![[OK]](Icons/tick.png) Basic Statistics
Basic Statistics
| Measure | Value |
|---|---|
| Filename | 140701_JD_B_9_ACTTGA_L004_R2_001.fastq.gz |
| File type | Conventional base calls |
| Encoding | Sanger / Illumina 1.9 |
| Total Sequences | 26710200 |
| Filtered Sequences | 0 |
| Sequence length | 51 |
| %GC | 44 |
![[OK]](Icons/tick.png) Per base sequence quality
Per base sequence quality
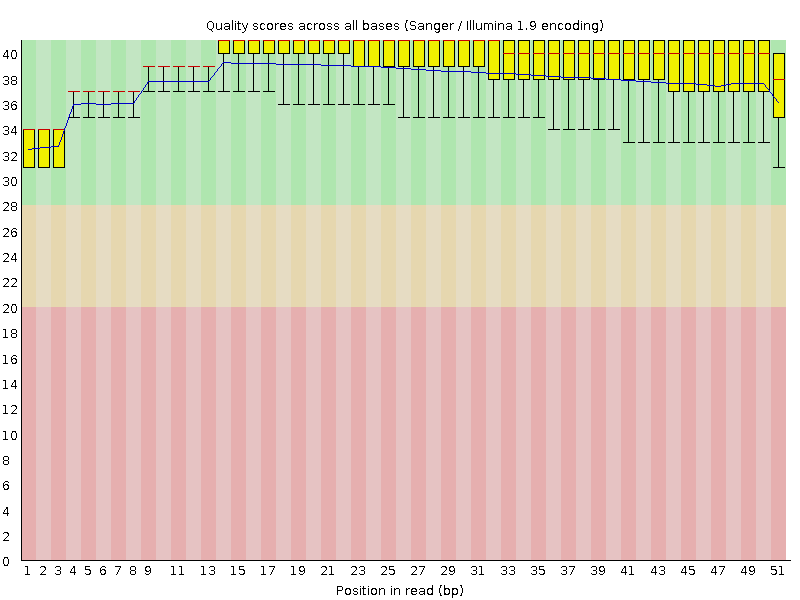
![[OK]](Icons/tick.png) Per sequence quality scores
Per sequence quality scores
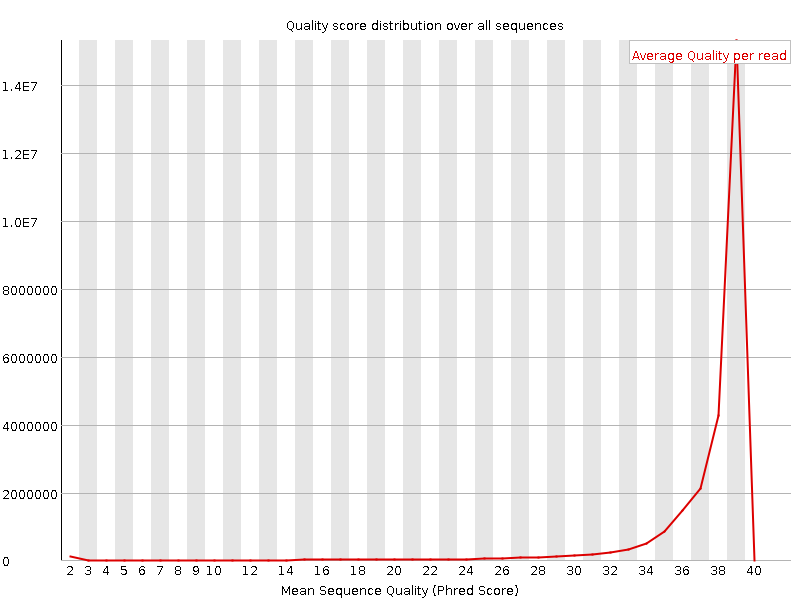
![[FAIL]](Icons/error.png) Per base sequence content
Per base sequence content

![[WARN]](Icons/warning.png) Per base GC content
Per base GC content

![[WARN]](Icons/warning.png) Per sequence GC content
Per sequence GC content

![[OK]](Icons/tick.png) Per base N content
Per base N content

![[OK]](Icons/tick.png) Sequence Length Distribution
Sequence Length Distribution
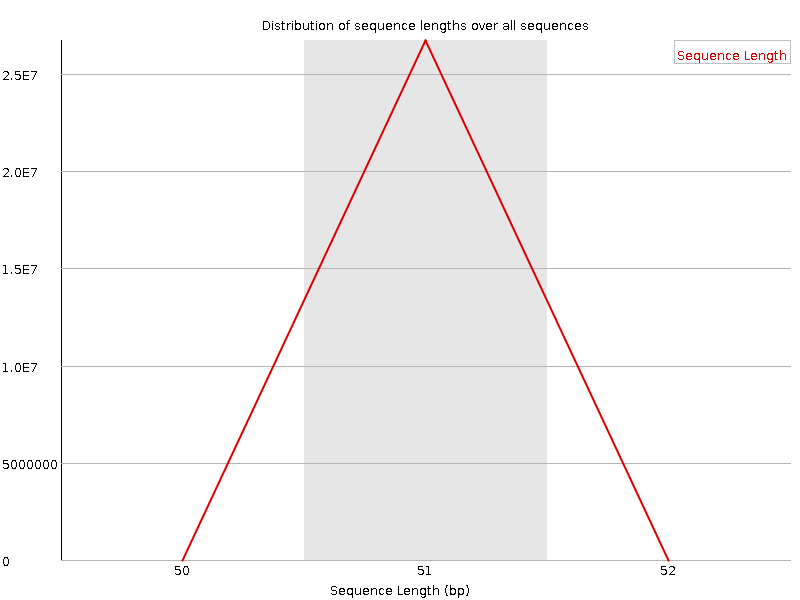
![[WARN]](Icons/warning.png) Sequence Duplication Levels
Sequence Duplication Levels
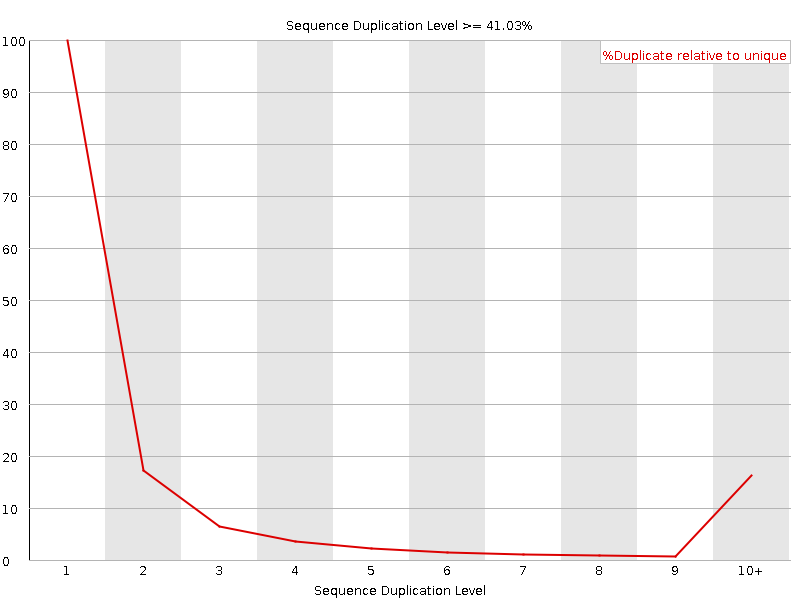
![[WARN]](Icons/warning.png) Overrepresented sequences
Overrepresented sequences
| Sequence | Count | Percentage | Possible Source |
|---|---|---|---|
| CTTTGTGTTTGATTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTT | 171582 | 0.6423838084327336 | No Hit |
| TTGTGTTTGATTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTT | 133447 | 0.4996106356373221 | No Hit |
| NNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNN | 131632 | 0.4928154787309717 | No Hit |
| TAACTTTGTGTTTGATTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTT | 79454 | 0.29746688530973187 | No Hit |
| AAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAA | 63886 | 0.23918203532732815 | No Hit |
| TGTGTTTGATTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTT | 51010 | 0.19097573211731847 | No Hit |
| TTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTT | 30022 | 0.11239900861843041 | No Hit |
![[FAIL]](Icons/error.png) Kmer Content
Kmer Content
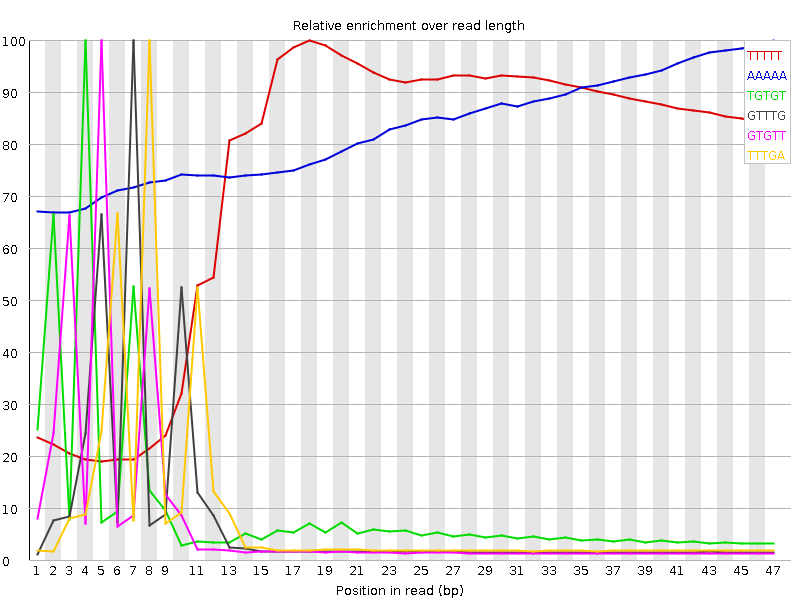
| Sequence | Count | Obs/Exp Overall | Obs/Exp Max | Max Obs/Exp Position |
|---|---|---|---|---|
| TTTTT | 32363175 | 10.76092 | 14.422021 | 18 |
| AAAAA | 9493745 | 6.778718 | 8.17986 | 47 |
| TGTGT | 8929615 | 4.8136263 | 48.571033 | 4 |
| GTTTG | 6995795 | 3.7711751 | 48.067993 | 7 |
| GTGTT | 6760850 | 3.6445248 | 48.232937 | 5 |
| TTTGA | 7267015 | 3.5847375 | 44.15633 | 8 |
| TGTTT | 7926095 | 3.3556619 | 38.221493 | 6 |
| TTGTG | 6130455 | 3.3047023 | 48.184326 | 3 |
| CTTTG | 5135000 | 3.127746 | 54.590427 | 1 |
| GGGGG | 2692040 | 2.995596 | 8.305384 | 13 |
| GAGGG | 2837390 | 2.8892322 | 17.592487 | 11 |
| TTGAG | 4412615 | 2.7715201 | 32.412083 | 9 |
| TTTGT | 5713800 | 2.419045 | 38.079147 | 2 |
| GAGAG | 2584325 | 2.408087 | 5.524552 | 11 |
| TGAGG | 2990340 | 2.3914607 | 20.028173 | 10 |
| AGGGG | 1976510 | 2.0126228 | 10.079486 | 12 |
| AACTT | 3091140 | 2.0074916 | 30.76396 | 2 |
| ACTTT | 3384930 | 1.8866982 | 25.787325 | 3 |
| TGAGC | 1744460 | 1.5763589 | 7.358854 | 10 |
| TAACT | 2413195 | 1.567211 | 30.55391 | 1 |
| ATTTT | 3854115 | 1.4931564 | 5.6192555 | 12 |
| TGAGA | 1987020 | 1.4541411 | 5.685767 | 10 |
| TTGAT | 2776040 | 1.3693895 | 11.696542 | 9 |
| GAGTG | 1656560 | 1.3247987 | 7.346048 | 11 |
| TGAGT | 2055855 | 1.2912621 | 9.006869 | 10 |
| TGATT | 2331820 | 1.1502606 | 7.8872247 | 10 |
| GATTT | 2268130 | 1.1188433 | 6.7307315 | 11 |
| TTGAC | 1554740 | 1.1033955 | 8.374096 | 9 |
| GTAAC | 1094745 | 0.90525043 | 9.031113 | 1 |
| GGTAA | 1093520 | 0.80026 | 5.3407173 | 1 |