![[OK]](Icons/tick.png) Basic Statistics
Basic Statistics
| Measure | Value |
|---|---|
| Filename | 140701_JD_D_15_TGACCA_L006_R2_001.fastq.gz |
| File type | Conventional base calls |
| Encoding | Sanger / Illumina 1.9 |
| Total Sequences | 28082252 |
| Filtered Sequences | 0 |
| Sequence length | 51 |
| %GC | 42 |
![[OK]](Icons/tick.png) Per base sequence quality
Per base sequence quality
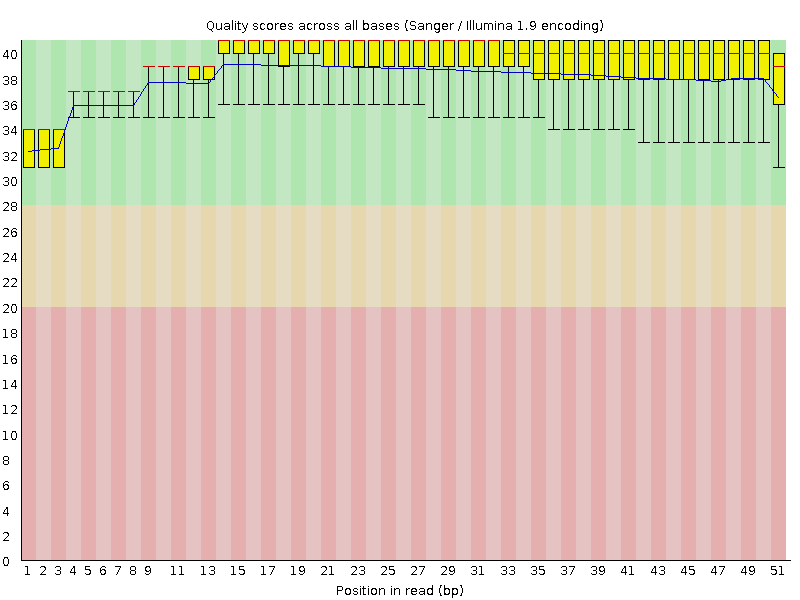
![[OK]](Icons/tick.png) Per sequence quality scores
Per sequence quality scores
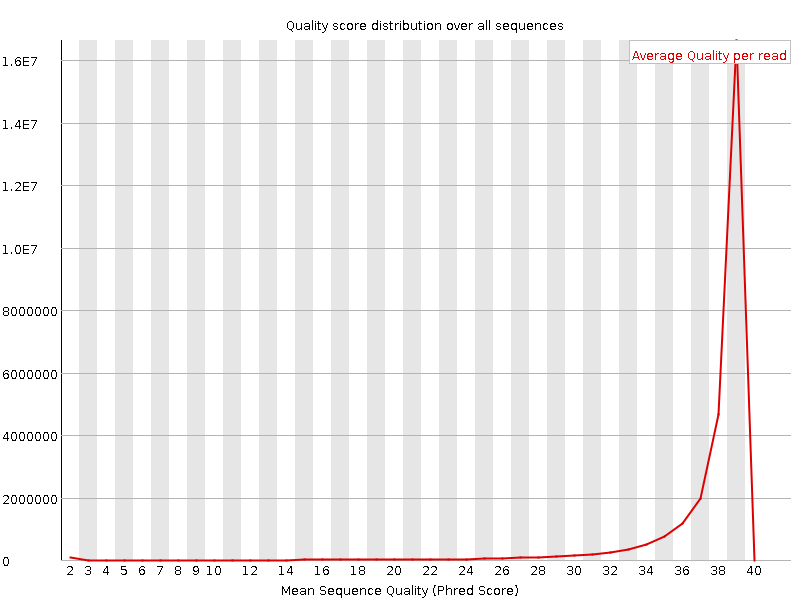
![[WARN]](Icons/warning.png) Per base sequence content
Per base sequence content

![[OK]](Icons/tick.png) Per base GC content
Per base GC content

![[OK]](Icons/tick.png) Per sequence GC content
Per sequence GC content

![[OK]](Icons/tick.png) Per base N content
Per base N content

![[OK]](Icons/tick.png) Sequence Length Distribution
Sequence Length Distribution
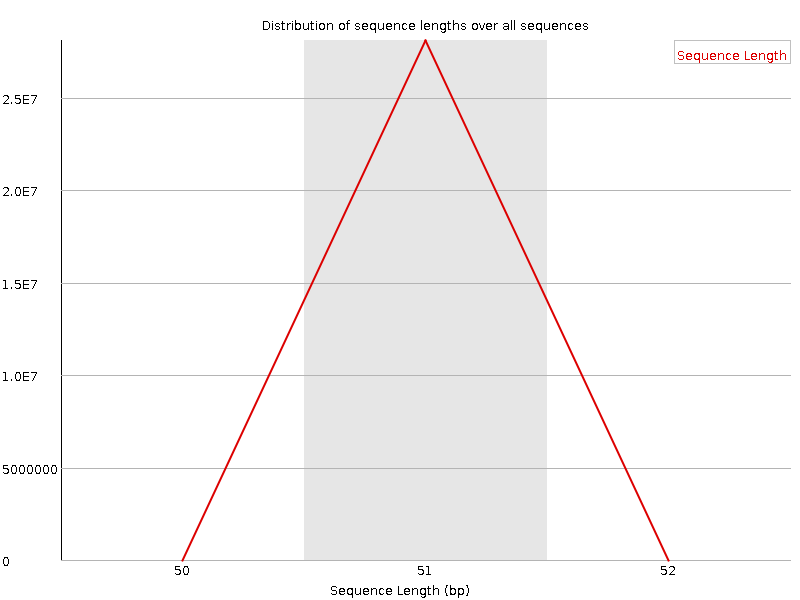
![[WARN]](Icons/warning.png) Sequence Duplication Levels
Sequence Duplication Levels
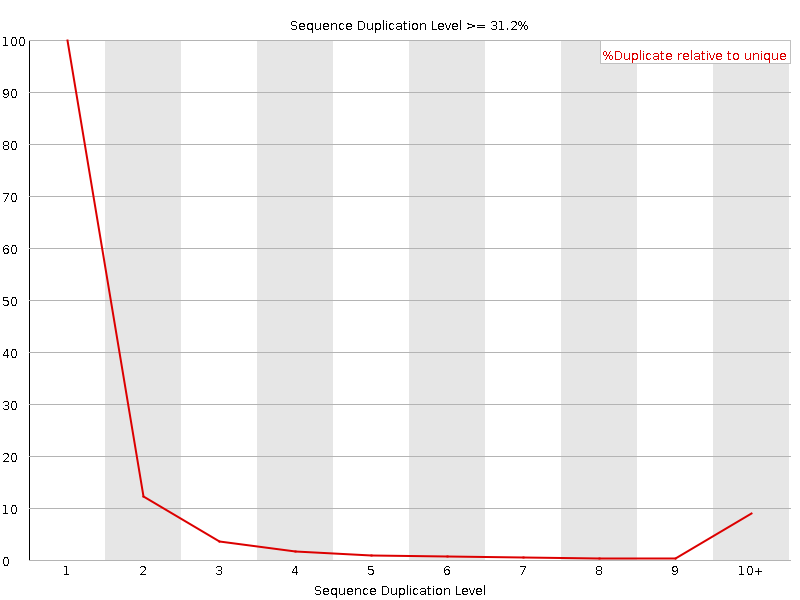
![[WARN]](Icons/warning.png) Overrepresented sequences
Overrepresented sequences
| Sequence | Count | Percentage | Possible Source |
|---|---|---|---|
| NNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNN | 118065 | 0.42042568380911904 | No Hit |
![[FAIL]](Icons/error.png) Kmer Content
Kmer Content
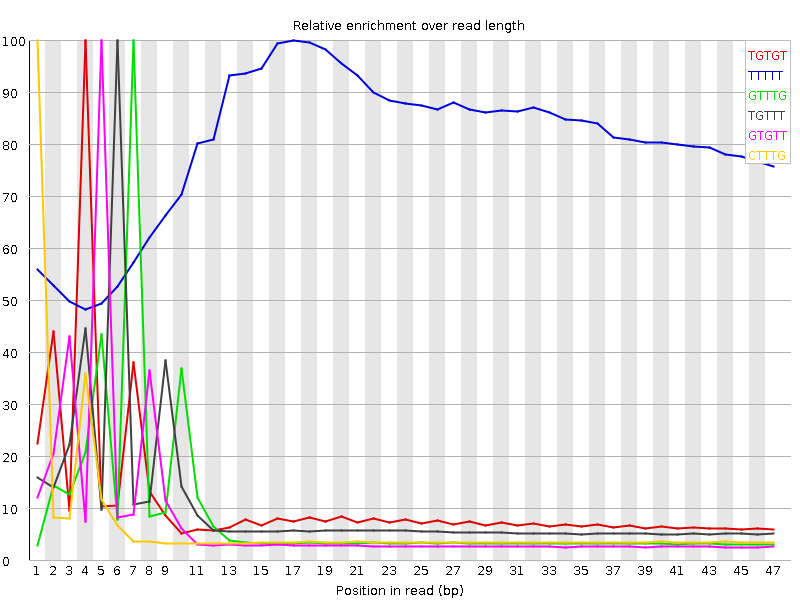
| Sequence | Count | Obs/Exp Overall | Obs/Exp Max | Max Obs/Exp Position |
|---|---|---|---|---|
| TGTGT | 6169850 | 3.805641 | 34.282314 | 4 |
| TTTTT | 9397180 | 3.318474 | 4.1365075 | 17 |
| GTTTG | 4597140 | 2.8355737 | 33.77171 | 7 |
| TGTTT | 5827800 | 2.719892 | 26.143421 | 6 |
| GTGTT | 4179170 | 2.5777645 | 33.651077 | 5 |
| CTTTG | 3696285 | 2.4499464 | 36.277718 | 1 |
| GGGGG | 1714445 | 2.4411542 | 5.4707627 | 13 |
| TTTGA | 4981405 | 2.440078 | 27.02467 | 8 |
| TTGTG | 3827690 | 2.3609674 | 33.629784 | 3 |
| GAGGG | 2028545 | 2.2937968 | 11.041254 | 11 |
| TTTGT | 4434715 | 2.0697255 | 25.761124 | 2 |
| TTGAG | 3192635 | 2.0668423 | 21.909908 | 9 |
| TGAGG | 2302285 | 1.969805 | 13.296688 | 10 |
| AGGGG | 1500845 | 1.6970949 | 6.014607 | 12 |
| AACTT | 2838680 | 1.5682344 | 11.498097 | 2 |
| TGAGC | 1629540 | 1.4981908 | 5.8539505 | 10 |
| ACTTT | 2747660 | 1.4462814 | 10.49847 | 3 |
| TGAGT | 1936870 | 1.253887 | 6.6325235 | 10 |
| GAGTG | 1437935 | 1.2302784 | 5.427689 | 11 |
| TAACT | 1980405 | 1.0940787 | 11.189317 | 1 |
| TTGAT | 2214415 | 1.0847031 | 5.959948 | 9 |
| TTGAC | 1425540 | 0.9916887 | 5.4171352 | 9 |