![[OK]](Icons/tick.png) Basic Statistics
Basic Statistics
| Measure | Value |
|---|---|
| Filename | 140701_JD_B_15_GGCTAC_L004_R2_001.fastq.gz |
| File type | Conventional base calls |
| Encoding | Sanger / Illumina 1.9 |
| Total Sequences | 24559423 |
| Filtered Sequences | 0 |
| Sequence length | 51 |
| %GC | 44 |
![[OK]](Icons/tick.png) Per base sequence quality
Per base sequence quality
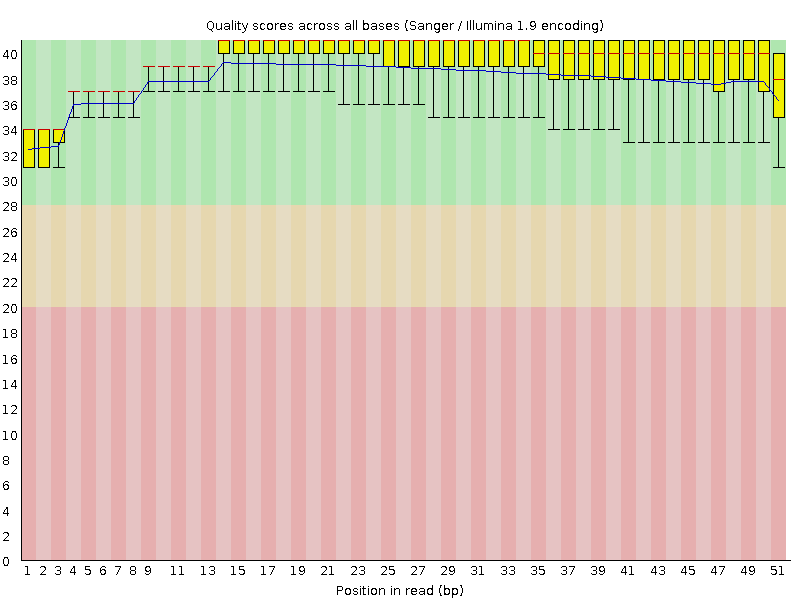
![[OK]](Icons/tick.png) Per sequence quality scores
Per sequence quality scores
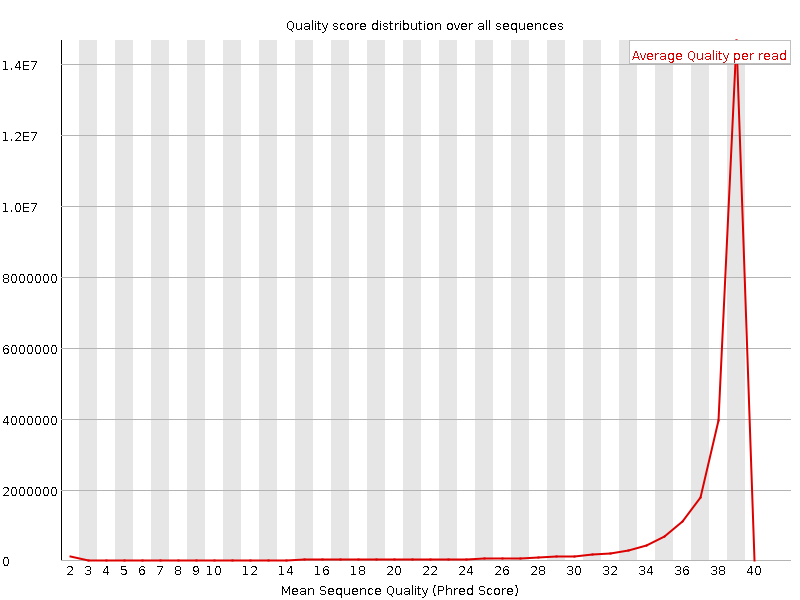
![[WARN]](Icons/warning.png) Per base sequence content
Per base sequence content

![[OK]](Icons/tick.png) Per base GC content
Per base GC content

![[OK]](Icons/tick.png) Per sequence GC content
Per sequence GC content

![[OK]](Icons/tick.png) Per base N content
Per base N content

![[OK]](Icons/tick.png) Sequence Length Distribution
Sequence Length Distribution
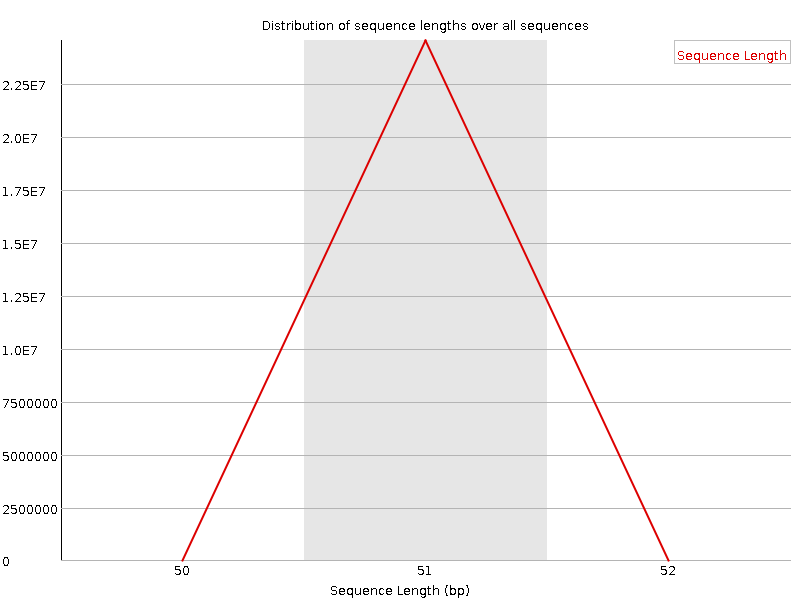
![[WARN]](Icons/warning.png) Sequence Duplication Levels
Sequence Duplication Levels
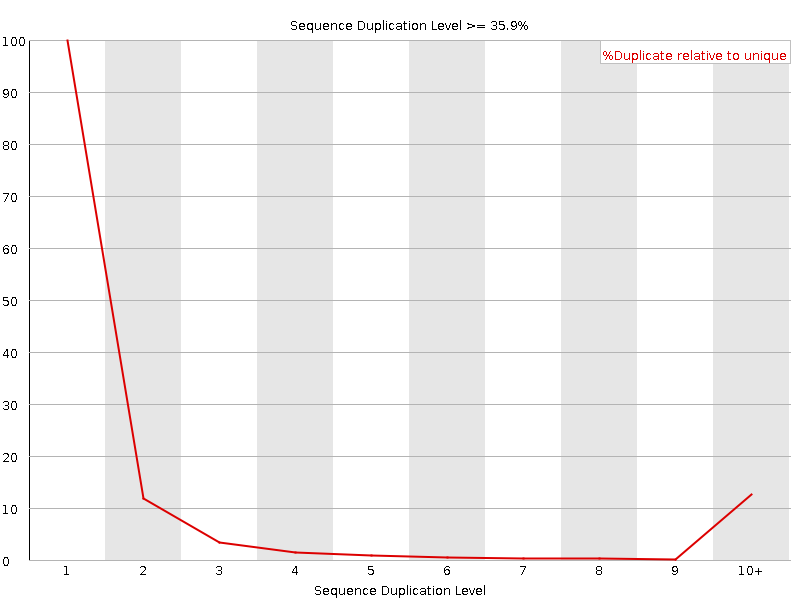
![[WARN]](Icons/warning.png) Overrepresented sequences
Overrepresented sequences
| Sequence | Count | Percentage | Possible Source |
|---|---|---|---|
| NNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNN | 118926 | 0.48423776079755626 | No Hit |
![[WARN]](Icons/warning.png) Kmer Content
Kmer Content
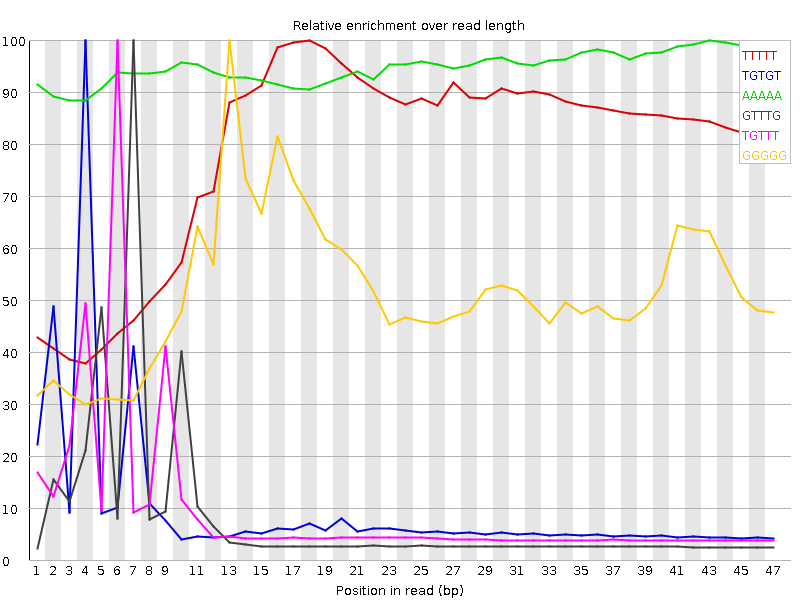
| Sequence | Count | Obs/Exp Overall | Obs/Exp Max | Max Obs/Exp Position |
|---|---|---|---|---|
| TTTTT | 9435680 | 4.4493656 | 5.6326346 | 18 |
| TGTGT | 5340030 | 3.767062 | 38.429535 | 4 |
| AAAAA | 5713950 | 3.6958747 | 3.8930511 | 43 |
| GTTTG | 4373930 | 3.0855381 | 38.14416 | 7 |
| TGTTT | 5166495 | 2.9798062 | 31.670082 | 6 |
| GGGGG | 2264500 | 2.9230256 | 5.6512637 | 13 |
| TTTGA | 4708515 | 2.892858 | 33.444683 | 8 |
| GTGTT | 3951310 | 2.7874057 | 37.979202 | 5 |
| CTTTG | 3439815 | 2.595127 | 40.773064 | 1 |
| TTGTG | 3583580 | 2.5279949 | 37.96055 | 3 |
| GAGGG | 2024680 | 2.2761495 | 11.031519 | 11 |
| TTTGT | 3876515 | 2.2358027 | 31.376093 | 2 |
| TTGAG | 2911665 | 2.188022 | 24.09541 | 9 |
| TGAGG | 2079945 | 1.9117416 | 13.89545 | 10 |
| AACTT | 2403990 | 1.682641 | 15.588832 | 2 |
| ACTTT | 2501095 | 1.6433793 | 14.2037 | 3 |
| AGGGG | 1431120 | 1.6088681 | 5.944777 | 12 |
| TGAGC | 1482500 | 1.4572581 | 5.7169213 | 10 |
| TGAGT | 1651435 | 1.2409999 | 6.963381 | 10 |
| TAACT | 1770460 | 1.2392099 | 15.353084 | 1 |
| TTGAT | 1994100 | 1.2251524 | 7.245362 | 9 |
| TTGAC | 1374805 | 1.1048819 | 7.454972 | 9 |