![[OK]](Icons/tick.png) Basic Statistics
Basic Statistics
| Measure | Value |
|---|---|
| Filename | 140701_JD_B_9_ACTTGA_L004_R1_001.fastq.gz |
| File type | Conventional base calls |
| Encoding | Sanger / Illumina 1.9 |
| Total Sequences | 26710200 |
| Filtered Sequences | 0 |
| Sequence length | 51 |
| %GC | 44 |
![[OK]](Icons/tick.png) Per base sequence quality
Per base sequence quality
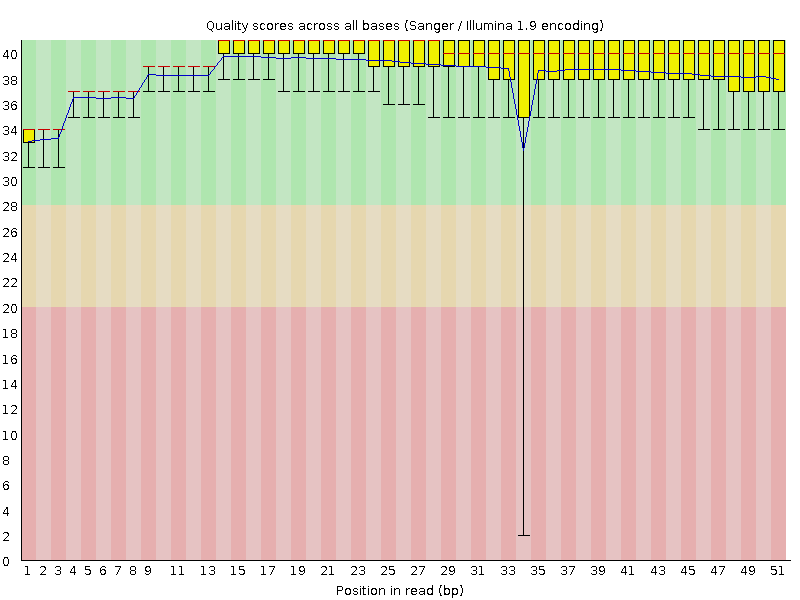
![[OK]](Icons/tick.png) Per sequence quality scores
Per sequence quality scores
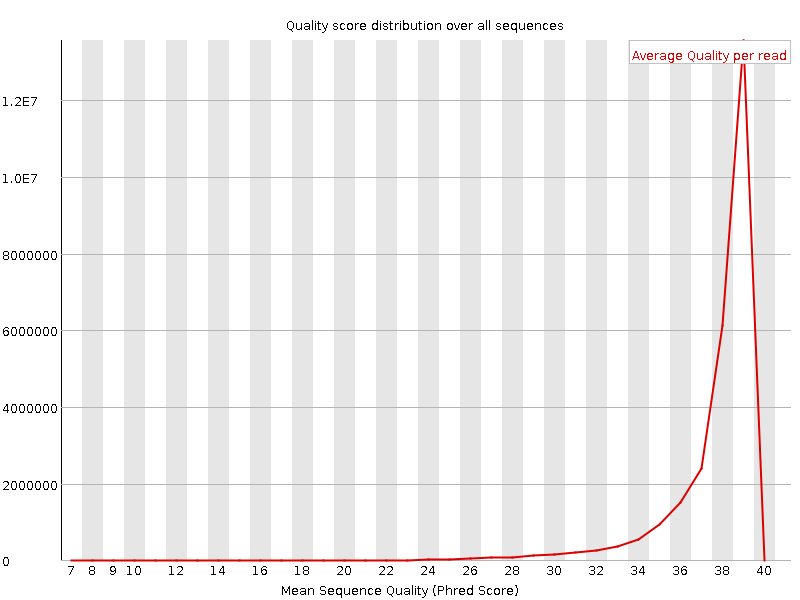
![[WARN]](Icons/warning.png) Per base sequence content
Per base sequence content

![[OK]](Icons/tick.png) Per base GC content
Per base GC content

![[WARN]](Icons/warning.png) Per sequence GC content
Per sequence GC content

![[WARN]](Icons/warning.png) Per base N content
Per base N content

![[OK]](Icons/tick.png) Sequence Length Distribution
Sequence Length Distribution
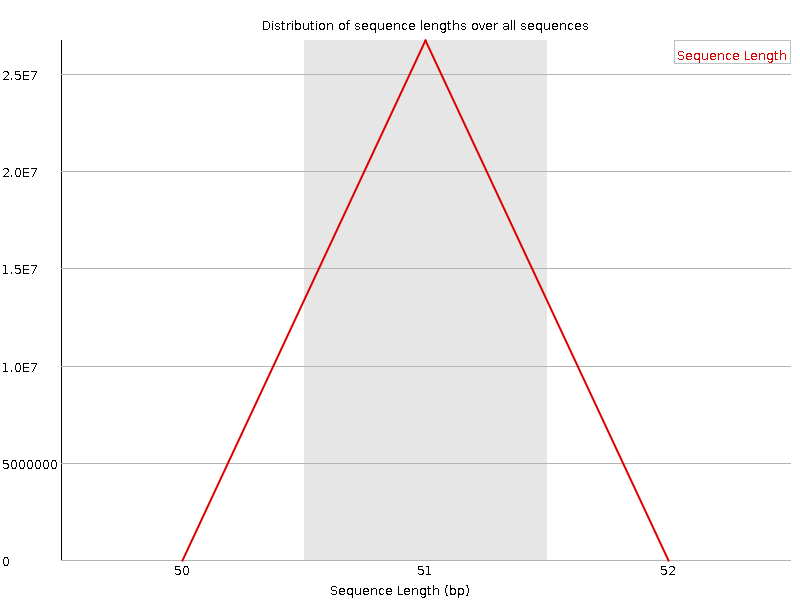
![[WARN]](Icons/warning.png) Sequence Duplication Levels
Sequence Duplication Levels
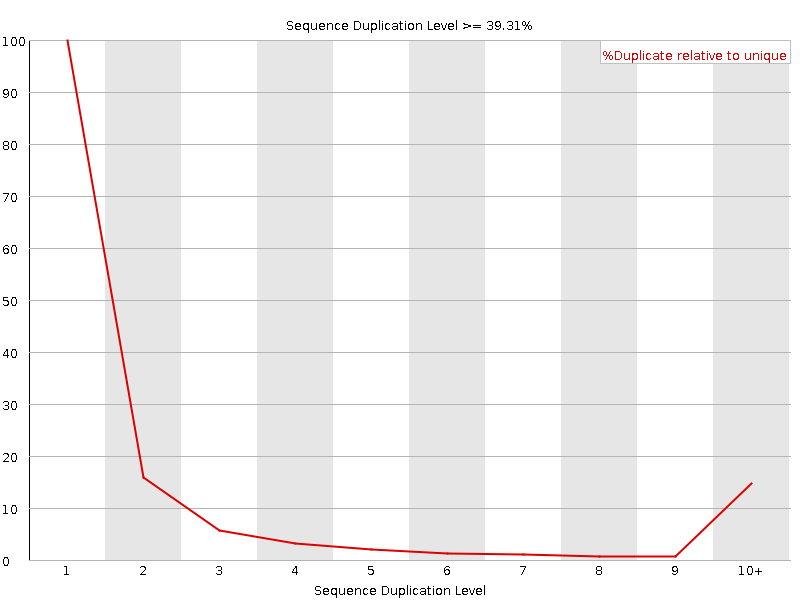
![[WARN]](Icons/warning.png) Overrepresented sequences
Overrepresented sequences
| Sequence | Count | Percentage | Possible Source |
|---|---|---|---|
| CTTTGTGTTTGATTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTT | 158140 | 0.5920584645566113 | No Hit |
| AAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAA | 82619 | 0.30931629115468995 | No Hit |
| TGTGTTTGATTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTT | 48219 | 0.18052654042275984 | No Hit |
| TTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTT | 37150 | 0.1390854430142792 | No Hit |
![[FAIL]](Icons/error.png) Kmer Content
Kmer Content
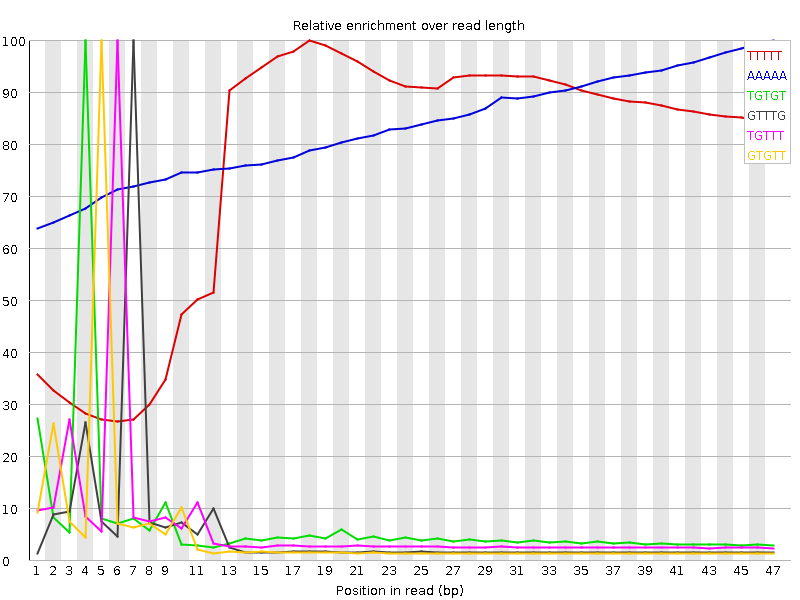
| Sequence | Count | Obs/Exp Overall | Obs/Exp Max | Max Obs/Exp Position |
|---|---|---|---|---|
| TTTTT | 24739215 | 9.988469 | 13.00193 | 18 |
| AAAAA | 11771840 | 7.5736833 | 9.106115 | 47 |
| TGTGT | 6224195 | 3.90355 | 56.038948 | 4 |
| GTTTG | 4860365 | 3.0482137 | 55.622253 | 7 |
| TGTTT | 5794705 | 2.9159236 | 45.117126 | 6 |
| GTGTT | 4597365 | 2.8832715 | 55.581337 | 5 |
| TTTGA | 5195790 | 2.8698971 | 49.191525 | 8 |
| CTTTG | 4184010 | 2.8120332 | 59.452 | 1 |
| TTGTG | 3945895 | 2.474697 | 55.476864 | 3 |
| GAGGG | 2289830 | 2.4485743 | 16.770914 | 11 |
| TTTGT | 4662955 | 2.3464215 | 44.7441 | 2 |
| GGGGG | 1899405 | 2.3061602 | 6.99837 | 13 |
| TTGAG | 3165520 | 2.1791725 | 34.14373 | 9 |
| GAGAG | 2242500 | 2.1119356 | 5.1700087 | 11 |
| TGAGG | 2394815 | 2.0547085 | 20.637424 | 10 |
| AGGGG | 1615275 | 1.7272552 | 9.596168 | 12 |
| ATTTT | 3545270 | 1.571203 | 6.948163 | 12 |
| TGAGC | 1601050 | 1.47209 | 7.4525084 | 10 |
| ACTTT | 2367950 | 1.4016463 | 5.5554156 | 5 |
| TGAGA | 1842880 | 1.3925587 | 5.754972 | 10 |
| AACTT | 2130935 | 1.3845417 | 6.0861497 | 4 |
| GAGTG | 1456310 | 1.249488 | 7.416614 | 11 |
| TTGAT | 2257895 | 1.2471493 | 13.693929 | 9 |
| TGAGT | 1756440 | 1.2091491 | 9.53137 | 10 |
| TGATT | 2006235 | 1.1081449 | 9.435777 | 10 |
| GATTT | 1975120 | 1.0909585 | 8.1214905 | 11 |
| TTGAC | 1329475 | 0.98079425 | 9.066769 | 9 |
| TAACT | 1448835 | 0.94135785 | 5.7832828 | 3 |
| GGTAA | 1091600 | 0.8248595 | 6.4768896 | 1 |
| GTAAC | 981425 | 0.7947394 | 6.956533 | 2 |