![[OK]](Icons/tick.png) Basic Statistics
Basic Statistics
| Measure | Value |
|---|---|
| Filename | 140701_JD_C_12_CAGATC_L007_R1_001.fastq.gz |
| File type | Conventional base calls |
| Encoding | Sanger / Illumina 1.9 |
| Total Sequences | 34775151 |
| Filtered Sequences | 0 |
| Sequence length | 51 |
| %GC | 44 |
![[OK]](Icons/tick.png) Per base sequence quality
Per base sequence quality
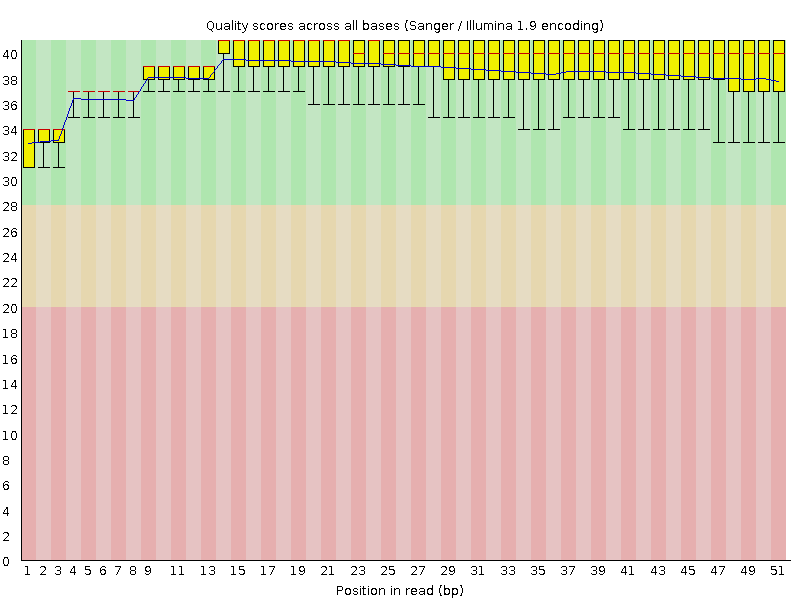
![[OK]](Icons/tick.png) Per sequence quality scores
Per sequence quality scores
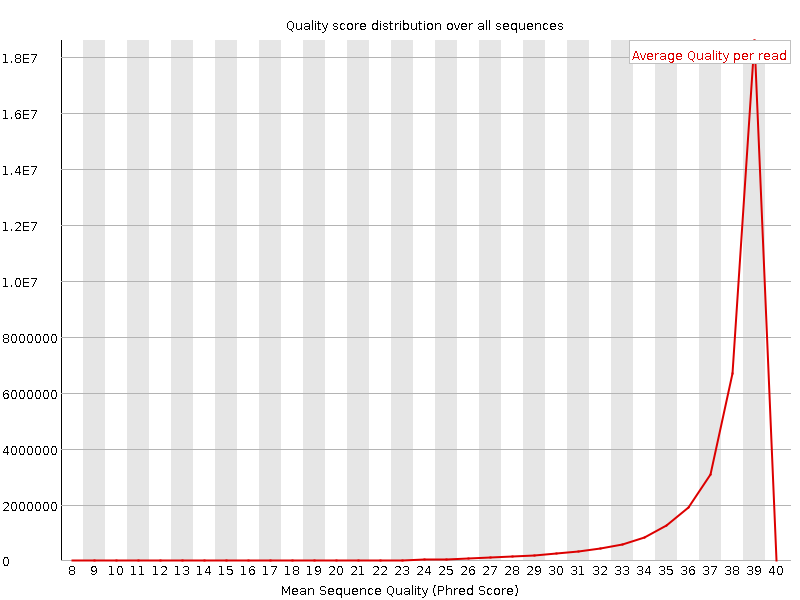
![[WARN]](Icons/warning.png) Per base sequence content
Per base sequence content

![[OK]](Icons/tick.png) Per base GC content
Per base GC content

![[OK]](Icons/tick.png) Per sequence GC content
Per sequence GC content

![[OK]](Icons/tick.png) Per base N content
Per base N content

![[OK]](Icons/tick.png) Sequence Length Distribution
Sequence Length Distribution
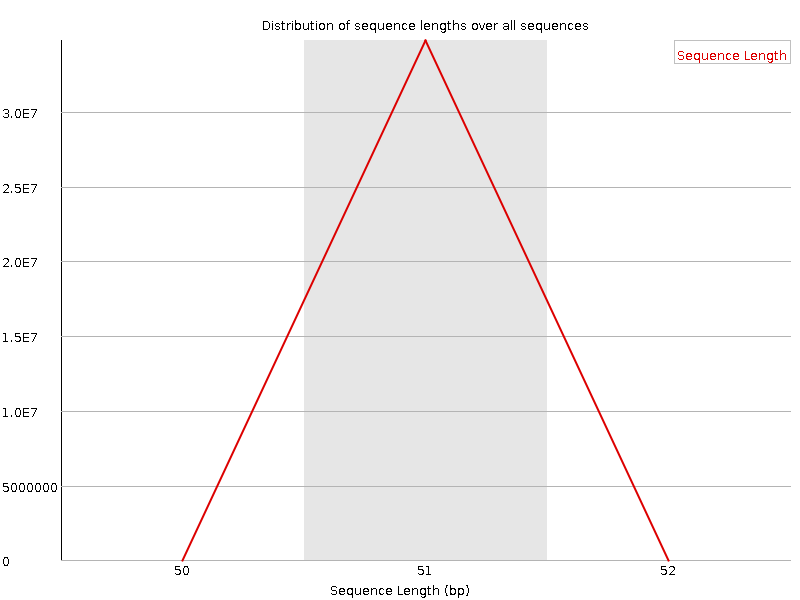
![[WARN]](Icons/warning.png) Sequence Duplication Levels
Sequence Duplication Levels
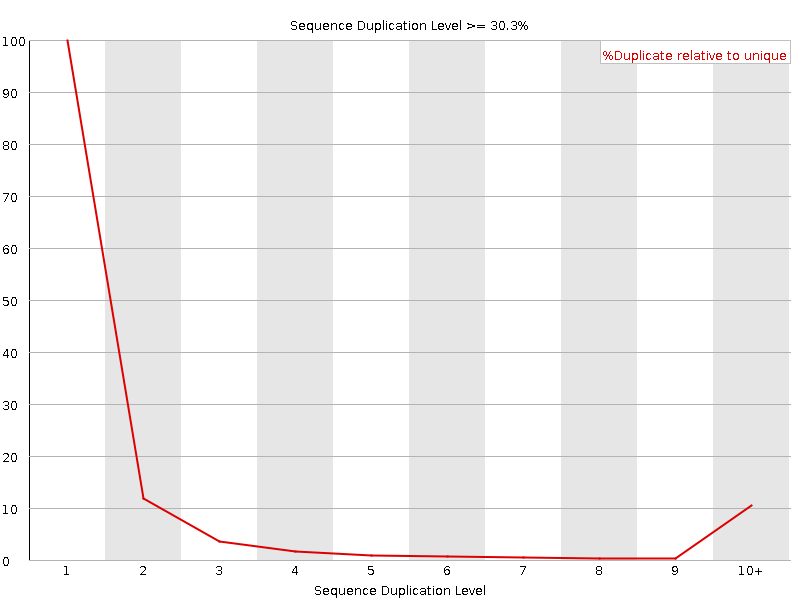
![[WARN]](Icons/warning.png) Overrepresented sequences
Overrepresented sequences
| Sequence | Count | Percentage | Possible Source |
|---|---|---|---|
| CTTTGTGTTTGATTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTT | 38247 | 0.10998370646902439 | No Hit |
![[WARN]](Icons/warning.png) Kmer Content
Kmer Content
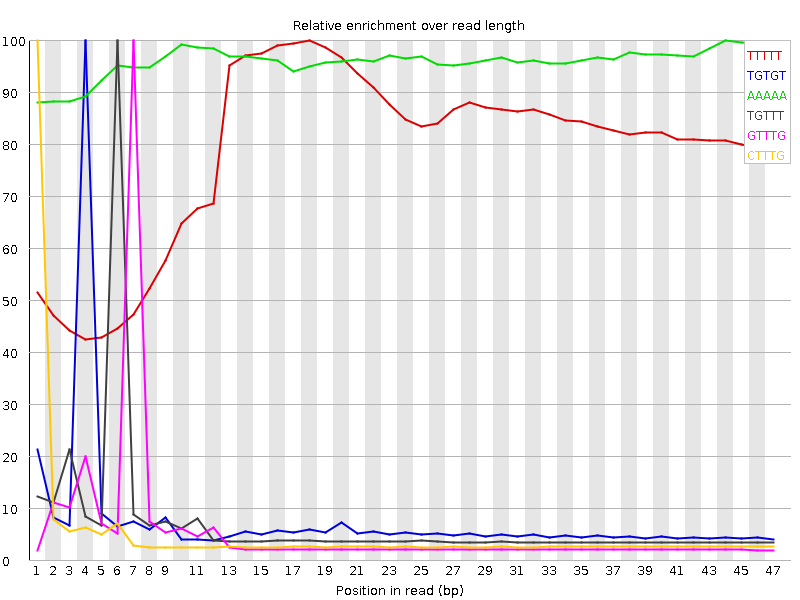
| Sequence | Count | Obs/Exp Overall | Obs/Exp Max | Max Obs/Exp Position |
|---|---|---|---|---|
| TTTTT | 12134085 | 4.0910363 | 5.202901 | 18 |
| TGTGT | 7261570 | 3.7000568 | 48.08881 | 4 |
| AAAAA | 7577010 | 3.295878 | 3.432467 | 44 |
| TGTTT | 6629620 | 2.7478354 | 39.11452 | 6 |
| GTTTG | 5211140 | 2.6552815 | 47.35654 | 7 |
| CTTTG | 4810375 | 2.5895305 | 49.97908 | 1 |
| TTTGA | 5783525 | 2.5224586 | 40.818058 | 8 |
| GTGTT | 4894500 | 2.493941 | 47.348038 | 5 |
| TTTGT | 5468025 | 2.266379 | 38.738586 | 2 |
| TTGTG | 4381090 | 2.2323384 | 47.327324 | 3 |
| GAGGG | 2573575 | 2.0854297 | 12.058115 | 11 |
| TTGAG | 3560675 | 1.909148 | 28.356714 | 9 |
| TGAGG | 2812440 | 1.8538141 | 16.48291 | 10 |
| AGGGG | 1878245 | 1.5219871 | 6.570871 | 12 |
| TGAGC | 2056090 | 1.431823 | 6.402751 | 10 |
| TGAGA | 2513790 | 1.4182917 | 5.291282 | 10 |
| GAGTG | 1883185 | 1.2412977 | 6.0551496 | 11 |
| TGAGT | 2293235 | 1.2295771 | 7.986811 | 10 |
| TTGAT | 2596875 | 1.1326156 | 9.471875 | 9 |
| TTGAC | 1738105 | 0.98457205 | 8.661533 | 9 |