![[OK]](Icons/tick.png) Basic Statistics
Basic Statistics
| Measure | Value |
|---|---|
| Filename | 140701_JD_C_14_TGACCA_L002_R2_001.fastq.gz |
| File type | Conventional base calls |
| Encoding | Sanger / Illumina 1.9 |
| Total Sequences | 20892507 |
| Filtered Sequences | 0 |
| Sequence length | 51 |
| %GC | 46 |
![[OK]](Icons/tick.png) Per base sequence quality
Per base sequence quality
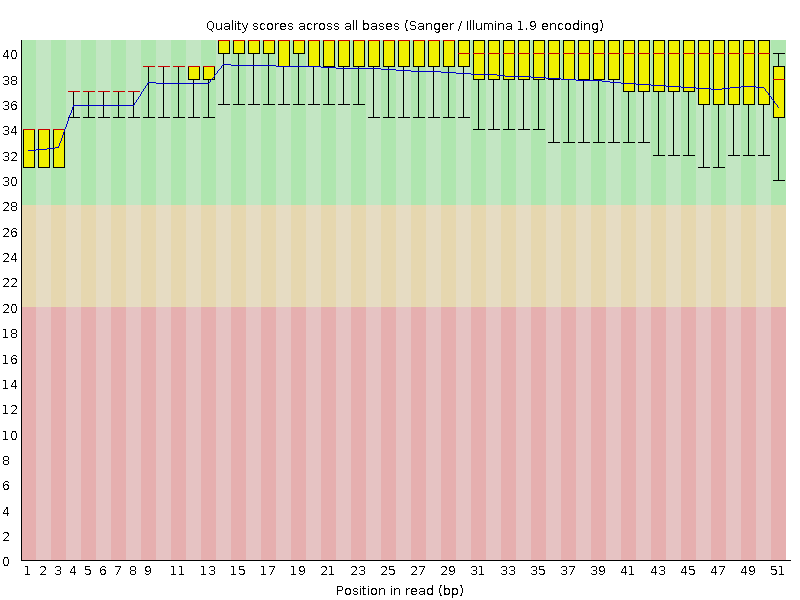
![[OK]](Icons/tick.png) Per sequence quality scores
Per sequence quality scores
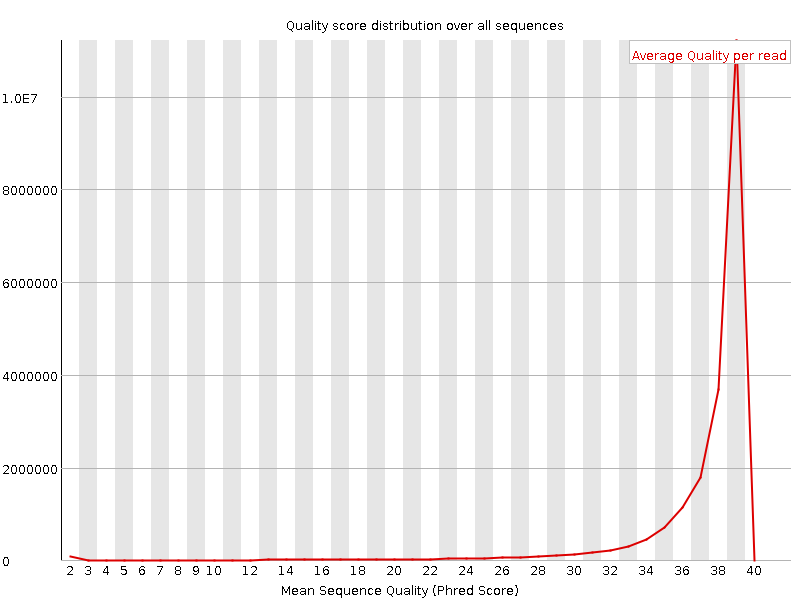
![[WARN]](Icons/warning.png) Per base sequence content
Per base sequence content

![[WARN]](Icons/warning.png) Per base GC content
Per base GC content

![[OK]](Icons/tick.png) Per sequence GC content
Per sequence GC content

![[OK]](Icons/tick.png) Per base N content
Per base N content

![[OK]](Icons/tick.png) Sequence Length Distribution
Sequence Length Distribution
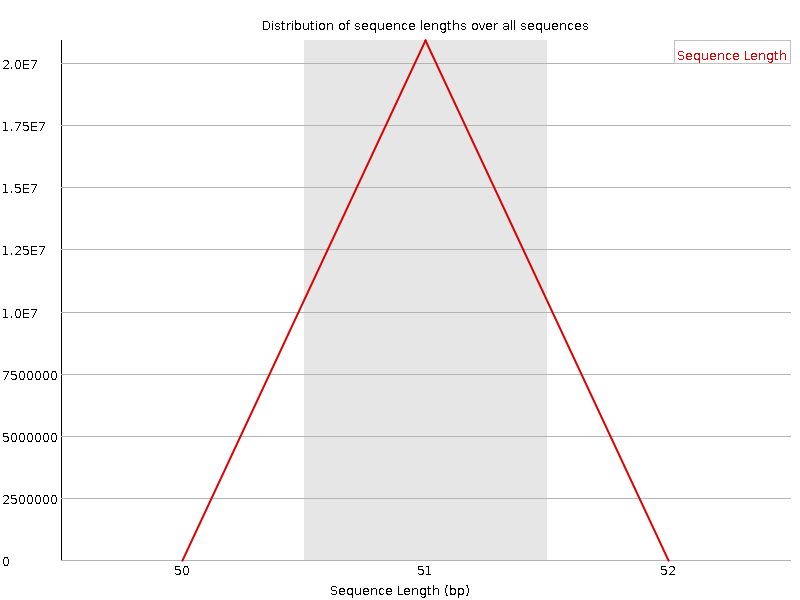
![[WARN]](Icons/warning.png) Sequence Duplication Levels
Sequence Duplication Levels
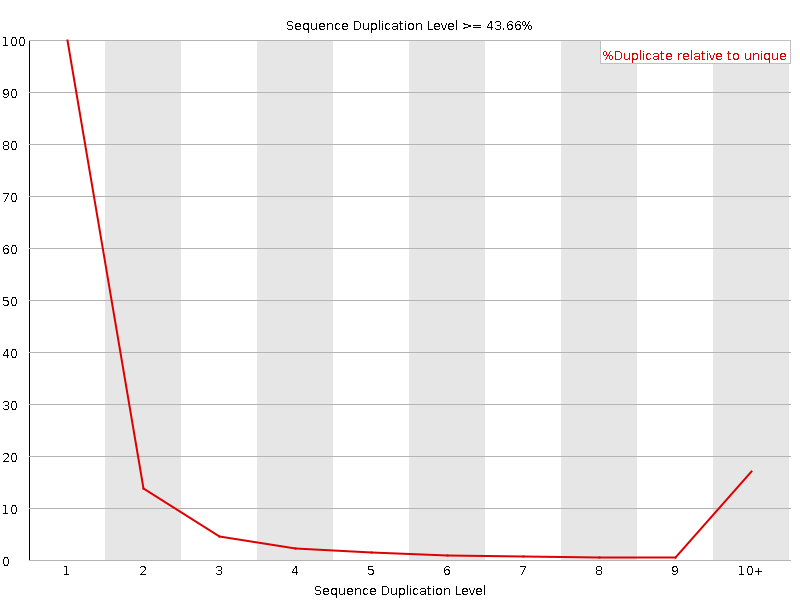
![[WARN]](Icons/warning.png) Overrepresented sequences
Overrepresented sequences
| Sequence | Count | Percentage | Possible Source |
|---|---|---|---|
| NNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNN | 96899 | 0.46379785824649955 | No Hit |
| CTTTGTGTTTGATTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTT | 30124 | 0.14418566426709825 | No Hit |
| CTTTGTGTTTGACGGGCGGTGTGTACAAAGGGCAGGGACTTAATCAACGCA | 24958 | 0.11945909602902131 | No Hit |
| TTGTGTTTGATTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTT | 23717 | 0.11351916742208103 | No Hit |
![[WARN]](Icons/warning.png) Kmer Content
Kmer Content
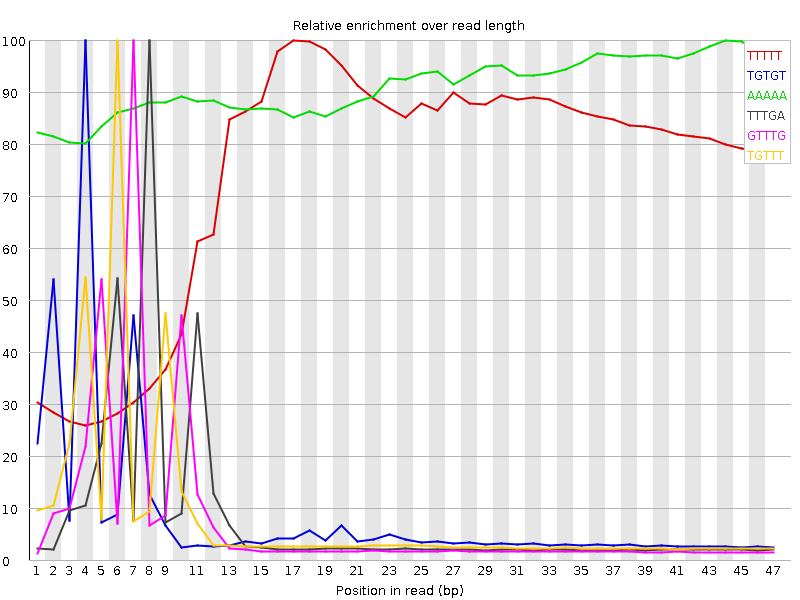
| Sequence | Count | Obs/Exp Overall | Obs/Exp Max | Max Obs/Exp Position |
|---|---|---|---|---|
| TTTTT | 10092440 | 5.789939 | 7.79181 | 17 |
| TGTGT | 5554375 | 4.341171 | 51.020653 | 4 |
| AAAAA | 4370175 | 4.137378 | 4.5329175 | 44 |
| TTTGA | 5119960 | 3.7896664 | 48.34736 | 8 |
| GTTTG | 4835425 | 3.779256 | 50.819218 | 7 |
| TGTTT | 5383990 | 3.605197 | 43.906487 | 6 |
| GTGTT | 4570805 | 3.5724351 | 50.73223 | 5 |
| TTGTG | 4076235 | 3.1858904 | 50.70595 | 3 |
| CTTTG | 3695265 | 3.1566968 | 55.61353 | 1 |
| TTTGT | 3964555 | 2.6547227 | 43.63302 | 2 |
| TTGAG | 3005790 | 2.5968084 | 31.479382 | 9 |
| GAGGG | 1955315 | 2.3013985 | 12.320443 | 11 |
| AACTT | 2346905 | 2.0987236 | 27.842546 | 2 |
| ACTTT | 2531895 | 2.0483105 | 24.43628 | 3 |
| TGAGG | 1988105 | 2.0047839 | 17.055468 | 10 |
| TAACT | 1762640 | 1.5762438 | 27.635504 | 1 |
| AGGGG | 1271790 | 1.4968922 | 6.124639 | 12 |
| TGAGC | 1352475 | 1.4906412 | 7.3548956 | 10 |
| TTGAT | 1963380 | 1.4532448 | 10.665794 | 9 |
| TGAGA | 1411420 | 1.347868 | 5.5760875 | 10 |
| TTGAC | 1364585 | 1.2885388 | 11.285001 | 9 |
| TGAGT | 1413950 | 1.2215617 | 8.423164 | 10 |
| GAGTG | 1158430 | 1.1681484 | 5.6051216 | 11 |
| TGATT | 1518415 | 1.1238929 | 5.1043744 | 10 |
| GTAAC | 831630 | 0.8680342 | 7.9879484 | 1 |
| TGACG | 576100 | 0.63495326 | 5.374183 | 10 |