![[OK]](Icons/tick.png) Basic Statistics
Basic Statistics
| Measure | Value |
|---|---|
| Filename | 140701_JD_C_12_CAGATC_L007_R2_001.fastq.gz |
| File type | Conventional base calls |
| Encoding | Sanger / Illumina 1.9 |
| Total Sequences | 34775151 |
| Filtered Sequences | 0 |
| Sequence length | 51 |
| %GC | 44 |
![[OK]](Icons/tick.png) Per base sequence quality
Per base sequence quality
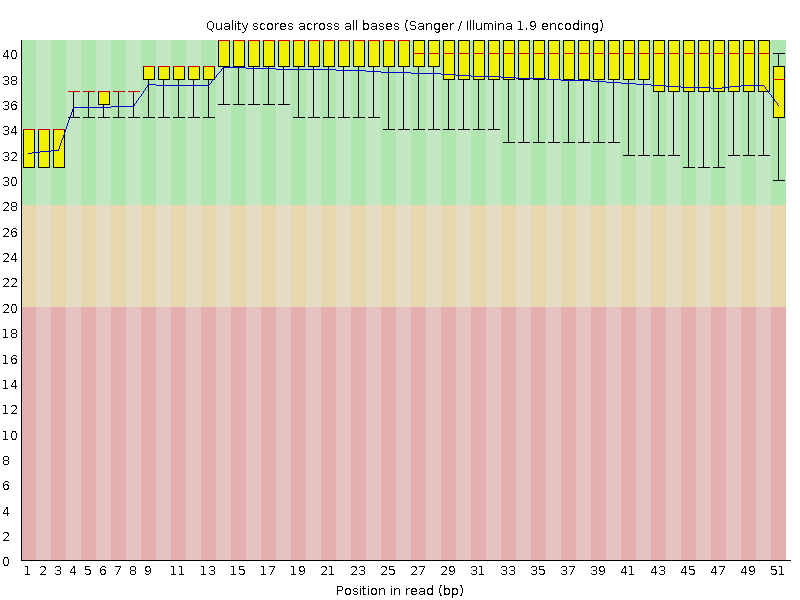
![[OK]](Icons/tick.png) Per sequence quality scores
Per sequence quality scores
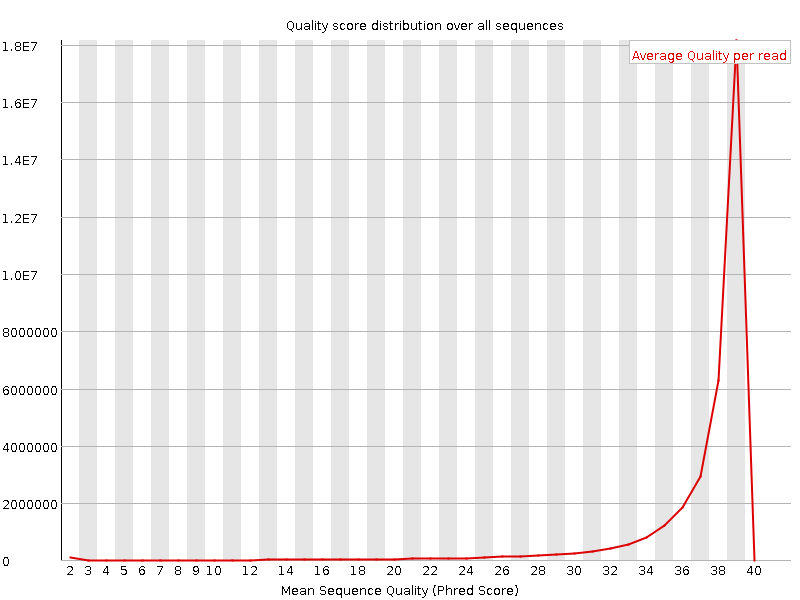
![[WARN]](Icons/warning.png) Per base sequence content
Per base sequence content

![[WARN]](Icons/warning.png) Per base GC content
Per base GC content

![[OK]](Icons/tick.png) Per sequence GC content
Per sequence GC content

![[OK]](Icons/tick.png) Per base N content
Per base N content

![[OK]](Icons/tick.png) Sequence Length Distribution
Sequence Length Distribution
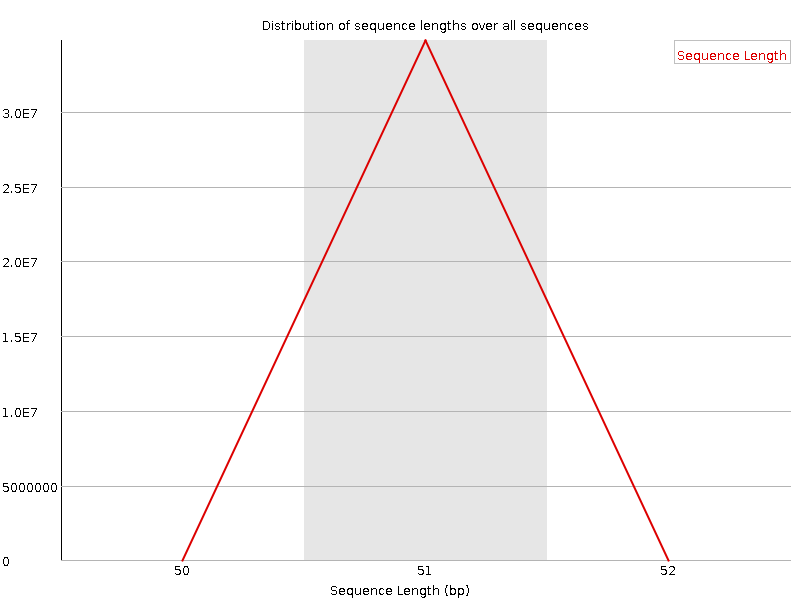
![[WARN]](Icons/warning.png) Sequence Duplication Levels
Sequence Duplication Levels
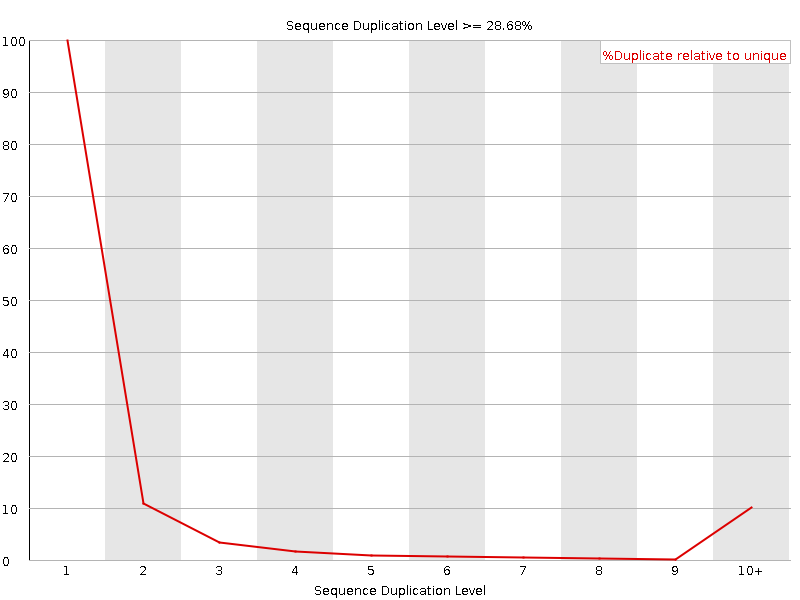
![[OK]](Icons/tick.png) Overrepresented sequences
Overrepresented sequences
No overrepresented sequences
![[FAIL]](Icons/error.png) Kmer Content
Kmer Content
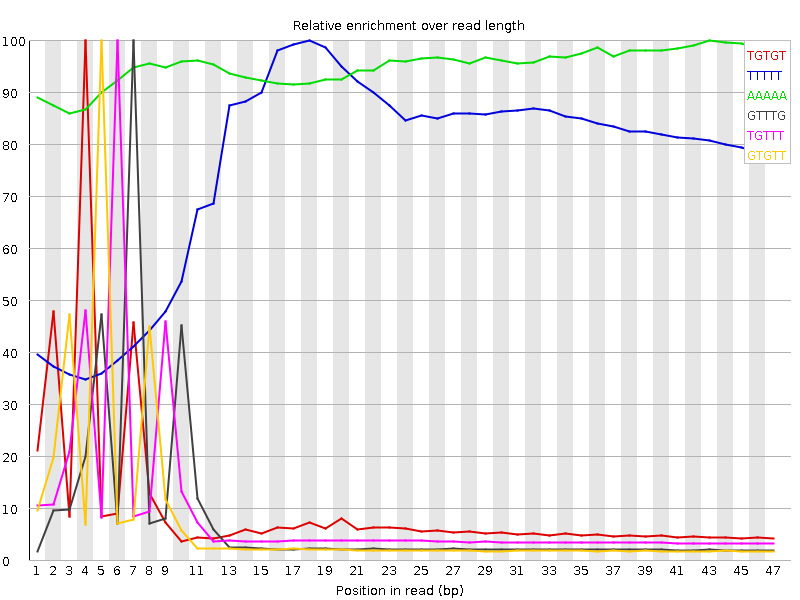
| Sequence | Count | Obs/Exp Overall | Obs/Exp Max | Max Obs/Exp Position |
|---|---|---|---|---|
| TGTGT | 9495565 | 4.5537047 | 46.0403 | 4 |
| TTTTT | 13327335 | 4.234609 | 5.560808 | 18 |
| AAAAA | 7660110 | 3.499263 | 3.6776595 | 43 |
| GTTTG | 7112260 | 3.410764 | 45.484642 | 7 |
| TGTTT | 8505475 | 3.320134 | 37.544735 | 6 |
| GTGTT | 6802105 | 3.262026 | 45.452267 | 5 |
| TTTGA | 7610970 | 3.1947095 | 40.007156 | 8 |
| CTTTG | 5743450 | 3.0096004 | 49.793285 | 1 |
| TTGTG | 6251340 | 2.9979002 | 45.46004 | 3 |
| GGGGG | 3005060 | 2.6721342 | 5.683897 | 13 |
| TTTGT | 6483920 | 2.5310147 | 37.239655 | 2 |
| TTGAG | 4629505 | 2.3873327 | 28.172224 | 9 |
| GAGGG | 2966750 | 2.3090503 | 12.166154 | 11 |
| TGAGG | 3292540 | 2.0859144 | 16.26188 | 10 |
| ACTTT | 3851125 | 1.7663263 | 19.31075 | 3 |
| AACTT | 3473450 | 1.7130839 | 21.396702 | 2 |
| AGGGG | 2143035 | 1.6679449 | 6.6826086 | 12 |
| TGAGC | 2170095 | 1.5022289 | 6.486358 | 10 |
| TGAGA | 2636590 | 1.4620267 | 5.2631855 | 10 |
| TAACT | 2715305 | 1.3391716 | 21.249712 | 1 |
| TGAGT | 2531780 | 1.3055826 | 7.9149117 | 10 |
| GAGTG | 2037435 | 1.2907709 | 6.0571346 | 11 |
| TTGAT | 2970205 | 1.2467455 | 8.947303 | 9 |
| TTGAC | 1978745 | 1.1149622 | 8.68537 | 9 |
| GTAAC | 1258200 | 0.7623498 | 6.2821393 | 1 |