![[OK]](Icons/tick.png) Basic Statistics
Basic Statistics
| Measure | Value |
|---|---|
| Filename | 140701_JD_C_6_GTCCGC_L007_R1_001.fastq.gz |
| File type | Conventional base calls |
| Encoding | Sanger / Illumina 1.9 |
| Total Sequences | 41850372 |
| Filtered Sequences | 0 |
| Sequence length | 51 |
| %GC | 44 |
![[OK]](Icons/tick.png) Per base sequence quality
Per base sequence quality
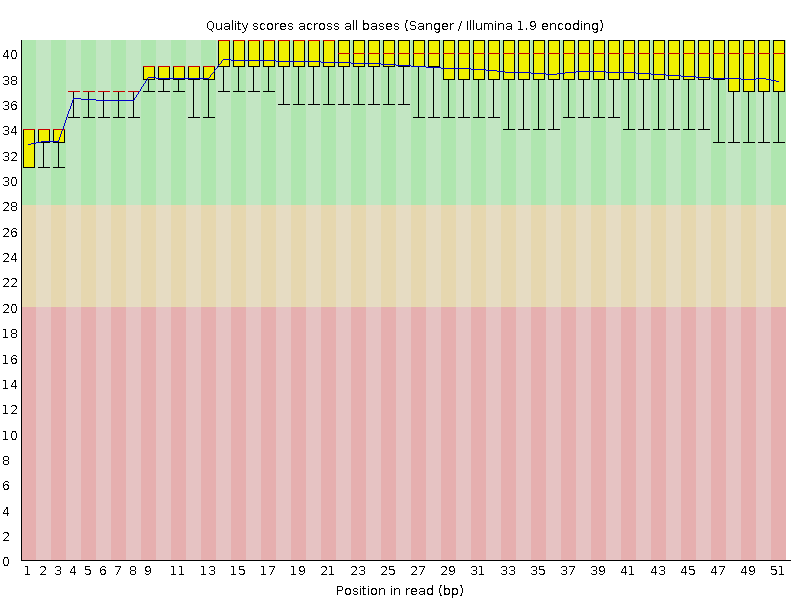
![[OK]](Icons/tick.png) Per sequence quality scores
Per sequence quality scores
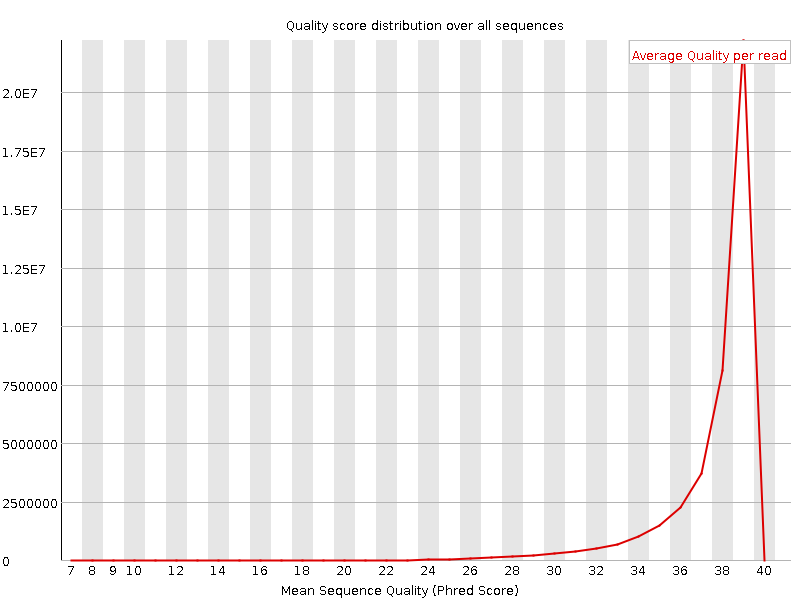
![[WARN]](Icons/warning.png) Per base sequence content
Per base sequence content

![[OK]](Icons/tick.png) Per base GC content
Per base GC content

![[OK]](Icons/tick.png) Per sequence GC content
Per sequence GC content

![[OK]](Icons/tick.png) Per base N content
Per base N content

![[OK]](Icons/tick.png) Sequence Length Distribution
Sequence Length Distribution
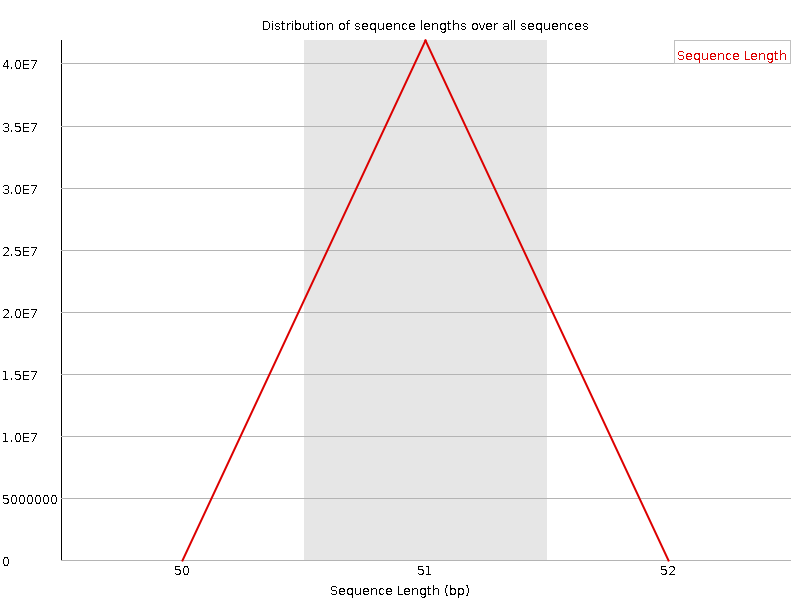
![[WARN]](Icons/warning.png) Sequence Duplication Levels
Sequence Duplication Levels
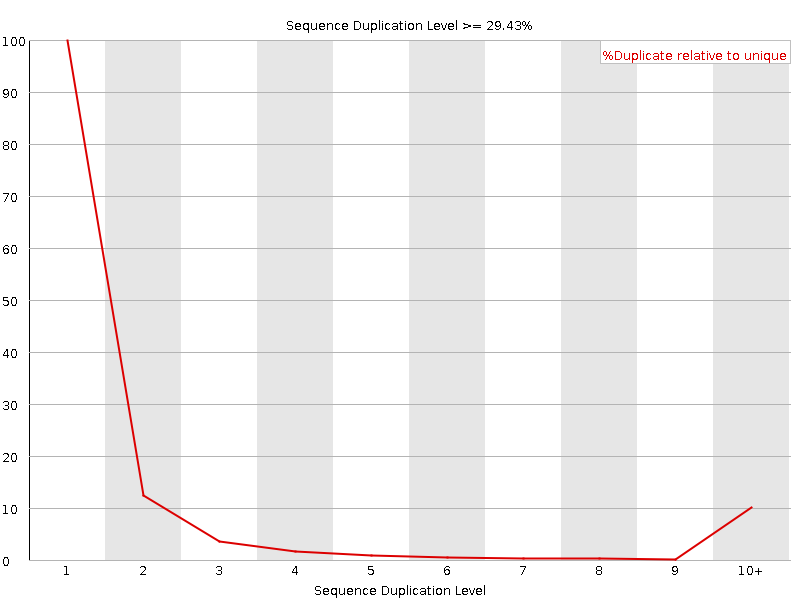
![[WARN]](Icons/warning.png) Overrepresented sequences
Overrepresented sequences
| Sequence | Count | Percentage | Possible Source |
|---|---|---|---|
| CTTTGTGTTTGATTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTT | 47046 | 0.11241477136690685 | No Hit |
![[WARN]](Icons/warning.png) Kmer Content
Kmer Content
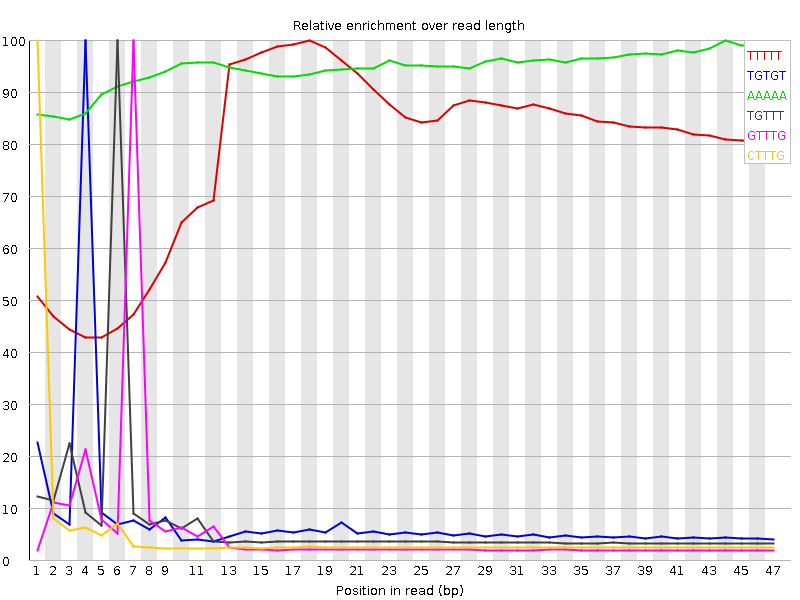
| Sequence | Count | Obs/Exp Overall | Obs/Exp Max | Max Obs/Exp Position |
|---|---|---|---|---|
| TTTTT | 14937555 | 4.024978 | 5.0911965 | 18 |
| TGTGT | 9253835 | 3.871551 | 49.941017 | 4 |
| AAAAA | 9448660 | 3.3222208 | 3.5052342 | 44 |
| TGTTT | 8372725 | 2.8111916 | 40.084923 | 6 |
| GTTTG | 6634605 | 2.7757368 | 49.224648 | 7 |
| CTTTG | 5955100 | 2.6462073 | 52.18934 | 1 |
| GTGTT | 6219880 | 2.602227 | 49.22366 | 5 |
| TTTGA | 7288115 | 2.5807958 | 41.906406 | 8 |
| TTGTG | 5535170 | 2.3157635 | 49.203903 | 3 |
| TTTGT | 6833435 | 2.2943656 | 39.689518 | 2 |
| GAGGG | 3126145 | 2.141738 | 13.117005 | 11 |
| GGGGG | 2572910 | 2.0826035 | 5.2831845 | 13 |
| TTGAG | 4534990 | 2.0010338 | 29.940855 | 9 |
| TGAGG | 3492775 | 1.9203848 | 17.635654 | 10 |
| AGGGG | 2268220 | 1.5539691 | 7.1775756 | 12 |
| TGAGC | 2493530 | 1.4561424 | 6.6169405 | 10 |
| TGAGA | 3118055 | 1.4510309 | 5.5482755 | 10 |
| TGAGT | 2862945 | 1.2632551 | 8.5103445 | 10 |
| GAGTG | 2295780 | 1.2622573 | 6.5510173 | 11 |
| TTGAT | 3188375 | 1.1290362 | 9.583297 | 9 |
| TTGAC | 2121275 | 0.994138 | 8.783105 | 9 |