![[OK]](Icons/tick.png) Basic Statistics
Basic Statistics
| Measure | Value |
|---|---|
| Filename | 140701_JD_B_4_GCCAAT_L008_R1_001.fastq.gz |
| File type | Conventional base calls |
| Encoding | Sanger / Illumina 1.9 |
| Total Sequences | 33178573 |
| Filtered Sequences | 0 |
| Sequence length | 51 |
| %GC | 43 |
![[OK]](Icons/tick.png) Per base sequence quality
Per base sequence quality
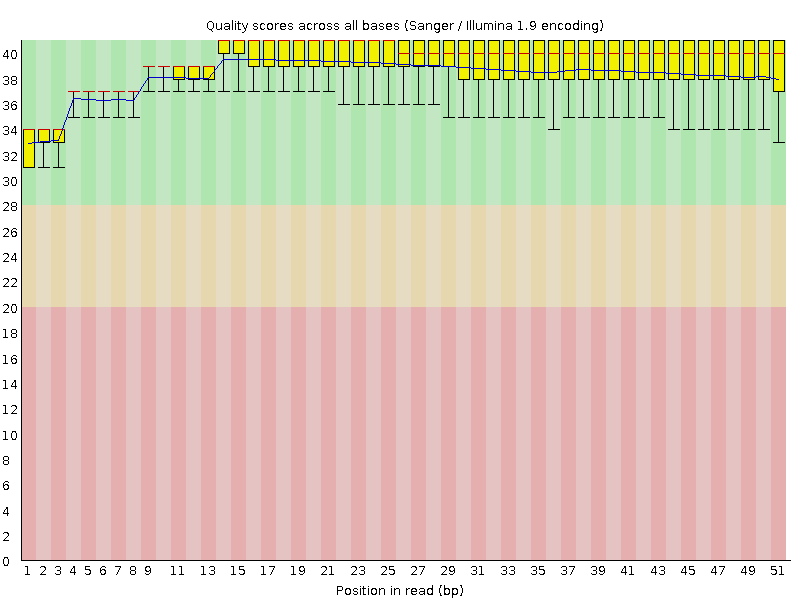
![[OK]](Icons/tick.png) Per sequence quality scores
Per sequence quality scores
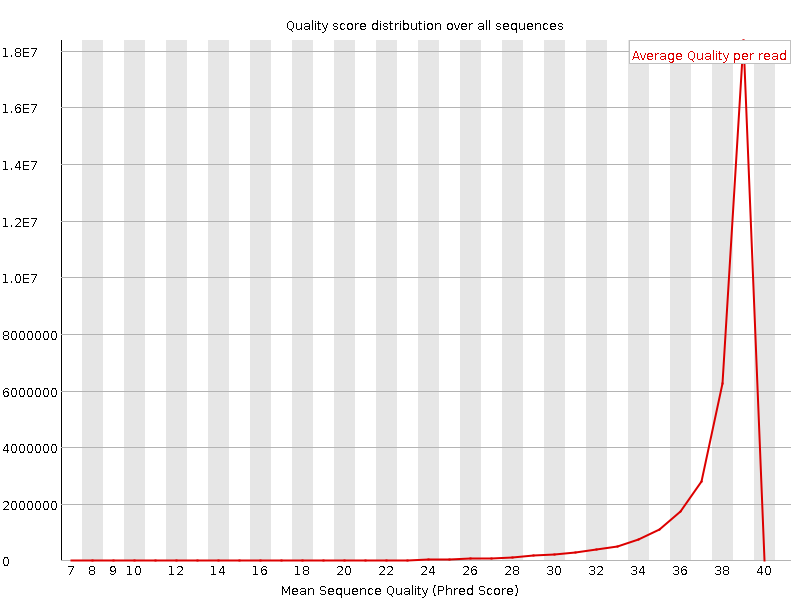
![[WARN]](Icons/warning.png) Per base sequence content
Per base sequence content

![[OK]](Icons/tick.png) Per base GC content
Per base GC content

![[OK]](Icons/tick.png) Per sequence GC content
Per sequence GC content

![[OK]](Icons/tick.png) Per base N content
Per base N content

![[OK]](Icons/tick.png) Sequence Length Distribution
Sequence Length Distribution
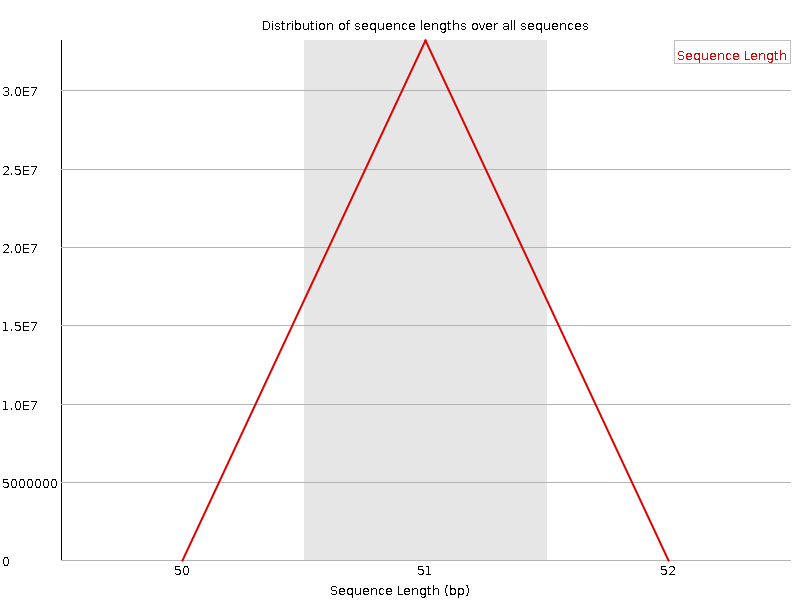
![[WARN]](Icons/warning.png) Sequence Duplication Levels
Sequence Duplication Levels
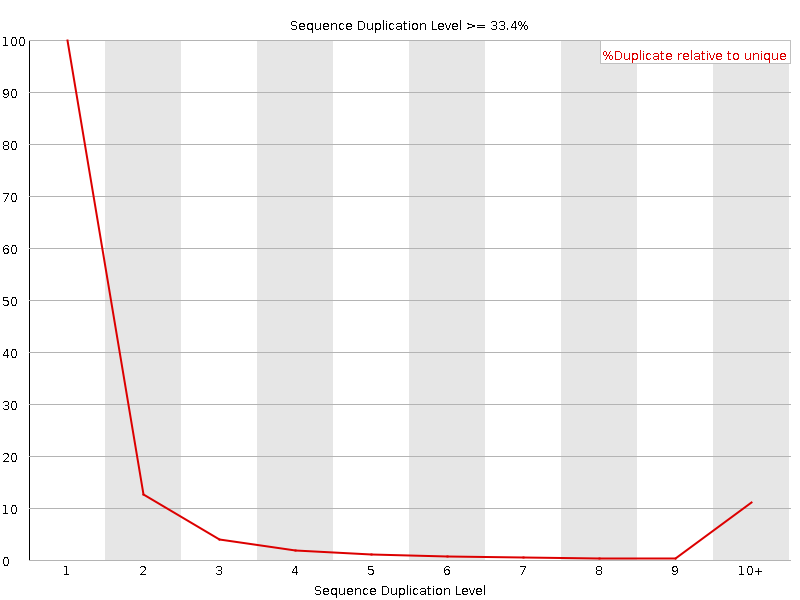
![[WARN]](Icons/warning.png) Overrepresented sequences
Overrepresented sequences
| Sequence | Count | Percentage | Possible Source |
|---|---|---|---|
| CTTTGTGTTTGATTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTT | 81263 | 0.24492614555785747 | No Hit |
![[WARN]](Icons/warning.png) Kmer Content
Kmer Content
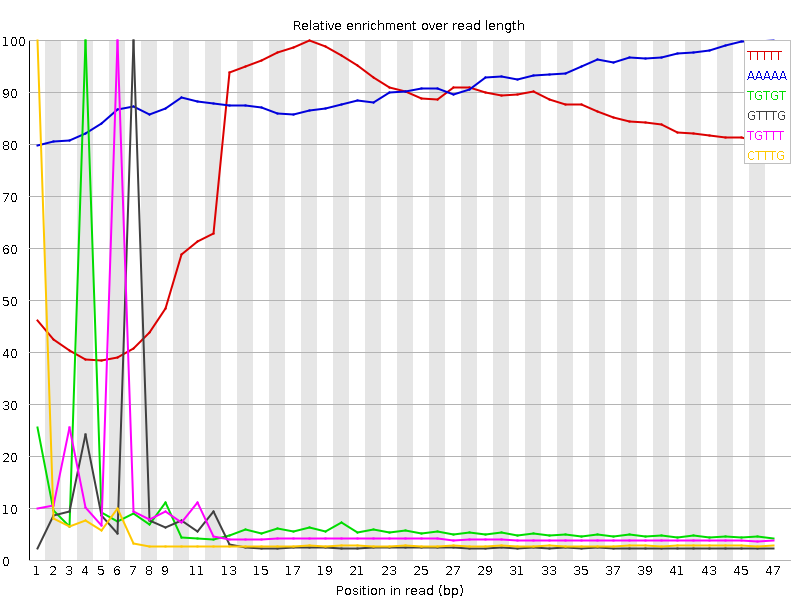
| Sequence | Count | Obs/Exp Overall | Obs/Exp Max | Max Obs/Exp Position |
|---|---|---|---|---|
| TTTTT | 15858740 | 5.0921187 | 6.4895306 | 18 |
| AAAAA | 8950625 | 3.6832616 | 4.058872 | 47 |
| TGTGT | 6629865 | 3.517048 | 43.122173 | 4 |
| GTTTG | 4845930 | 2.5706959 | 42.562416 | 7 |
| TGTTT | 6191280 | 2.5552425 | 33.665844 | 6 |
| CTTTG | 4369630 | 2.446783 | 44.877274 | 1 |
| GTGTT | 4485725 | 2.3796124 | 42.426605 | 5 |
| TTTGA | 5326150 | 2.3100162 | 35.01415 | 8 |
| GAGGG | 2415925 | 2.225106 | 13.060331 | 11 |
| GGGGG | 1874800 | 2.1120014 | 5.5030594 | 13 |
| TTGTG | 3979895 | 2.1112769 | 42.444675 | 3 |
| TTTGT | 5081235 | 2.0971088 | 33.390835 | 2 |
| TTGAG | 3398855 | 1.8947685 | 26.591604 | 9 |
| TGAGG | 2627555 | 1.8827696 | 16.240572 | 10 |
| AGGGG | 1731925 | 1.5951309 | 7.131492 | 12 |
| TGAGC | 1937895 | 1.4657259 | 6.4378104 | 10 |
| GAGTG | 1688710 | 1.210042 | 5.98336 | 11 |
| TGAGT | 2160065 | 1.2041768 | 7.61618 | 10 |
| TTGAT | 2461750 | 1.067691 | 8.31154 | 9 |
| TTGAC | 1597215 | 0.93986183 | 6.9103503 | 9 |