![[OK]](Icons/tick.png) Basic Statistics
Basic Statistics
| Measure | Value |
|---|---|
| Filename | 140701_JD_C_1_ACAGTG_L002_R1_001.fastq.gz |
| File type | Conventional base calls |
| Encoding | Sanger / Illumina 1.9 |
| Total Sequences | 35774422 |
| Filtered Sequences | 0 |
| Sequence length | 51 |
| %GC | 44 |
![[OK]](Icons/tick.png) Per base sequence quality
Per base sequence quality
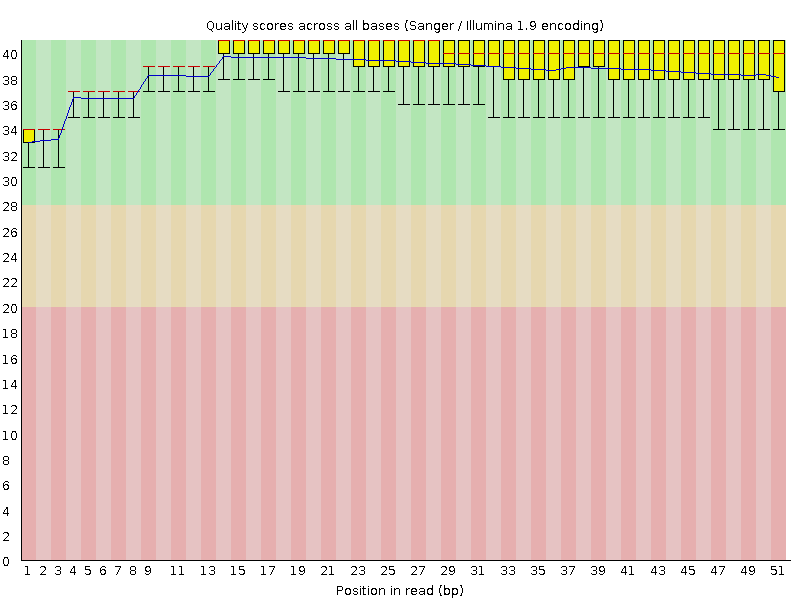
![[OK]](Icons/tick.png) Per sequence quality scores
Per sequence quality scores
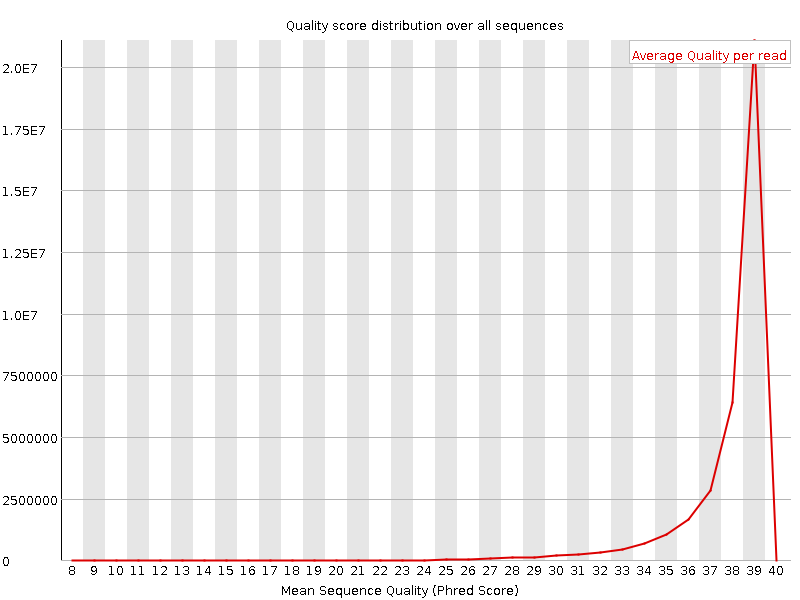
![[OK]](Icons/tick.png) Per base sequence content
Per base sequence content

![[OK]](Icons/tick.png) Per base GC content
Per base GC content

![[OK]](Icons/tick.png) Per sequence GC content
Per sequence GC content

![[OK]](Icons/tick.png) Per base N content
Per base N content

![[OK]](Icons/tick.png) Sequence Length Distribution
Sequence Length Distribution
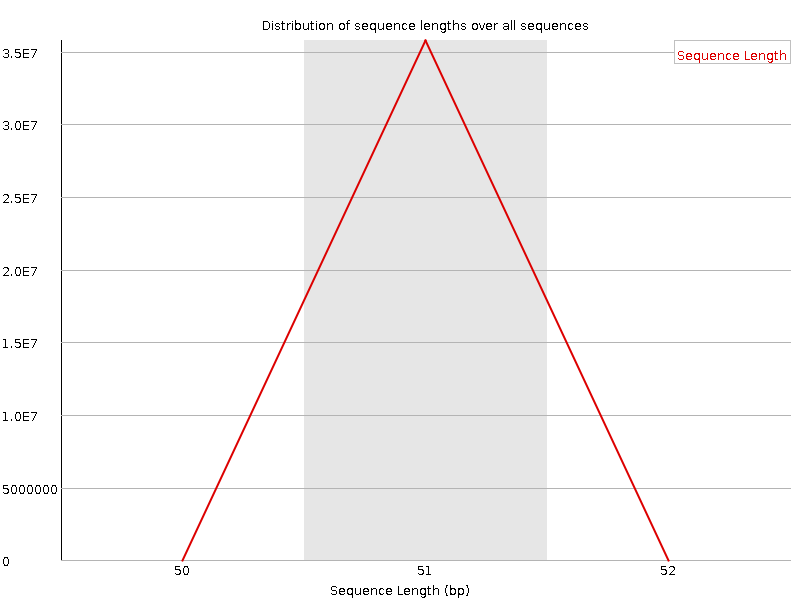
![[WARN]](Icons/warning.png) Sequence Duplication Levels
Sequence Duplication Levels
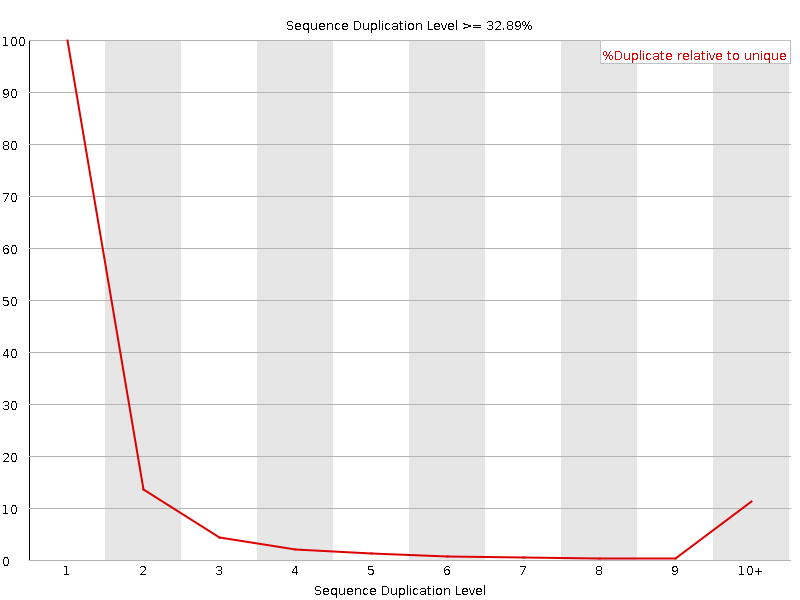
![[OK]](Icons/tick.png) Overrepresented sequences
Overrepresented sequences
No overrepresented sequences
![[WARN]](Icons/warning.png) Kmer Content
Kmer Content
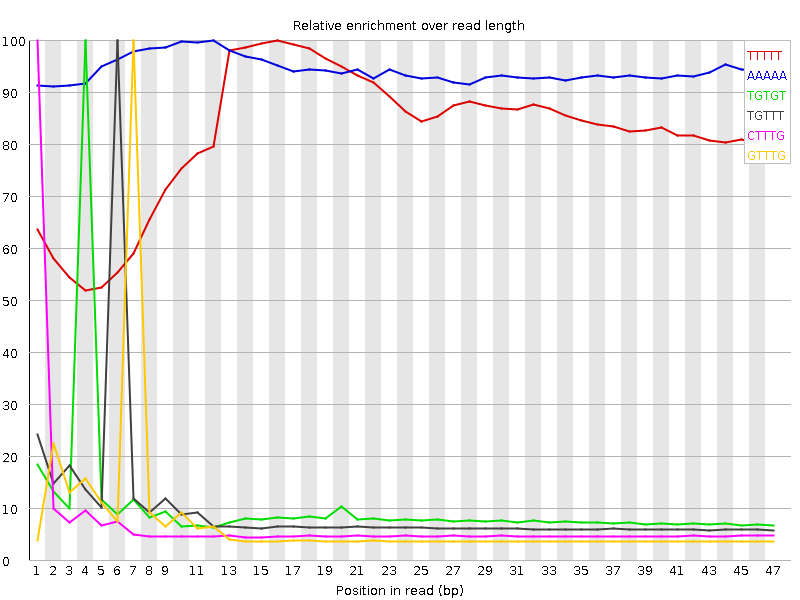
| Sequence | Count | Obs/Exp Overall | Obs/Exp Max | Max Obs/Exp Position |
|---|---|---|---|---|
| TTTTT | 11073845 | 3.8389113 | 4.6886115 | 16 |
| AAAAA | 7589540 | 3.071283 | 3.2537317 | 12 |
| TGTGT | 5722940 | 2.9767087 | 29.06864 | 4 |
| TGTTT | 5463930 | 2.3201635 | 23.813234 | 6 |
| CTTTG | 3910235 | 2.1078963 | 29.360735 | 1 |
| GTTTG | 4004705 | 2.0829926 | 28.331743 | 7 |
| TTTGA | 4634920 | 2.0299933 | 24.181534 | 8 |
| TTTGT | 4552500 | 1.9331404 | 23.444473 | 2 |
| GAGGG | 2375105 | 1.9118173 | 7.5280476 | 11 |
| GTGTT | 3634645 | 1.8905108 | 28.311556 | 5 |
| TTGTG | 3405980 | 1.7715739 | 28.302185 | 3 |
| TGAGG | 2547160 | 1.673848 | 9.953265 | 10 |
| TTGAG | 2934925 | 1.5745379 | 16.843313 | 9 |
| TGAGT | 2169285 | 1.1637849 | 5.102827 | 10 |
| TTGAT | 2409625 | 1.0553629 | 5.8665905 | 9 |
| TTGAC | 1588055 | 0.8829801 | 5.217125 | 9 |