![[OK]](Icons/tick.png) Basic Statistics
Basic Statistics
| Measure | Value |
|---|---|
| Filename | 140701_JD_C_9_ACTTGA_L003_R1_001.fastq.gz |
| File type | Conventional base calls |
| Encoding | Sanger / Illumina 1.9 |
| Total Sequences | 24153543 |
| Filtered Sequences | 0 |
| Sequence length | 51 |
| %GC | 43 |
![[OK]](Icons/tick.png) Per base sequence quality
Per base sequence quality
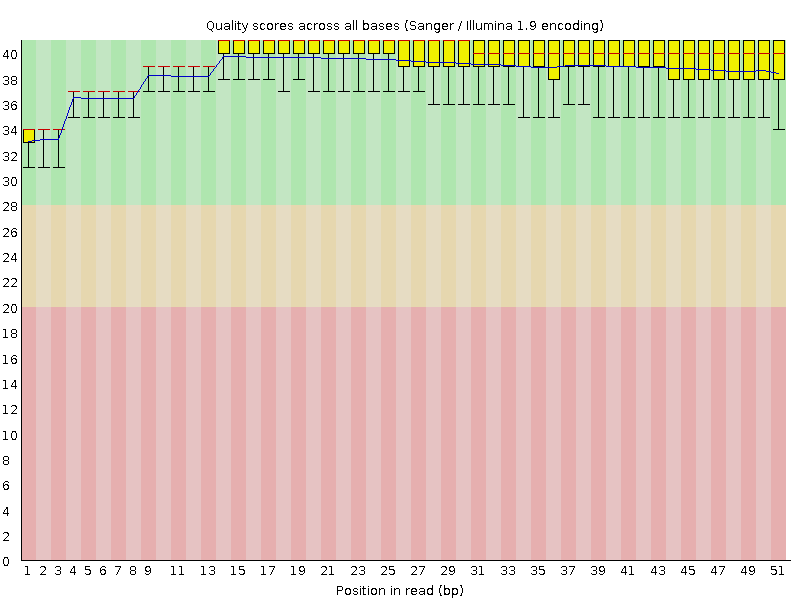
![[OK]](Icons/tick.png) Per sequence quality scores
Per sequence quality scores
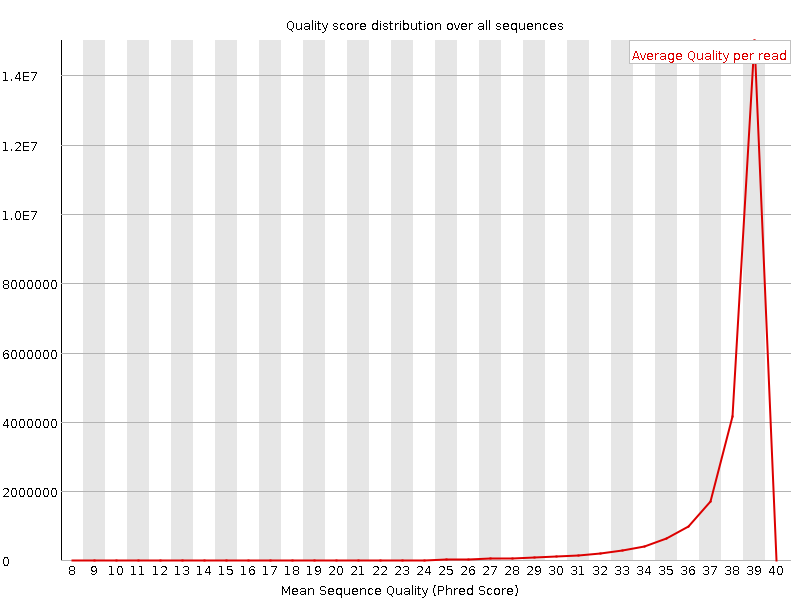
![[WARN]](Icons/warning.png) Per base sequence content
Per base sequence content

![[OK]](Icons/tick.png) Per base GC content
Per base GC content

![[OK]](Icons/tick.png) Per sequence GC content
Per sequence GC content

![[OK]](Icons/tick.png) Per base N content
Per base N content

![[OK]](Icons/tick.png) Sequence Length Distribution
Sequence Length Distribution
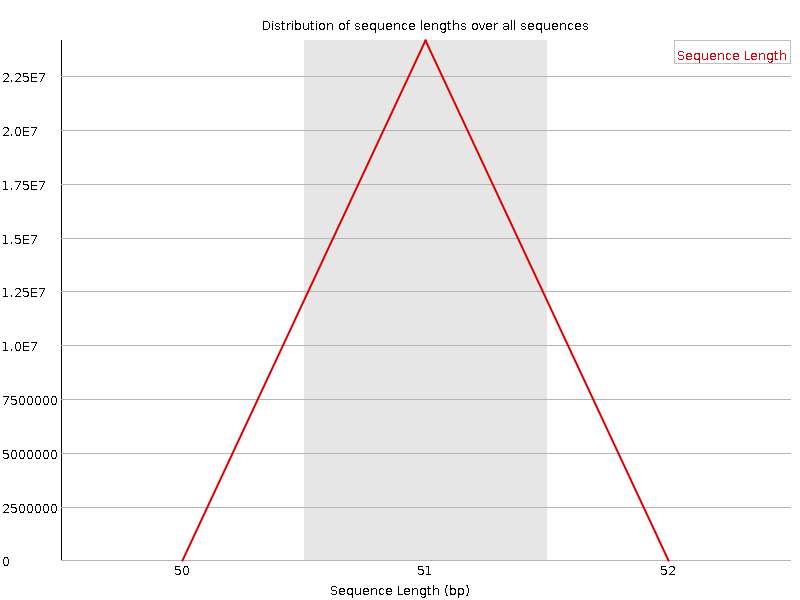
![[WARN]](Icons/warning.png) Sequence Duplication Levels
Sequence Duplication Levels
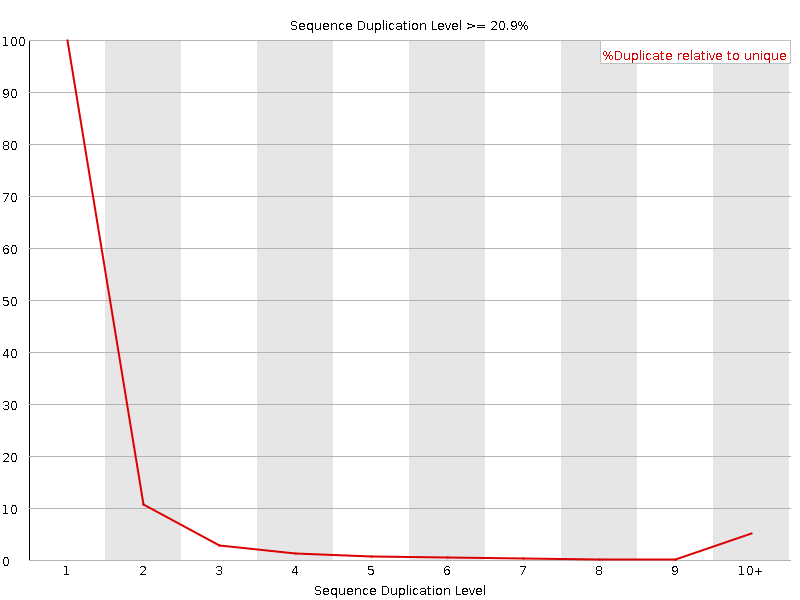
![[WARN]](Icons/warning.png) Overrepresented sequences
Overrepresented sequences
| Sequence | Count | Percentage | Possible Source |
|---|---|---|---|
| CTTTGTGTTTGATTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTT | 27739 | 0.11484443503795695 | No Hit |
![[WARN]](Icons/warning.png) Kmer Content
Kmer Content
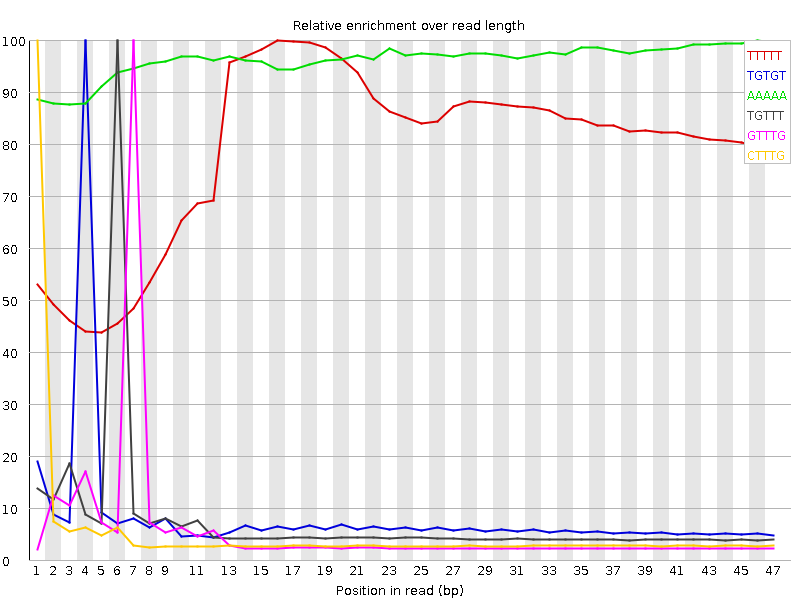
| Sequence | Count | Obs/Exp Overall | Obs/Exp Max | Max Obs/Exp Position |
|---|---|---|---|---|
| TTTTT | 9264165 | 3.8796535 | 4.9007897 | 16 |
| TGTGT | 5334975 | 3.8551264 | 46.084595 | 4 |
| AAAAA | 5846790 | 3.119271 | 3.2369685 | 46 |
| TGTTT | 4787910 | 2.6338584 | 35.095196 | 6 |
| GTTTG | 3613750 | 2.6113455 | 45.171577 | 7 |
| CTTTG | 3310140 | 2.5401418 | 48.03999 | 1 |
| GTGTT | 3365035 | 2.4316206 | 45.22509 | 5 |
| TTTGA | 3975460 | 2.2954278 | 36.28045 | 8 |
| TTGTG | 3105835 | 2.2443192 | 45.19541 | 3 |
| TTTGT | 3952200 | 2.1741292 | 34.73682 | 2 |
| GAGGG | 1655035 | 2.1660213 | 13.826559 | 11 |
| GAGAG | 2041715 | 2.135112 | 5.0488977 | 11 |
| TGAGG | 2015120 | 2.0076902 | 17.611328 | 10 |
| GGGGG | 1182810 | 1.9373161 | 5.96076 | 13 |
| TTGAG | 2551075 | 1.9349033 | 28.4499 | 9 |
| AGGGG | 1278940 | 1.6738085 | 7.9593205 | 12 |
| TGAGA | 1867590 | 1.4867821 | 5.4048166 | 10 |
| TGAGC | 1390115 | 1.4707942 | 6.142729 | 10 |
| GAGTG | 1319985 | 1.3151182 | 6.9930744 | 11 |
| TGAGT | 1707620 | 1.2951713 | 8.539096 | 10 |
| TTGAT | 1805505 | 1.0424973 | 8.705555 | 9 |
| TTGAC | 1145395 | 0.9225645 | 6.5997567 | 9 |