![[OK]](Icons/tick.png) Basic Statistics
Basic Statistics
| Measure | Value |
|---|---|
| Filename | 140701_JD_C_10_ACTTGA_L007_R2_001.fastq.gz |
| File type | Conventional base calls |
| Encoding | Sanger / Illumina 1.9 |
| Total Sequences | 38097996 |
| Filtered Sequences | 0 |
| Sequence length | 51 |
| %GC | 44 |
![[OK]](Icons/tick.png) Per base sequence quality
Per base sequence quality
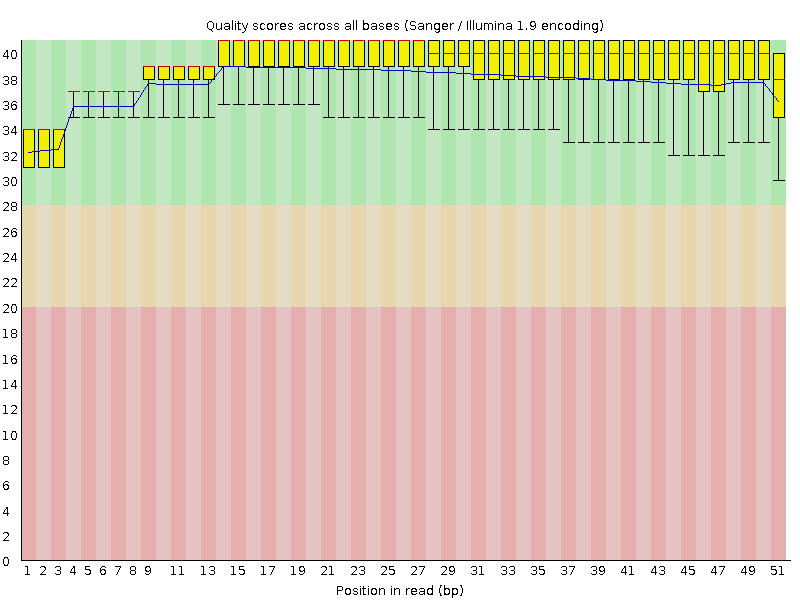
![[OK]](Icons/tick.png) Per sequence quality scores
Per sequence quality scores
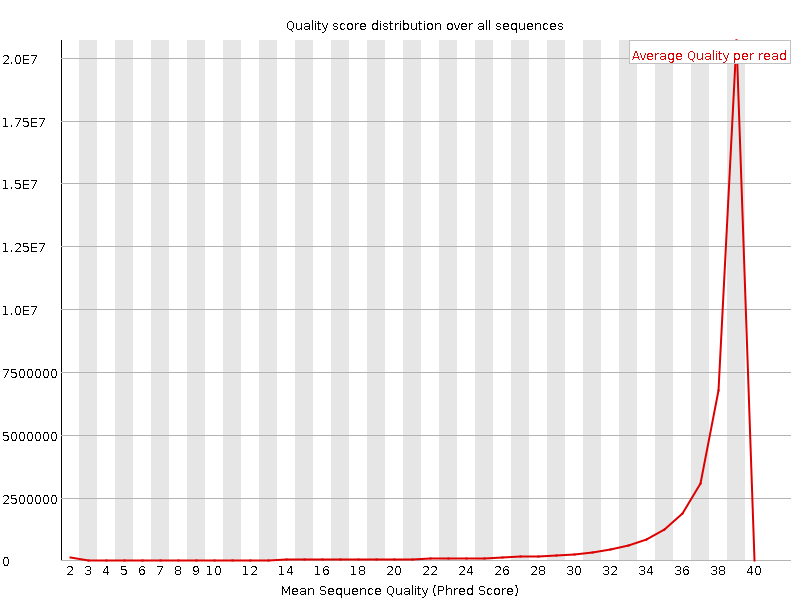
![[WARN]](Icons/warning.png) Per base sequence content
Per base sequence content

![[OK]](Icons/tick.png) Per base GC content
Per base GC content

![[OK]](Icons/tick.png) Per sequence GC content
Per sequence GC content

![[OK]](Icons/tick.png) Per base N content
Per base N content

![[OK]](Icons/tick.png) Sequence Length Distribution
Sequence Length Distribution
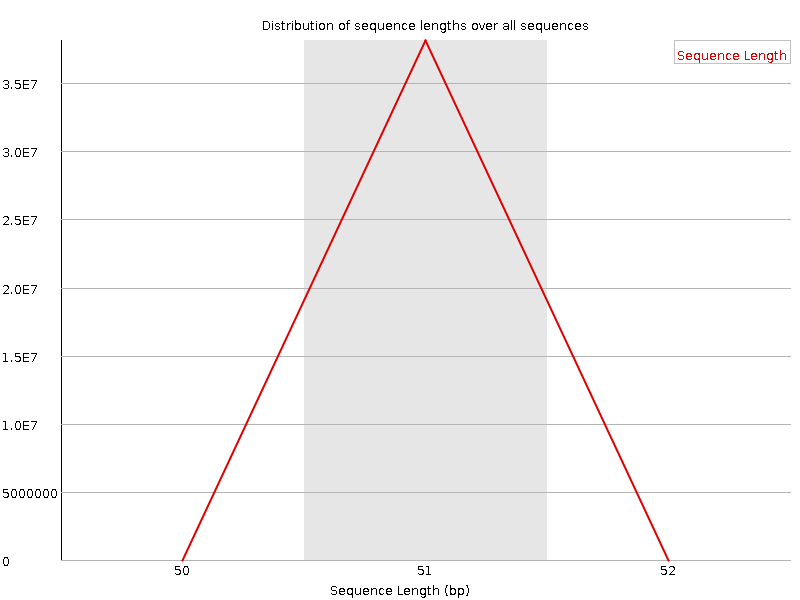
![[WARN]](Icons/warning.png) Sequence Duplication Levels
Sequence Duplication Levels
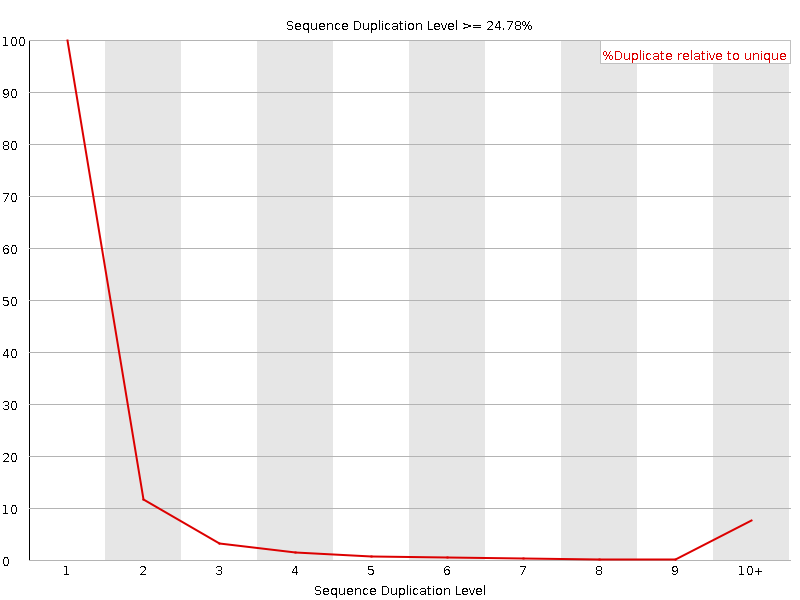
![[OK]](Icons/tick.png) Overrepresented sequences
Overrepresented sequences
No overrepresented sequences
![[FAIL]](Icons/error.png) Kmer Content
Kmer Content
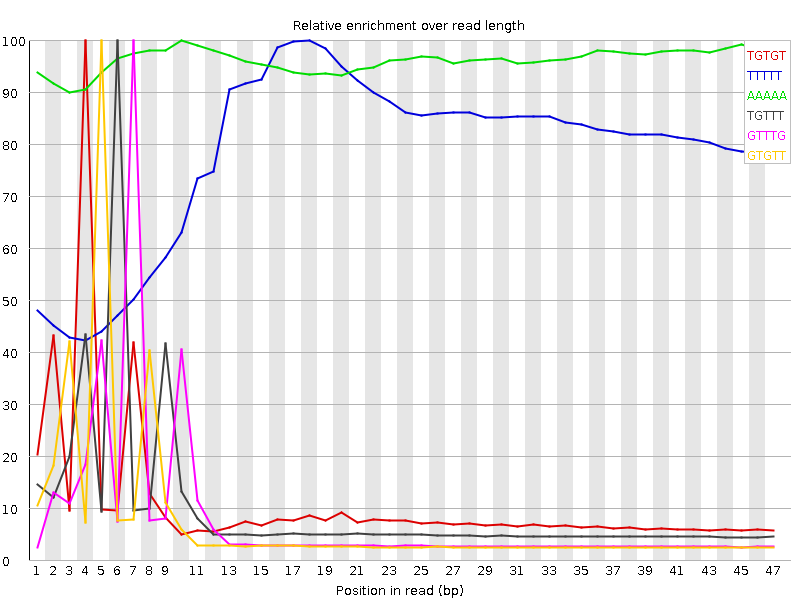
| Sequence | Count | Obs/Exp Overall | Obs/Exp Max | Max Obs/Exp Position |
|---|---|---|---|---|
| TGTGT | 9040680 | 4.1296597 | 37.642567 | 4 |
| TTTTT | 12543020 | 3.6222484 | 4.629211 | 18 |
| AAAAA | 8301425 | 3.0962448 | 3.2155762 | 10 |
| TGTTT | 8080640 | 2.9348803 | 30.042213 | 6 |
| GTTTG | 6372125 | 2.9107 | 37.021973 | 7 |
| GTGTT | 6059895 | 2.7680779 | 36.978928 | 5 |
| CTTTG | 5447680 | 2.6716015 | 39.834545 | 1 |
| TTTGA | 6970445 | 2.6645658 | 31.175259 | 8 |
| TTGTG | 5686525 | 2.5975273 | 37.01211 | 3 |
| GGGGG | 2783955 | 2.5297692 | 5.236898 | 13 |
| TTTGT | 6308510 | 2.2912445 | 29.702868 | 2 |
| GAGGG | 2907640 | 2.2111192 | 10.585115 | 11 |
| TTGAG | 4390945 | 2.1110206 | 22.986597 | 9 |
| TGAGG | 3326170 | 2.0111637 | 13.798611 | 10 |
| AGGGG | 2208005 | 1.6790807 | 6.0402665 | 12 |
| ACTTT | 3760450 | 1.5433084 | 13.294392 | 3 |
| TGAGC | 2278280 | 1.4789621 | 5.3223124 | 10 |
| AACTT | 3381385 | 1.4605919 | 14.377586 | 2 |
| GAGTG | 2156250 | 1.3037733 | 5.3318305 | 11 |
| TGAGT | 2708135 | 1.3019814 | 6.7429757 | 10 |
| TAACT | 2617840 | 1.1307778 | 14.224715 | 1 |
| TTGAT | 2939455 | 1.1236544 | 7.1277204 | 9 |
| TTGAC | 1917885 | 0.98992884 | 6.37162 | 9 |