![[OK]](Icons/tick.png) Basic Statistics
Basic Statistics
| Measure | Value |
|---|---|
| Filename | 140701_JD_D_9_ATCACG_L006_R1_001.fastq.gz |
| File type | Conventional base calls |
| Encoding | Sanger / Illumina 1.9 |
| Total Sequences | 25704617 |
| Filtered Sequences | 0 |
| Sequence length | 51 |
| %GC | 44 |
![[OK]](Icons/tick.png) Per base sequence quality
Per base sequence quality
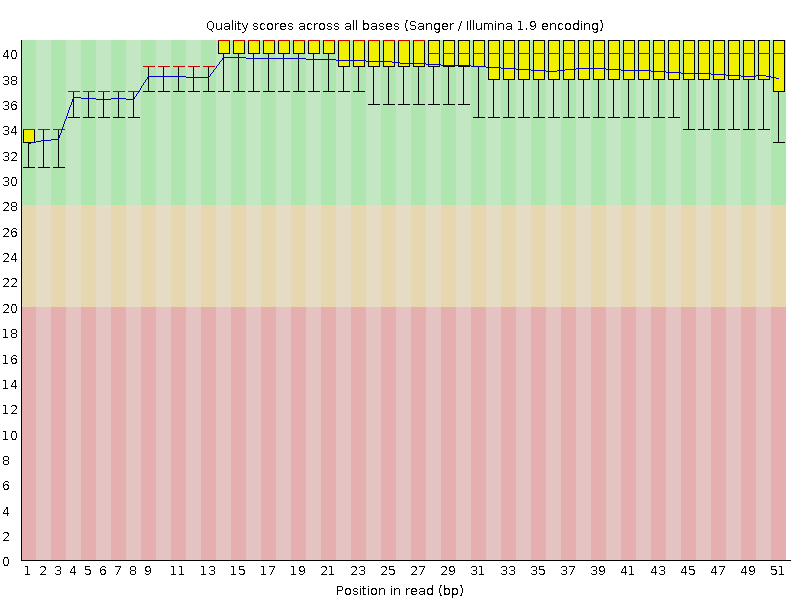
![[OK]](Icons/tick.png) Per sequence quality scores
Per sequence quality scores
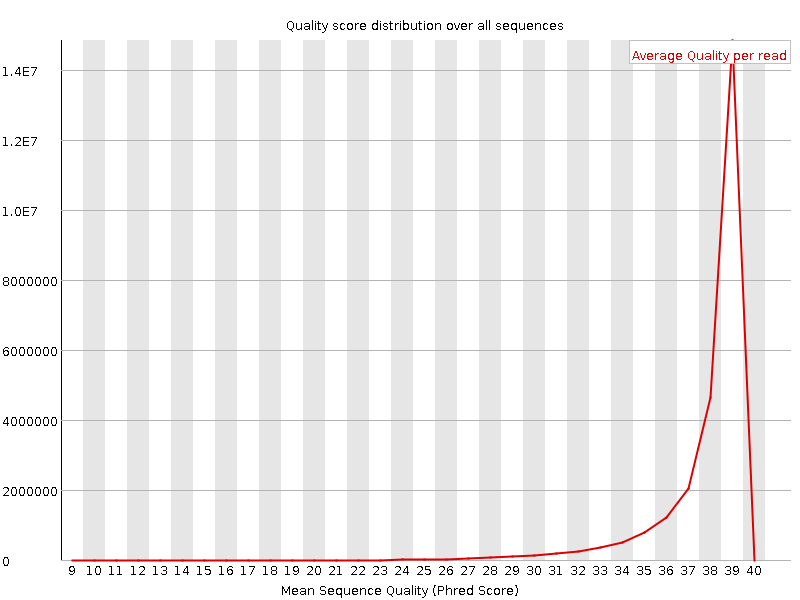
![[WARN]](Icons/warning.png) Per base sequence content
Per base sequence content

![[OK]](Icons/tick.png) Per base GC content
Per base GC content

![[OK]](Icons/tick.png) Per sequence GC content
Per sequence GC content

![[OK]](Icons/tick.png) Per base N content
Per base N content

![[OK]](Icons/tick.png) Sequence Length Distribution
Sequence Length Distribution
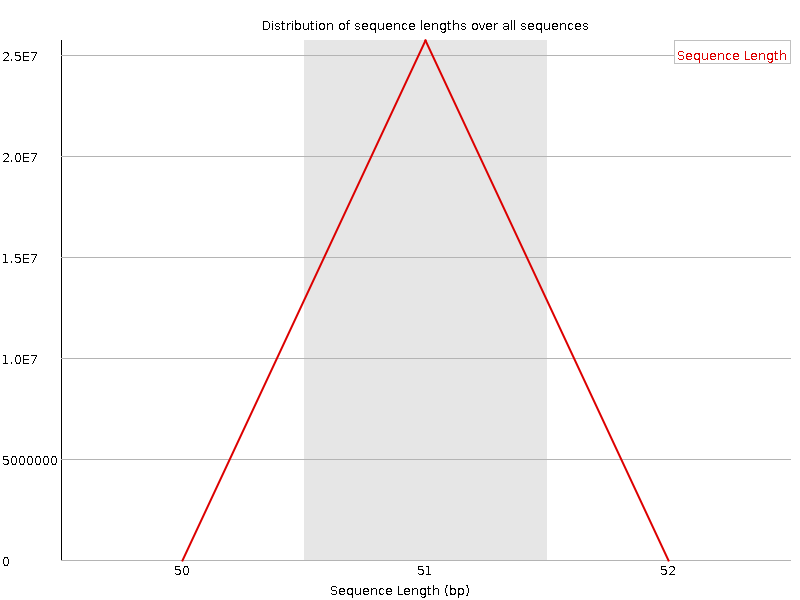
![[WARN]](Icons/warning.png) Sequence Duplication Levels
Sequence Duplication Levels
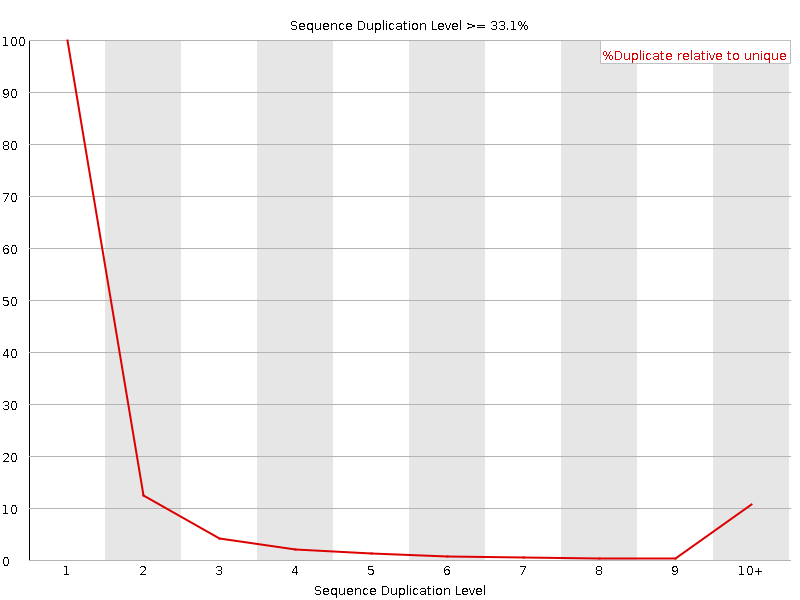
![[OK]](Icons/tick.png) Overrepresented sequences
Overrepresented sequences
No overrepresented sequences
![[WARN]](Icons/warning.png) Kmer Content
Kmer Content
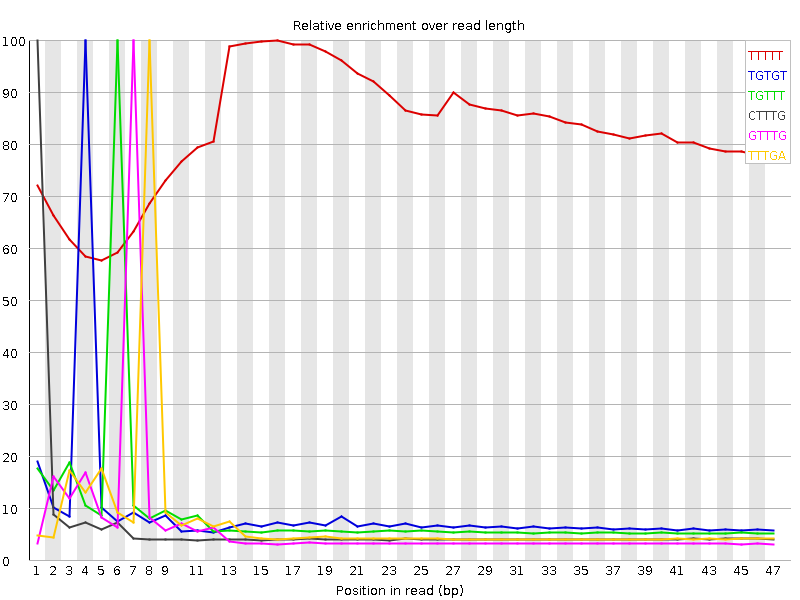
| Sequence | Count | Obs/Exp Overall | Obs/Exp Max | Max Obs/Exp Position |
|---|---|---|---|---|
| TTTTT | 7740470 | 3.5868351 | 4.3406186 | 16 |
| TGTGT | 4167270 | 3.0000243 | 32.982574 | 4 |
| TGTTT | 4048600 | 2.3383715 | 26.570139 | 6 |
| CTTTG | 2920580 | 2.1817725 | 33.507507 | 1 |
| GTTTG | 2996210 | 2.1569762 | 32.26548 | 7 |
| TTTGA | 3446320 | 2.053298 | 26.98084 | 8 |
| GAGGG | 1717955 | 1.9819912 | 8.635853 | 11 |
| GTGTT | 2732685 | 1.9672643 | 32.225567 | 5 |
| TTTGT | 3360365 | 1.9408638 | 26.190495 | 2 |
| TTGTG | 2512205 | 1.8085403 | 32.184837 | 3 |
| TGAGG | 1845845 | 1.7085221 | 11.423341 | 10 |
| TTGAG | 2156785 | 1.6016505 | 19.212976 | 9 |
| TGAGT | 1533730 | 1.1389635 | 5.61615 | 10 |
| TTGAT | 1770150 | 1.0546453 | 6.3229523 | 9 |
| TTGAC | 1203710 | 0.92757684 | 5.9031343 | 9 |