![[OK]](Icons/tick.png) Basic Statistics
Basic Statistics
| Measure | Value |
|---|---|
| Filename | 140701_JD_C_2_GATCAG_L006_R2_001.fastq.gz |
| File type | Conventional base calls |
| Encoding | Sanger / Illumina 1.9 |
| Total Sequences | 34096341 |
| Filtered Sequences | 0 |
| Sequence length | 51 |
| %GC | 44 |
![[OK]](Icons/tick.png) Per base sequence quality
Per base sequence quality
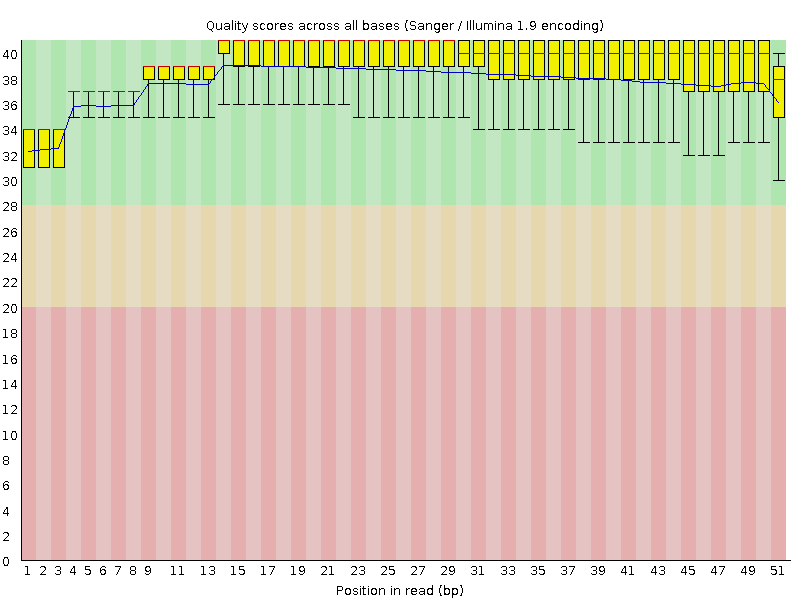
![[OK]](Icons/tick.png) Per sequence quality scores
Per sequence quality scores
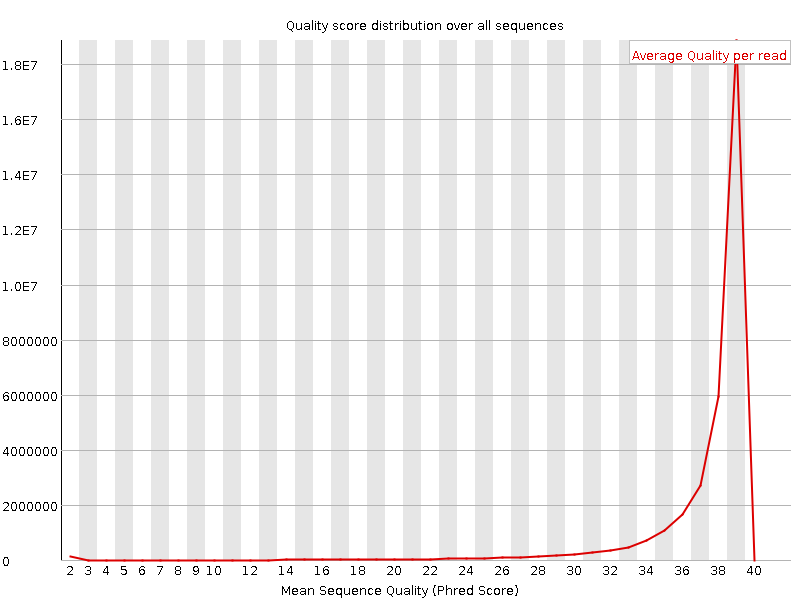
![[WARN]](Icons/warning.png) Per base sequence content
Per base sequence content

![[OK]](Icons/tick.png) Per base GC content
Per base GC content

![[OK]](Icons/tick.png) Per sequence GC content
Per sequence GC content

![[OK]](Icons/tick.png) Per base N content
Per base N content

![[OK]](Icons/tick.png) Sequence Length Distribution
Sequence Length Distribution
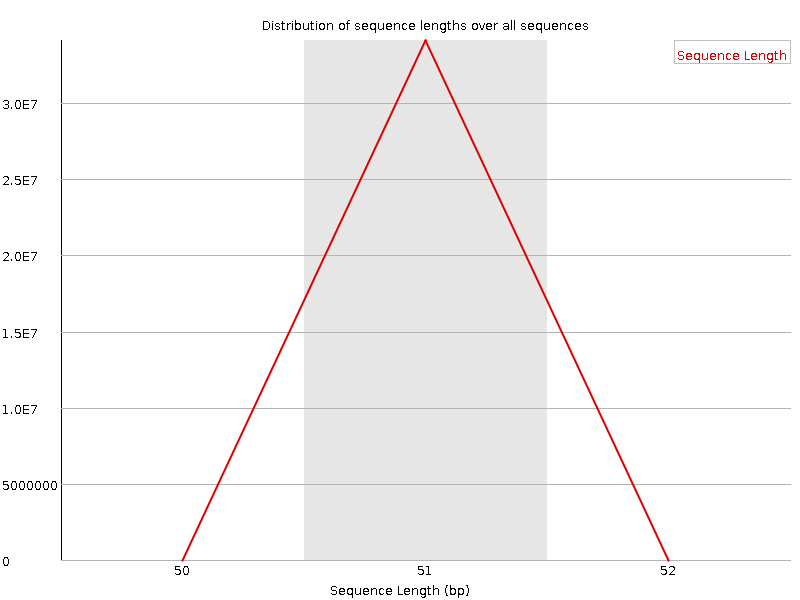
![[WARN]](Icons/warning.png) Sequence Duplication Levels
Sequence Duplication Levels
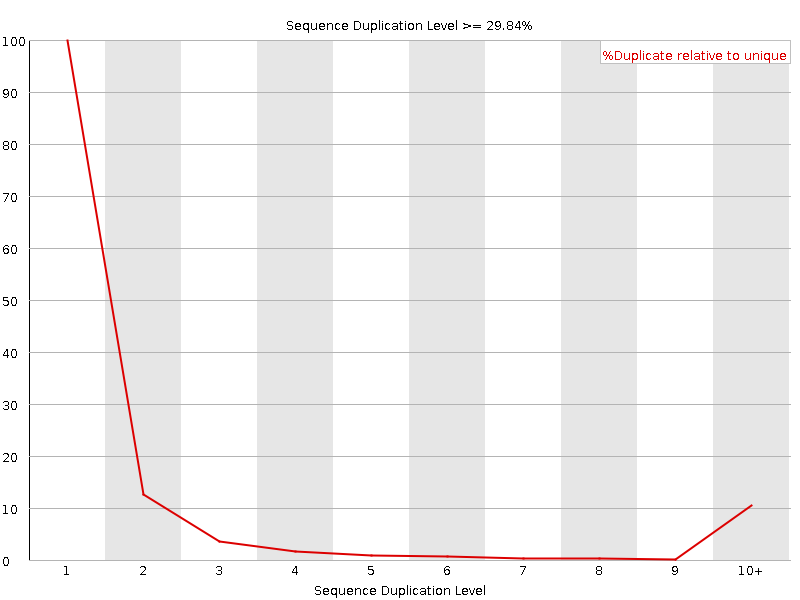
![[WARN]](Icons/warning.png) Overrepresented sequences
Overrepresented sequences
| Sequence | Count | Percentage | Possible Source |
|---|---|---|---|
| NNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNN | 142912 | 0.4191417489636205 | No Hit |
![[FAIL]](Icons/error.png) Kmer Content
Kmer Content
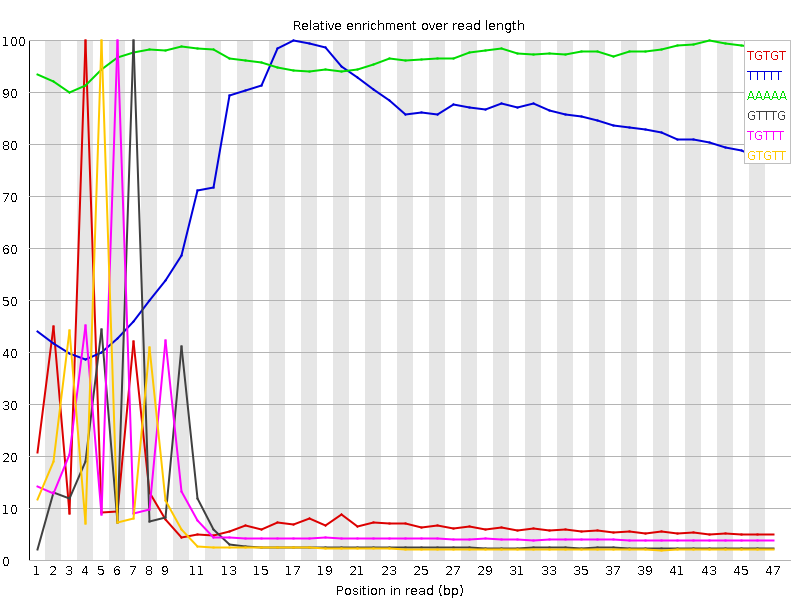
| Sequence | Count | Obs/Exp Overall | Obs/Exp Max | Max Obs/Exp Position |
|---|---|---|---|---|
| TGTGT | 8600355 | 4.308103 | 41.310043 | 4 |
| TTTTT | 12504950 | 4.162648 | 5.3621373 | 17 |
| AAAAA | 7524890 | 3.4427872 | 3.55675 | 43 |
| GTTTG | 6269125 | 3.1403399 | 40.717903 | 7 |
| TGTTT | 7649575 | 3.1236768 | 33.739418 | 6 |
| GTGTT | 5927300 | 2.9691122 | 40.670216 | 5 |
| TTTGA | 6767520 | 2.9449825 | 35.582283 | 8 |
| CTTTG | 5130455 | 2.780785 | 44.17829 | 1 |
| TTGTG | 5460475 | 2.7352695 | 40.696434 | 3 |
| GGGGG | 2734265 | 2.5283265 | 5.33458 | 13 |
| TTTGT | 5857275 | 2.391797 | 33.434593 | 2 |
| TTGAG | 4230800 | 2.258482 | 25.628786 | 9 |
| GAGGG | 2780080 | 2.2332294 | 11.143549 | 11 |
| TGAGG | 3076545 | 2.0146444 | 14.718448 | 10 |
| ACTTT | 3492210 | 1.6443539 | 15.515175 | 3 |
| AGGGG | 2044150 | 1.6420591 | 6.2777743 | 12 |
| AACTT | 3171195 | 1.5912645 | 17.049292 | 2 |
| TGAGC | 2094900 | 1.4843634 | 5.724689 | 10 |
| TGAGT | 2457130 | 1.311663 | 7.505667 | 10 |
| GAGTG | 1971110 | 1.2907612 | 5.7695446 | 11 |
| TAACT | 2467090 | 1.2379537 | 16.891928 | 1 |
| TTGAT | 2743610 | 1.1939209 | 7.9215827 | 9 |
| TTGAC | 1796840 | 1.0378757 | 7.370726 | 9 |
| GTAAC | 1193715 | 0.7347864 | 5.318273 | 1 |