![[OK]](Icons/tick.png) Basic Statistics
Basic Statistics
| Measure | Value |
|---|---|
| Filename | 140701_JD_B_6_CTTGTA_L008_R1_001.fastq.gz |
| File type | Conventional base calls |
| Encoding | Sanger / Illumina 1.9 |
| Total Sequences | 34987315 |
| Filtered Sequences | 0 |
| Sequence length | 51 |
| %GC | 45 |
![[OK]](Icons/tick.png) Per base sequence quality
Per base sequence quality
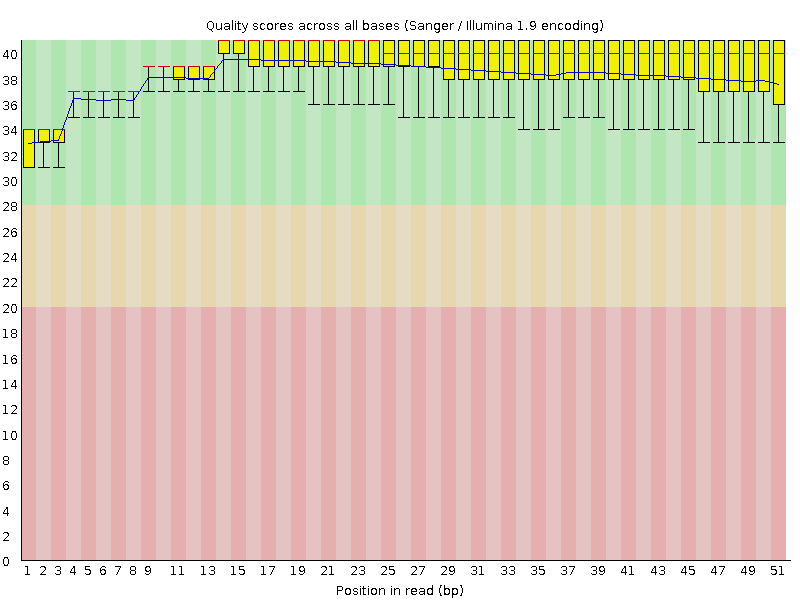
![[OK]](Icons/tick.png) Per sequence quality scores
Per sequence quality scores
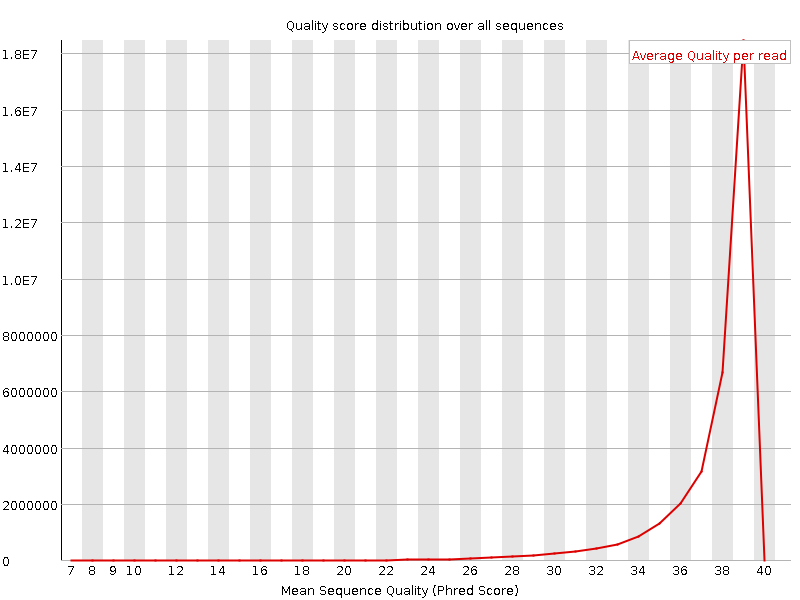
![[WARN]](Icons/warning.png) Per base sequence content
Per base sequence content

![[OK]](Icons/tick.png) Per base GC content
Per base GC content

![[OK]](Icons/tick.png) Per sequence GC content
Per sequence GC content

![[OK]](Icons/tick.png) Per base N content
Per base N content

![[OK]](Icons/tick.png) Sequence Length Distribution
Sequence Length Distribution
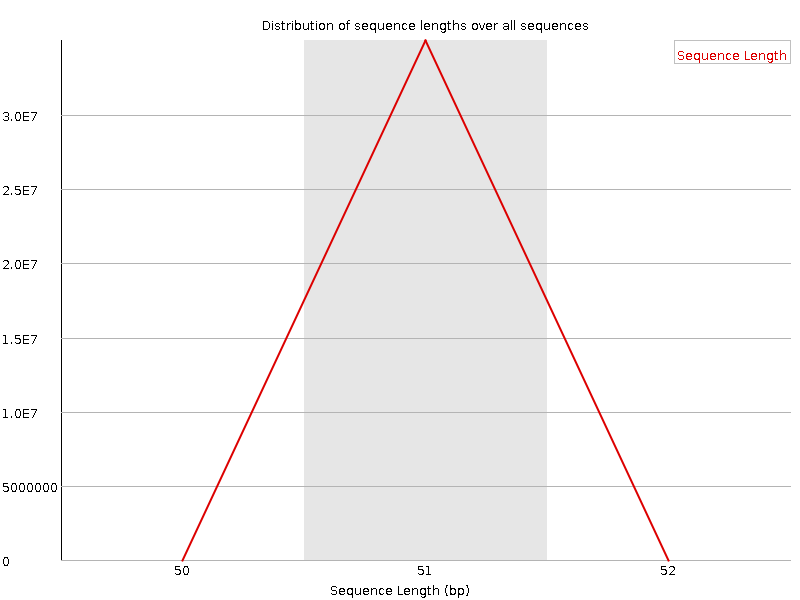
![[WARN]](Icons/warning.png) Sequence Duplication Levels
Sequence Duplication Levels
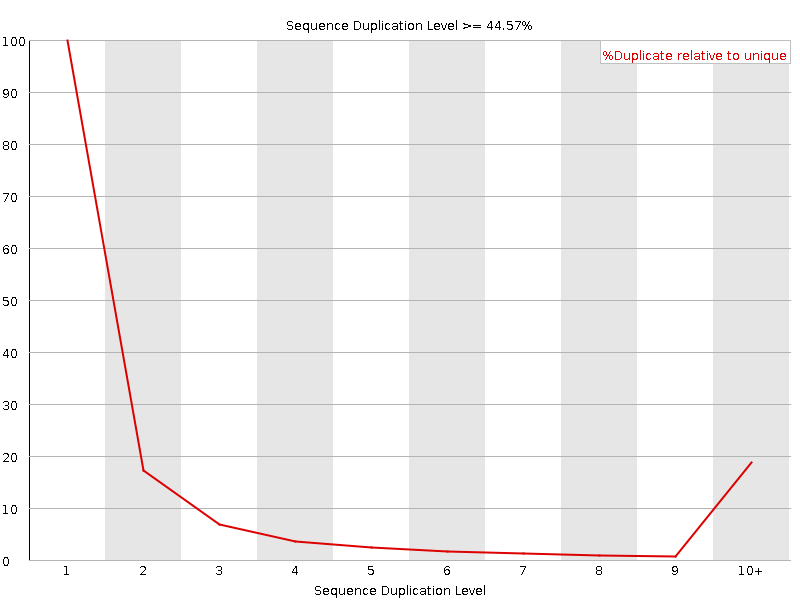
![[WARN]](Icons/warning.png) Overrepresented sequences
Overrepresented sequences
| Sequence | Count | Percentage | Possible Source |
|---|---|---|---|
| CTTTGTGTTTGATTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTT | 121477 | 0.34720297913686715 | No Hit |
| AAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAA | 46633 | 0.13328544931212927 | No Hit |
![[WARN]](Icons/warning.png) Kmer Content
Kmer Content
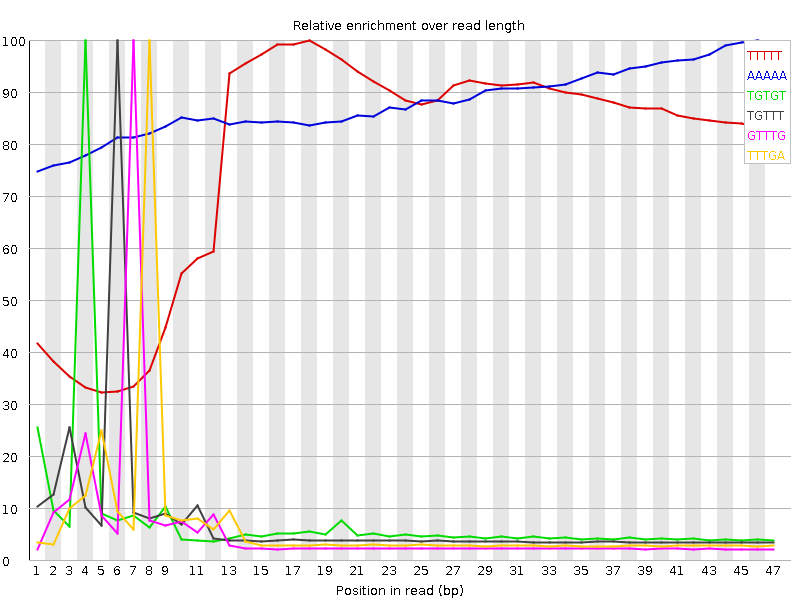
| Sequence | Count | Obs/Exp Overall | Obs/Exp Max | Max Obs/Exp Position |
|---|---|---|---|---|
| TTTTT | 21521165 | 7.503721 | 9.599831 | 18 |
| AAAAA | 10082430 | 4.766026 | 5.4079633 | 46 |
| TGTGT | 6279975 | 3.1873515 | 41.665623 | 4 |
| TGTTT | 6073610 | 2.554985 | 34.684956 | 6 |
| GTTTG | 4866770 | 2.4700906 | 41.236374 | 7 |
| TTTGA | 5429420 | 2.4273417 | 36.584167 | 8 |
| CTTTG | 4500095 | 2.3900003 | 43.19886 | 1 |
| GTGTT | 4562670 | 2.315747 | 41.151363 | 5 |
| TTTGT | 5022435 | 2.1127875 | 34.46347 | 2 |
| GAGGG | 2681280 | 2.1052873 | 10.50386 | 11 |
| TTGTG | 3990580 | 2.0253873 | 41.1392 | 3 |
| TTGAG | 3290735 | 1.7750108 | 23.919945 | 9 |
| TGAGG | 2665425 | 1.7346231 | 13.810067 | 10 |
| ATTTT | 4593835 | 1.7022461 | 5.129668 | 12 |
| AGGGG | 1899025 | 1.4910764 | 5.7088513 | 12 |
| TGAGC | 2067825 | 1.4081752 | 5.835283 | 10 |
| TTGAT | 2649850 | 1.1846739 | 9.916866 | 9 |
| TGAGT | 2035885 | 1.0981492 | 6.5302787 | 10 |
| GATTT | 2402825 | 1.0742359 | 5.392791 | 11 |
| TGATT | 2402515 | 1.0740973 | 6.446577 | 10 |
| TTGAC | 1668475 | 0.94174224 | 7.554342 | 9 |