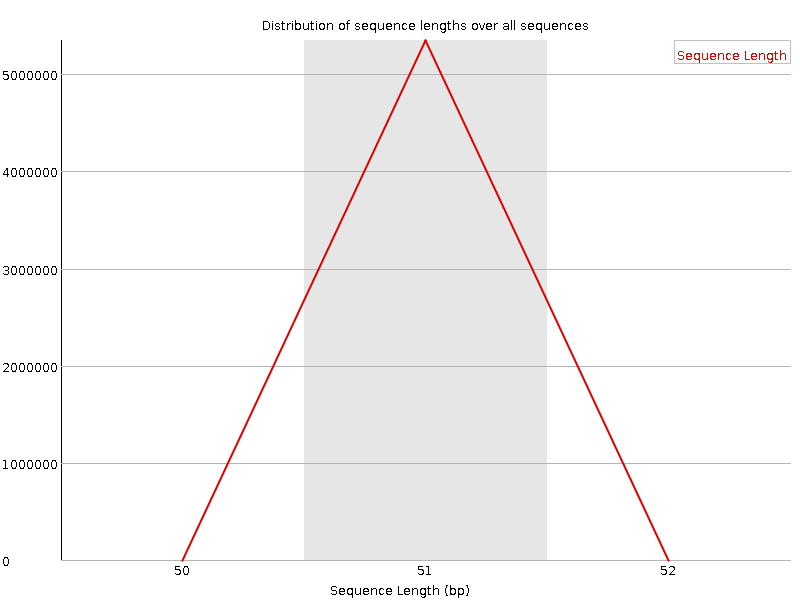
![[FAIL]](Icons/error.png) Sequence Duplication Levels
Sequence Duplication Levels
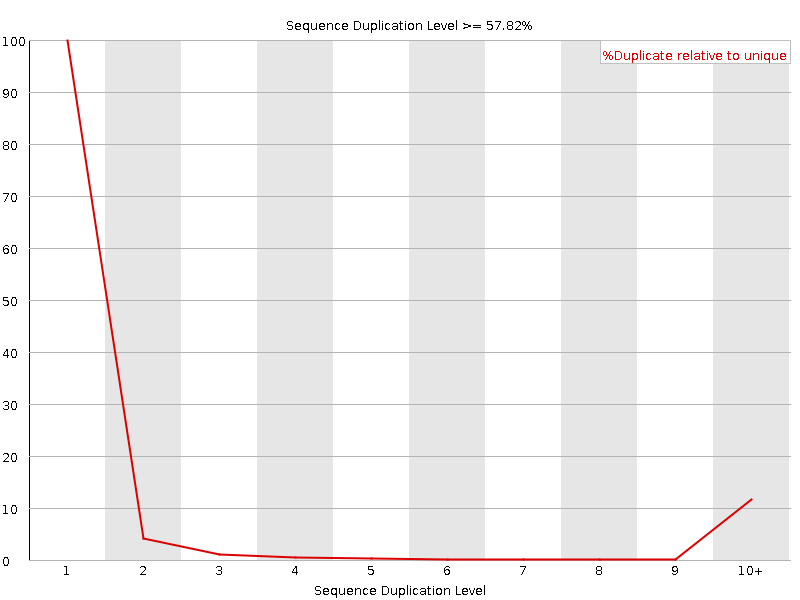
![[OK]](Icons/tick.png) Overrepresented sequences
Overrepresented sequences
No overrepresented sequences
![[WARN]](Icons/warning.png) Kmer Content
Kmer Content
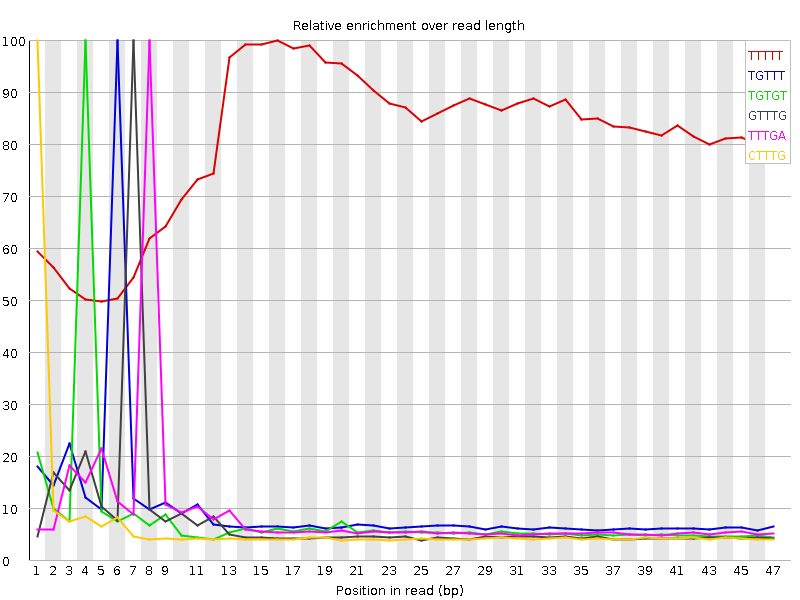
| Sequence | Count | Obs/Exp Overall | Obs/Exp Max | Max Obs/Exp Position |
|---|---|---|---|---|
| TTTTT | 1276045 | 3.59604 | 4.443601 | 16 |
| TGTTT | 684375 | 2.2821295 | 23.234575 | 6 |
| TGTGT | 551280 | 2.1752338 | 26.714806 | 4 |
| GTTTG | 539430 | 2.1284766 | 26.799728 | 7 |
| TTTGA | 612540 | 1.9424441 | 21.798687 | 8 |
| CTTTG | 449860 | 1.835021 | 27.356136 | 1 |
| TTTGT | 542915 | 1.8104143 | 22.757082 | 2 |
| GTGTT | 448395 | 1.7692715 | 26.518196 | 5 |
| GAGGG | 298970 | 1.5707407 | 6.1694617 | 11 |
| TTGAG | 413280 | 1.5507653 | 14.9029 | 9 |
| TTGTG | 392655 | 1.5493335 | 26.48059 | 3 |
| TGAGG | 323800 | 1.4376929 | 8.305667 | 10 |
| TTGAC | 365210 | 1.4166881 | 5.052034 | 9 |
| TTGAT | 338015 | 1.0718896 | 5.533931 | 9 |