![[OK]](Icons/tick.png) Basic Statistics
Basic Statistics
| Measure | Value |
|---|---|
| Filename | 140701_JD_B_1_CGATGT_L003_R2_001.fastq.gz |
| File type | Conventional base calls |
| Encoding | Sanger / Illumina 1.9 |
| Total Sequences | 25095039 |
| Filtered Sequences | 0 |
| Sequence length | 51 |
| %GC | 44 |
![[OK]](Icons/tick.png) Per base sequence quality
Per base sequence quality
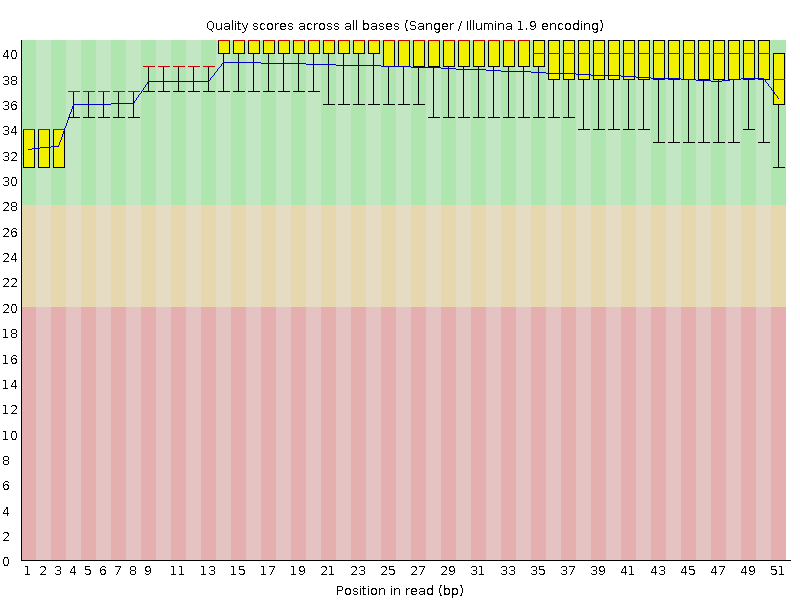
![[OK]](Icons/tick.png) Per sequence quality scores
Per sequence quality scores
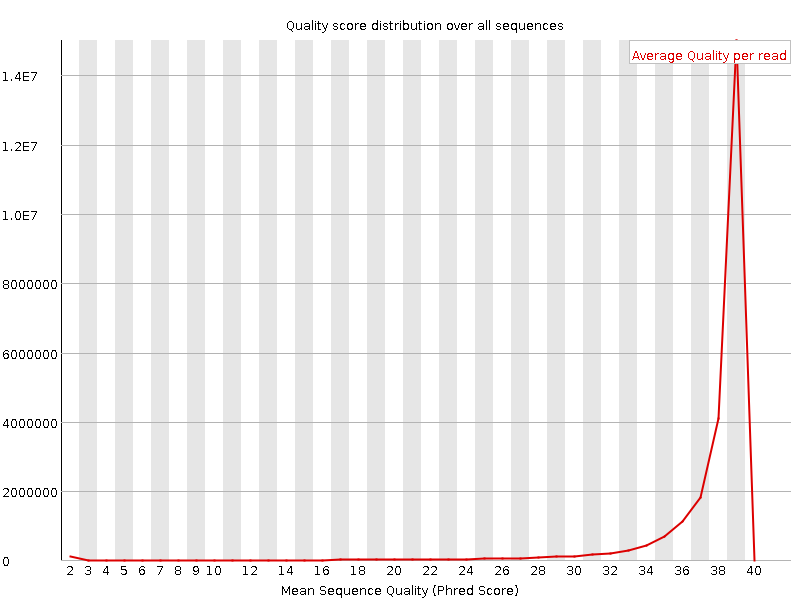
![[WARN]](Icons/warning.png) Per base sequence content
Per base sequence content

![[OK]](Icons/tick.png) Per base GC content
Per base GC content

![[OK]](Icons/tick.png) Per sequence GC content
Per sequence GC content

![[OK]](Icons/tick.png) Per base N content
Per base N content

![[OK]](Icons/tick.png) Sequence Length Distribution
Sequence Length Distribution
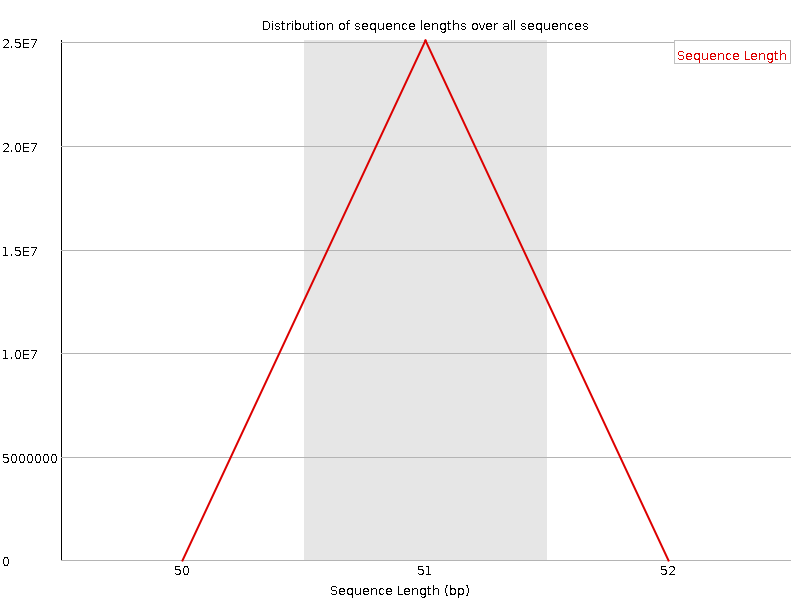
![[WARN]](Icons/warning.png) Sequence Duplication Levels
Sequence Duplication Levels
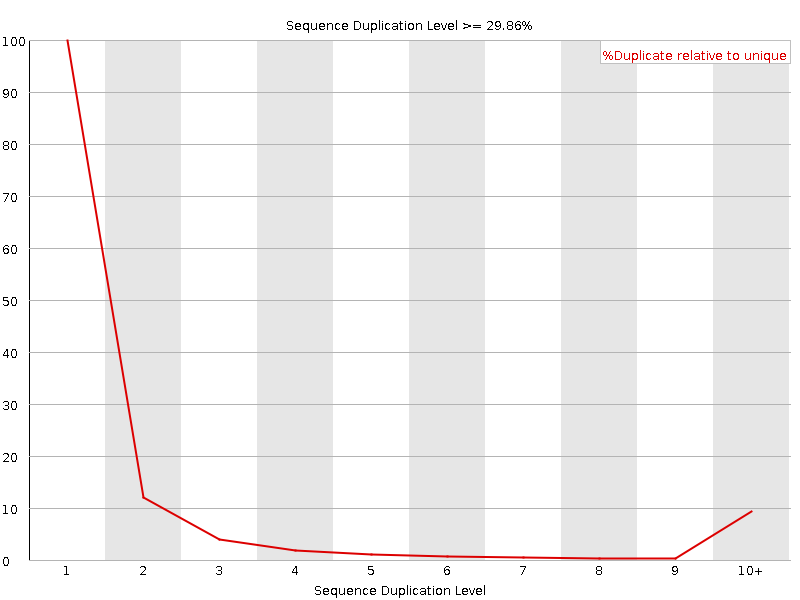
![[WARN]](Icons/warning.png) Overrepresented sequences
Overrepresented sequences
| Sequence | Count | Percentage | Possible Source |
|---|---|---|---|
| NNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNN | 110383 | 0.4398598464023108 | No Hit |
| CTTTGTGTTTGATTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTT | 47301 | 0.18848745363575647 | No Hit |
| TTGTGTTTGATTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTT | 31346 | 0.12490915037031822 | No Hit |
![[WARN]](Icons/warning.png) Kmer Content
Kmer Content
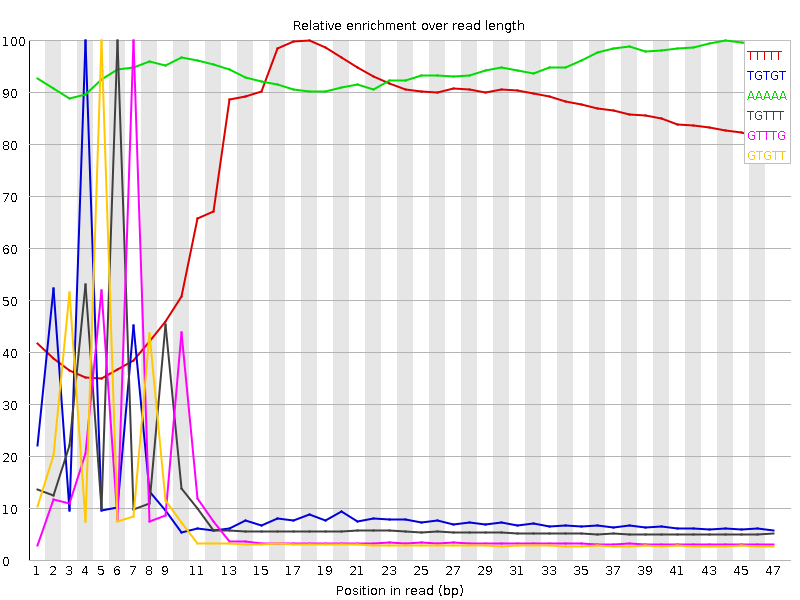
| Sequence | Count | Obs/Exp Overall | Obs/Exp Max | Max Obs/Exp Position |
|---|---|---|---|---|
| TTTTT | 12818800 | 5.7252717 | 7.3416505 | 18 |
| TGTGT | 5230475 | 3.6471717 | 31.787731 | 4 |
| AAAAA | 6051825 | 3.6053236 | 3.809186 | 44 |
| TGTTT | 4860785 | 2.7126157 | 25.582178 | 6 |
| GTTTG | 3837630 | 2.675951 | 31.163576 | 7 |
| GTGTT | 3623405 | 2.5265737 | 31.242018 | 5 |
| TTTGA | 4200130 | 2.4829414 | 26.637743 | 8 |
| CTTTG | 3298045 | 2.4416635 | 33.377018 | 1 |
| TTGTG | 3389805 | 2.363686 | 31.1664 | 3 |
| GAGGG | 1899670 | 2.1906927 | 9.357859 | 11 |
| TTTGT | 3767675 | 2.1025934 | 25.344814 | 2 |
| TTGAG | 2635420 | 1.946645 | 19.326975 | 9 |
| TGAGG | 2081585 | 1.9211631 | 11.7481365 | 10 |
| AGGGG | 1408300 | 1.6240466 | 5.288771 | 12 |
| ACTTT | 2417660 | 1.5174456 | 12.3122835 | 3 |
| AACTT | 2260610 | 1.5030221 | 13.386969 | 2 |
| TGAGT | 1615350 | 1.1931733 | 5.4727216 | 10 |
| TTGAT | 1892850 | 1.1189739 | 6.599864 | 9 |
| TAACT | 1652790 | 1.098898 | 13.098339 | 1 |
| TTGAC | 1221535 | 0.9579812 | 5.3119082 | 9 |