![[OK]](Icons/tick.png) Basic Statistics
Basic Statistics
| Measure | Value |
|---|---|
| Filename | 140701_JD_C_5_CTTGTA_L002_R1_001.fastq.gz |
| File type | Conventional base calls |
| Encoding | Sanger / Illumina 1.9 |
| Total Sequences | 25970033 |
| Filtered Sequences | 0 |
| Sequence length | 51 |
| %GC | 44 |
![[OK]](Icons/tick.png) Per base sequence quality
Per base sequence quality
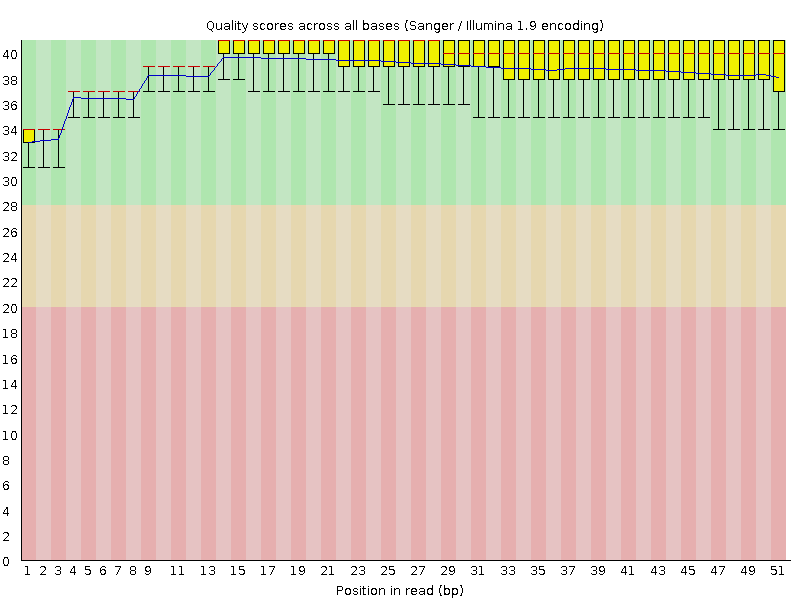
![[OK]](Icons/tick.png) Per sequence quality scores
Per sequence quality scores
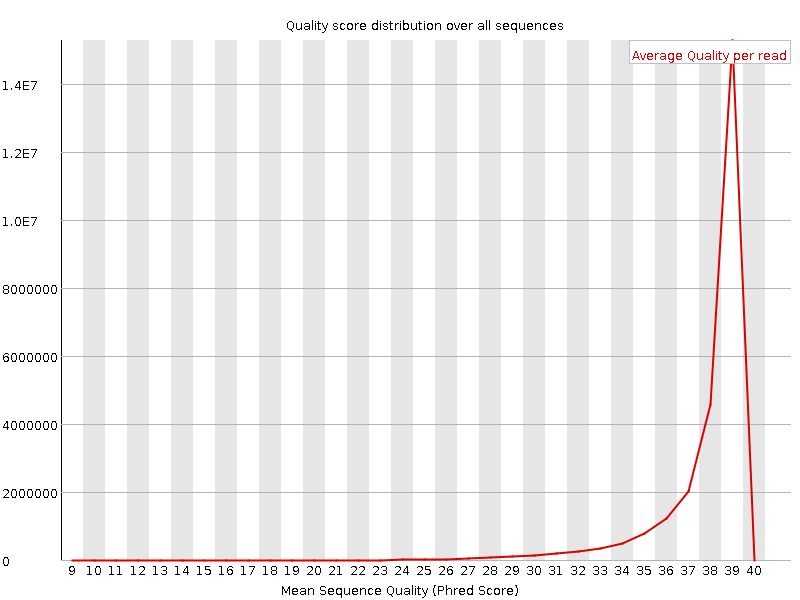
![[WARN]](Icons/warning.png) Per base sequence content
Per base sequence content

![[OK]](Icons/tick.png) Per base GC content
Per base GC content

![[OK]](Icons/tick.png) Per sequence GC content
Per sequence GC content

![[OK]](Icons/tick.png) Per base N content
Per base N content

![[OK]](Icons/tick.png) Sequence Length Distribution
Sequence Length Distribution
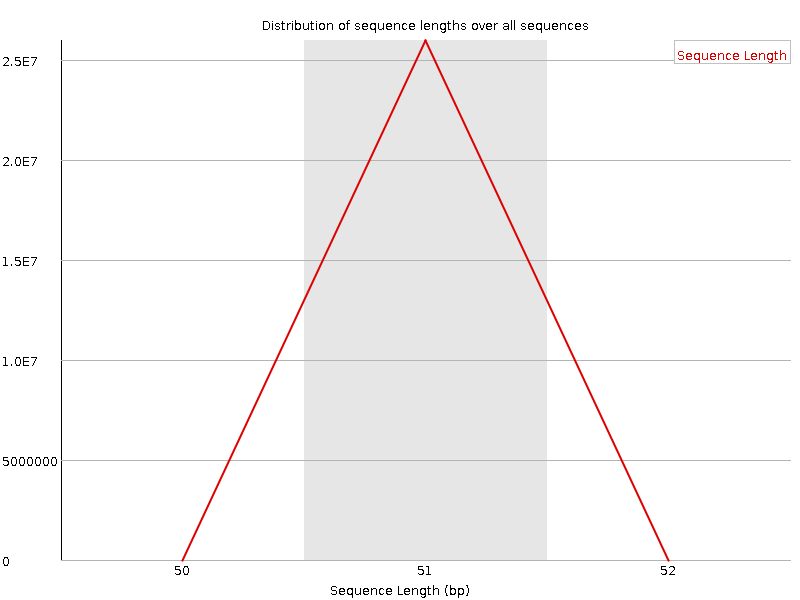
![[WARN]](Icons/warning.png) Sequence Duplication Levels
Sequence Duplication Levels
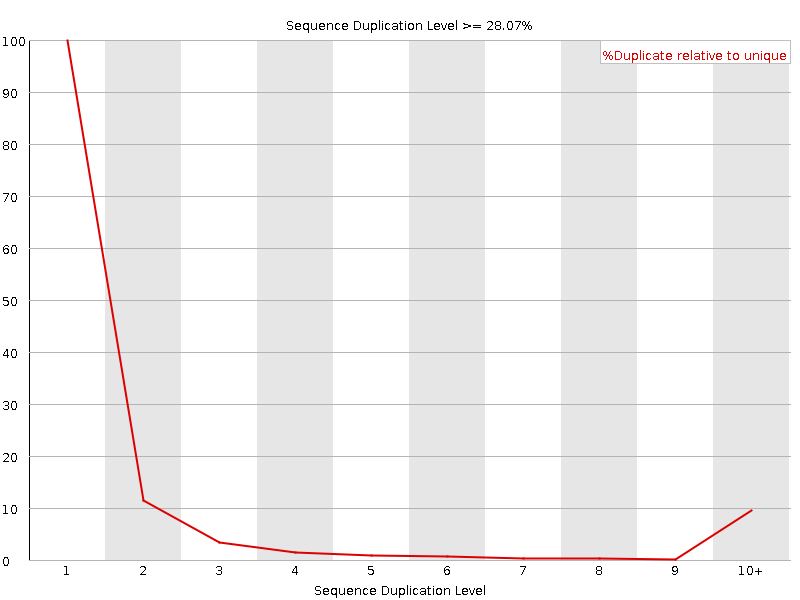
![[WARN]](Icons/warning.png) Overrepresented sequences
Overrepresented sequences
| Sequence | Count | Percentage | Possible Source |
|---|---|---|---|
| CTTTGTGTTTGATTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTT | 42372 | 0.16315728208739666 | No Hit |
![[WARN]](Icons/warning.png) Kmer Content
Kmer Content
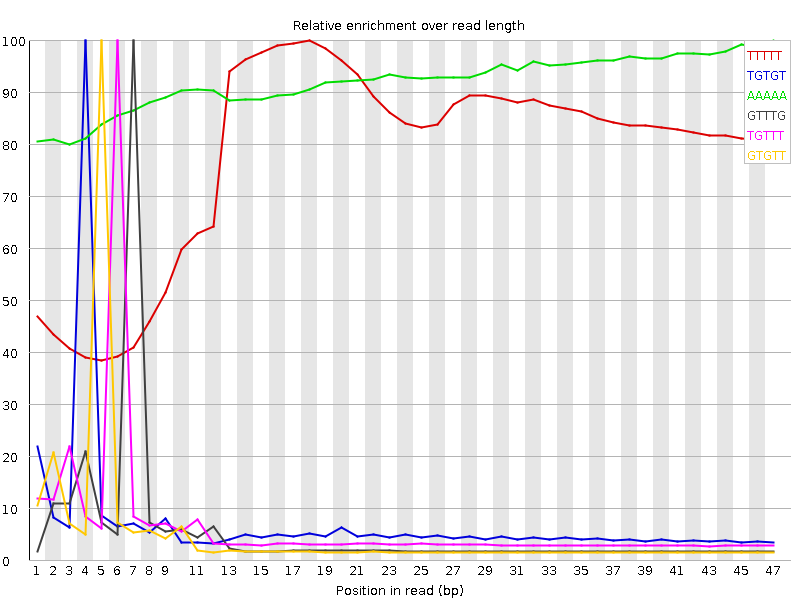
| Sequence | Count | Obs/Exp Overall | Obs/Exp Max | Max Obs/Exp Position |
|---|---|---|---|---|
| TTTTT | 11513670 | 4.6887984 | 6.0177817 | 18 |
| TGTGT | 6253920 | 4.081467 | 56.709526 | 4 |
| AAAAA | 6421425 | 3.6359248 | 3.9467196 | 47 |
| GTTTG | 4646250 | 3.0322611 | 56.07625 | 7 |
| TGTTT | 5700115 | 2.9385939 | 44.831554 | 6 |
| GTGTT | 4307190 | 2.8109818 | 56.012276 | 5 |
| CTTTG | 4005585 | 2.807797 | 59.85662 | 1 |
| TTTGA | 5006880 | 2.7570853 | 47.532593 | 8 |
| TTGTG | 3800795 | 2.4804952 | 55.96377 | 3 |
| TTTGT | 4608940 | 2.3760576 | 44.374184 | 2 |
| GAGGG | 2003365 | 2.2380378 | 15.26909 | 11 |
| TTGAG | 3062570 | 2.134898 | 34.327843 | 9 |
| GGGGG | 1610585 | 2.1324162 | 5.94606 | 13 |
| GAGAG | 2237740 | 2.1092887 | 5.472687 | 11 |
| TGAGG | 2278415 | 2.0106297 | 20.233246 | 10 |
| AGGGG | 1439320 | 1.6079209 | 8.3236065 | 12 |
| TGAGC | 1547125 | 1.4664266 | 7.600314 | 10 |
| TGAGA | 1948410 | 1.4507703 | 6.3999257 | 10 |
| TGAGT | 1835665 | 1.2796304 | 9.702513 | 10 |
| GAGTG | 1443510 | 1.2738522 | 7.464317 | 11 |
| TGATG | 1721650 | 1.2001513 | 5.1136723 | 10 |
| TTGAT | 2130060 | 1.1729374 | 11.256774 | 9 |
| TTGAC | 1395225 | 1.0446506 | 9.863699 | 9 |
| TGATT | 1775980 | 0.97796005 | 5.4219375 | 10 |