![[OK]](Icons/tick.png) Basic Statistics
Basic Statistics
| Measure | Value |
|---|---|
| Filename | 140701_JD_C_9_ACTTGA_L003_R2_001.fastq.gz |
| File type | Conventional base calls |
| Encoding | Sanger / Illumina 1.9 |
| Total Sequences | 24153543 |
| Filtered Sequences | 0 |
| Sequence length | 51 |
| %GC | 43 |
![[OK]](Icons/tick.png) Per base sequence quality
Per base sequence quality
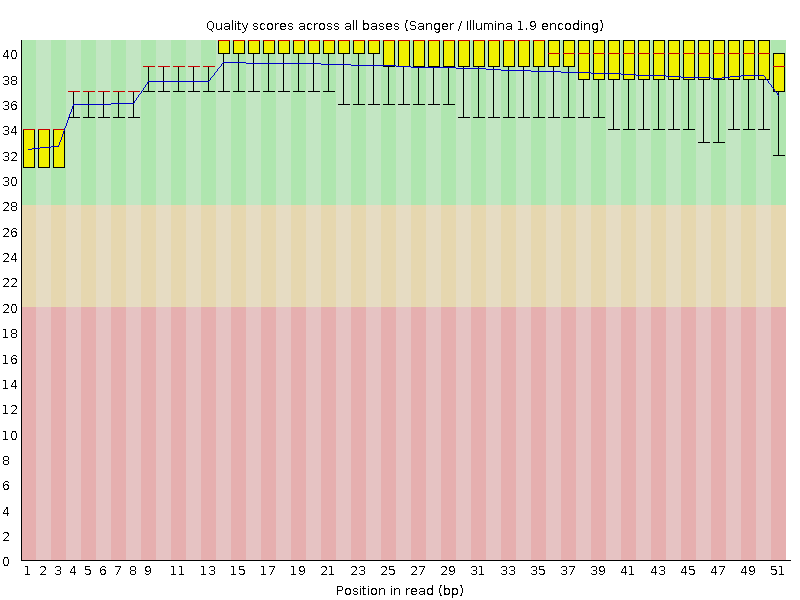
![[OK]](Icons/tick.png) Per sequence quality scores
Per sequence quality scores
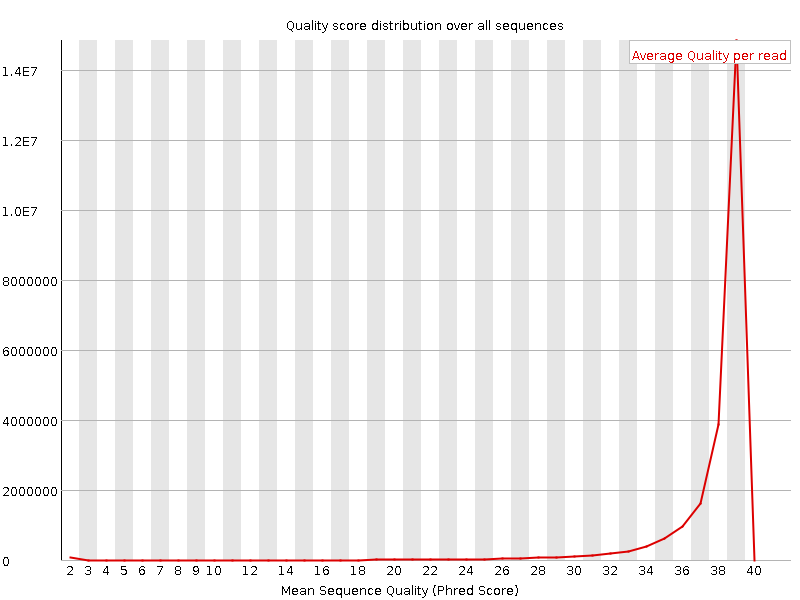
![[WARN]](Icons/warning.png) Per base sequence content
Per base sequence content

![[OK]](Icons/tick.png) Per base GC content
Per base GC content

![[OK]](Icons/tick.png) Per sequence GC content
Per sequence GC content

![[OK]](Icons/tick.png) Per base N content
Per base N content

![[OK]](Icons/tick.png) Sequence Length Distribution
Sequence Length Distribution
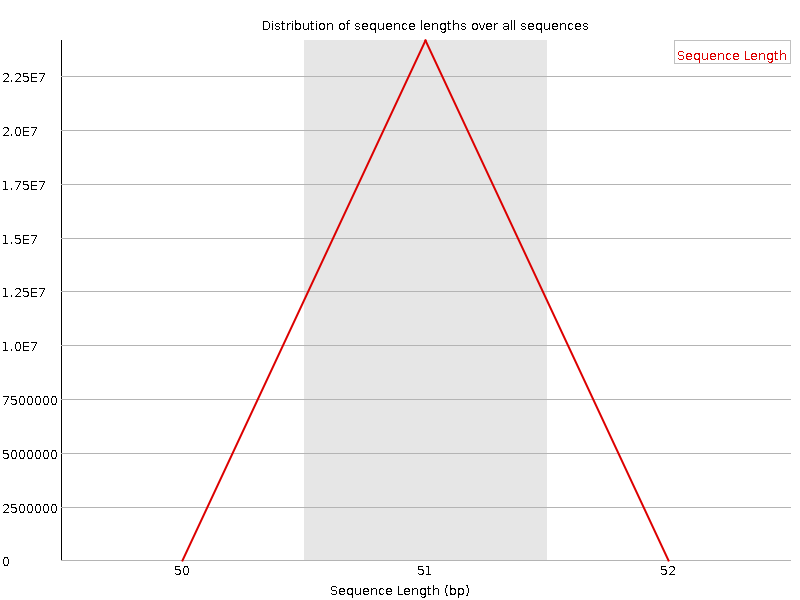
![[WARN]](Icons/warning.png) Sequence Duplication Levels
Sequence Duplication Levels
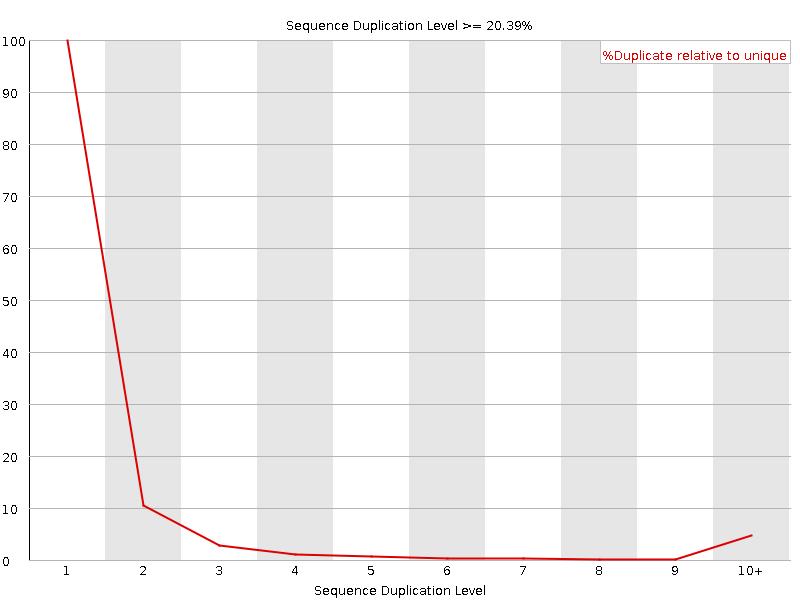
![[WARN]](Icons/warning.png) Overrepresented sequences
Overrepresented sequences
| Sequence | Count | Percentage | Possible Source |
|---|---|---|---|
| NNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNN | 106573 | 0.44123133405314496 | No Hit |
![[FAIL]](Icons/error.png) Kmer Content
Kmer Content
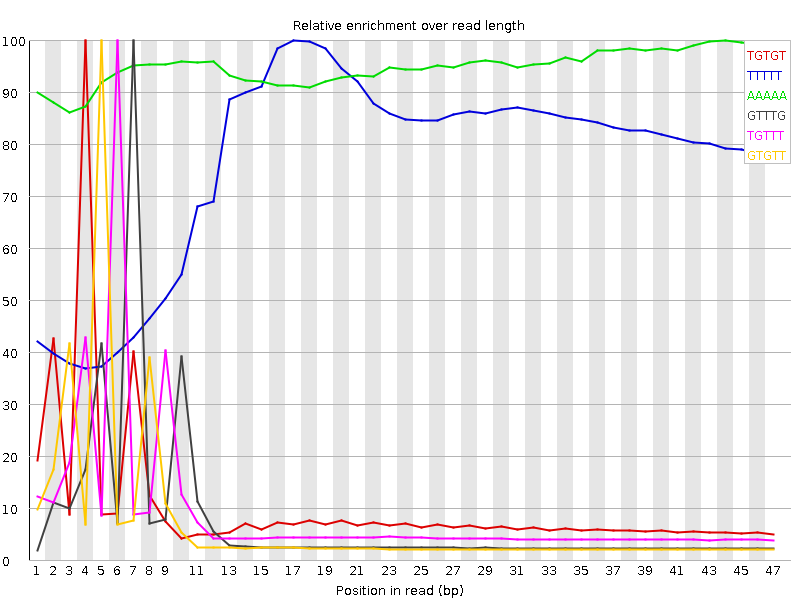
| Sequence | Count | Obs/Exp Overall | Obs/Exp Max | Max Obs/Exp Position |
|---|---|---|---|---|
| TGTGT | 6615995 | 4.547103 | 44.014675 | 4 |
| TTTTT | 9867860 | 3.977284 | 5.1981344 | 17 |
| AAAAA | 5865560 | 3.2969513 | 3.4710255 | 44 |
| GTTTG | 4686585 | 3.2210402 | 43.30671 | 7 |
| TGTTT | 5830860 | 3.068911 | 33.76014 | 6 |
| GTGTT | 4449350 | 3.0579908 | 43.30685 | 5 |
| CTTTG | 3830715 | 2.8823526 | 47.414192 | 1 |
| TTGTG | 4161715 | 2.8603027 | 43.309986 | 3 |
| TTTGA | 5011765 | 2.819228 | 35.59454 | 8 |
| GGGGG | 1644975 | 2.5174623 | 6.506168 | 13 |
| GAGGG | 1893140 | 2.3712988 | 13.600996 | 11 |
| TTTGT | 4504165 | 2.3706417 | 33.40008 | 2 |
| TTGAG | 3164980 | 2.3248692 | 27.764343 | 9 |
| GAGAG | 2193945 | 2.2492056 | 5.0451655 | 11 |
| TGAGG | 2288580 | 2.1952379 | 16.98825 | 10 |
| AGGGG | 1449970 | 1.8161955 | 7.817147 | 12 |
| ACTTT | 2496390 | 1.537374 | 14.940065 | 3 |
| TGAGC | 1450775 | 1.5235025 | 6.20474 | 10 |
| TGAGA | 1935330 | 1.519394 | 5.254938 | 10 |
| AACTT | 2286240 | 1.5047932 | 16.444496 | 2 |
| GAGTG | 1423980 | 1.3659017 | 6.8772187 | 11 |
| TGAGT | 1857440 | 1.3644018 | 8.353156 | 10 |
| TAACT | 1767540 | 1.1633871 | 16.272804 | 1 |
| TTGAT | 2022585 | 1.1377486 | 8.336813 | 9 |
| TTGAC | 1254575 | 1.0089085 | 6.6996017 | 9 |
| GTAAC | 840320 | 0.7222502 | 5.4154806 | 1 |