![[OK]](Icons/tick.png) Basic Statistics
Basic Statistics
| Measure | Value |
|---|---|
| Filename | lane2_Undetermined_L002_R1_001.fastq.gz |
| File type | Conventional base calls |
| Encoding | Sanger / Illumina 1.9 |
| Total Sequences | 5341076 |
| Filtered Sequences | 0 |
| Sequence length | 51 |
| %GC | 44 |
![[OK]](Icons/tick.png) Per base sequence quality
Per base sequence quality
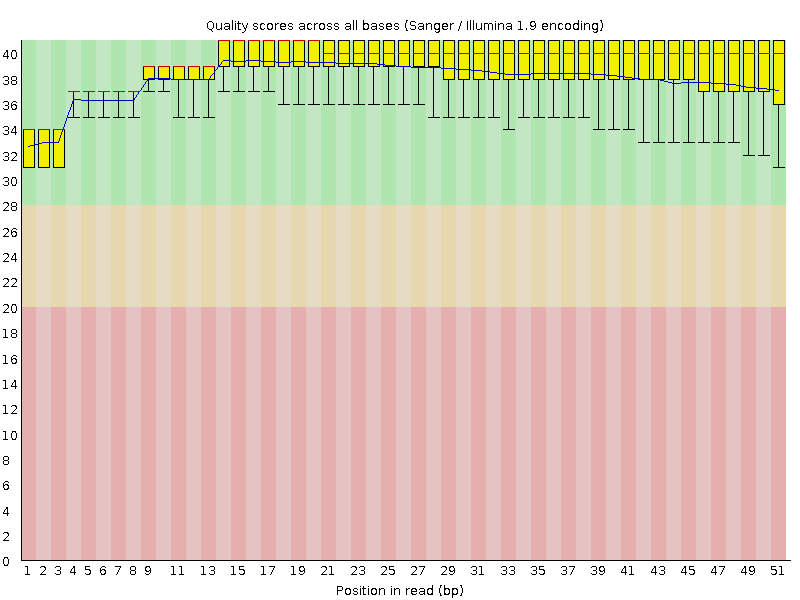
![[OK]](Icons/tick.png) Per sequence quality scores
Per sequence quality scores
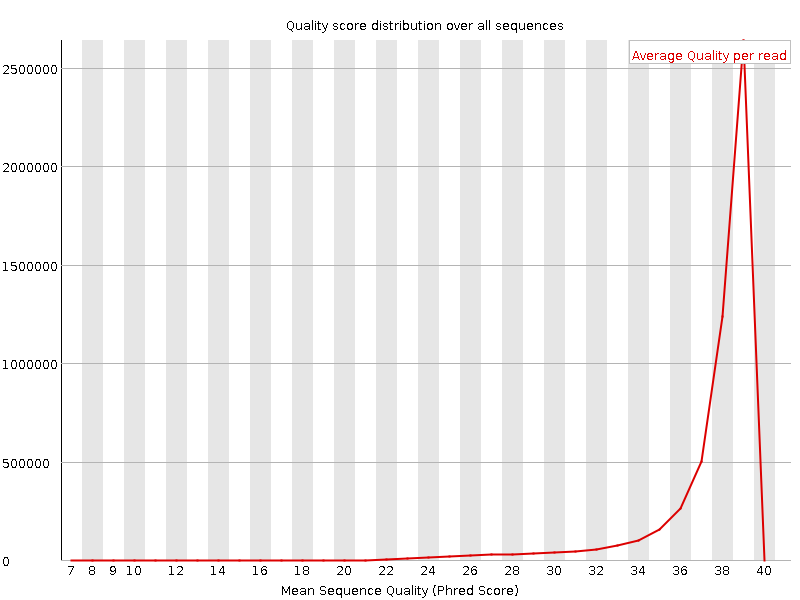
![[WARN]](Icons/warning.png) Per base sequence content
Per base sequence content
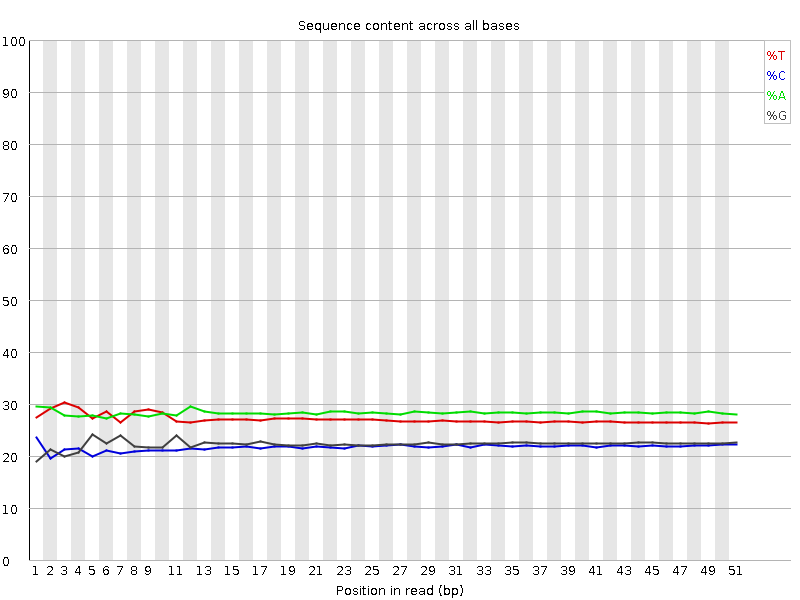
![[OK]](Icons/tick.png) Per base GC content
Per base GC content
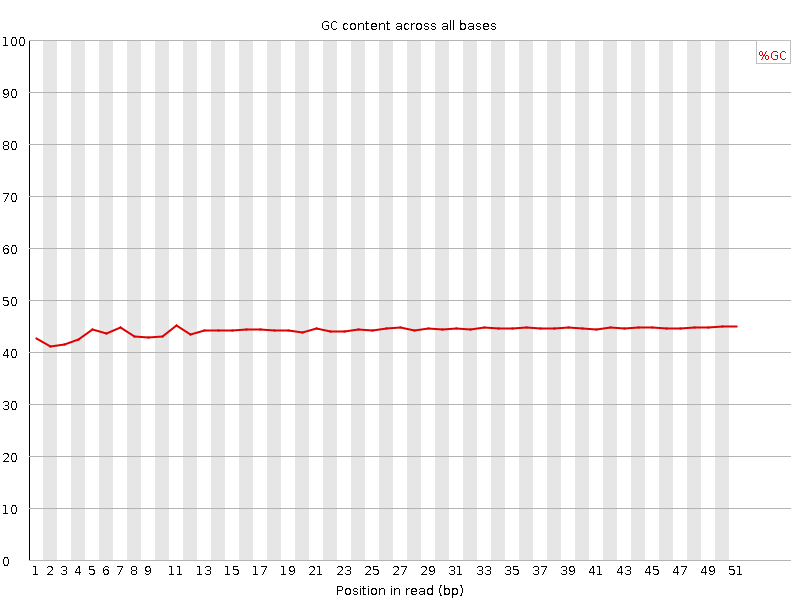
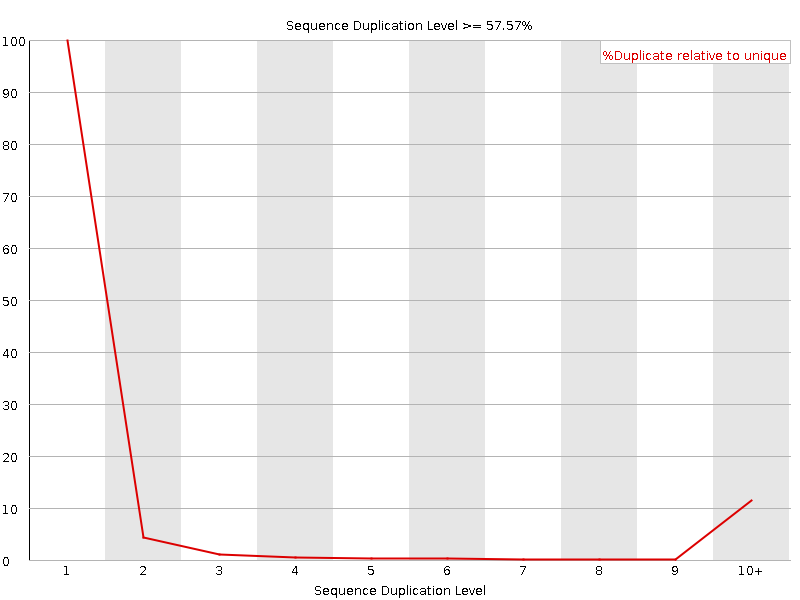
![[WARN]](Icons/warning.png) Per sequence GC content
Per sequence GC content
![[OK]](Icons/tick.png) Per base N content
Per base N content
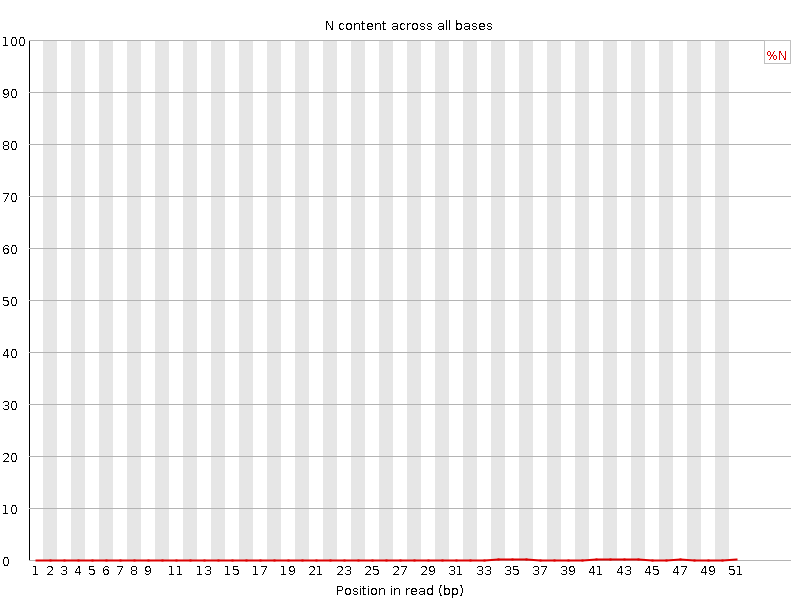
![[OK]](Icons/tick.png) Sequence Length Distribution
Sequence Length Distribution
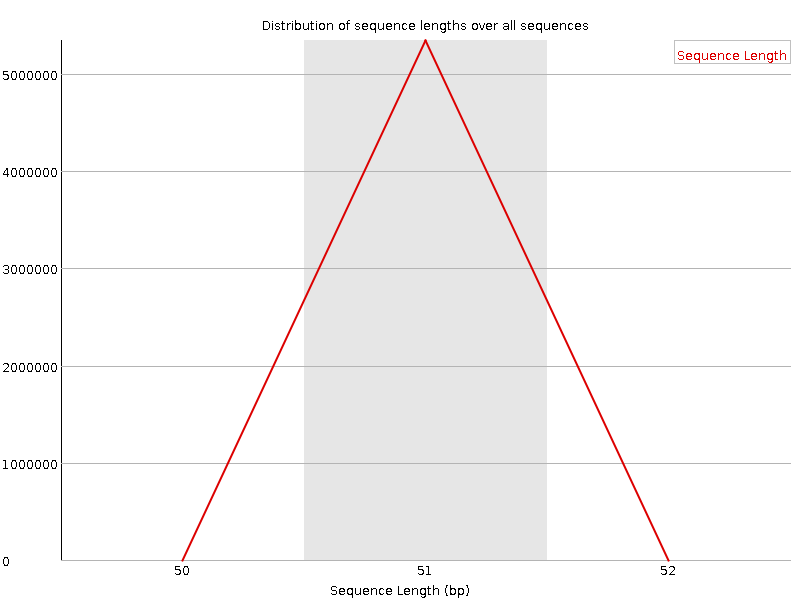
![[FAIL]](Icons/error.png) Sequence Duplication Levels
Sequence Duplication Levels
![[WARN]](Icons/warning.png) Overrepresented sequences
Overrepresented sequences
| Sequence | Count | Percentage | Possible Source |
|---|---|---|---|
| CTTTGTGTTTGATTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTT | 5493 | 0.10284444557613484 | No Hit |
![[WARN]](Icons/warning.png) Kmer Content
Kmer Content
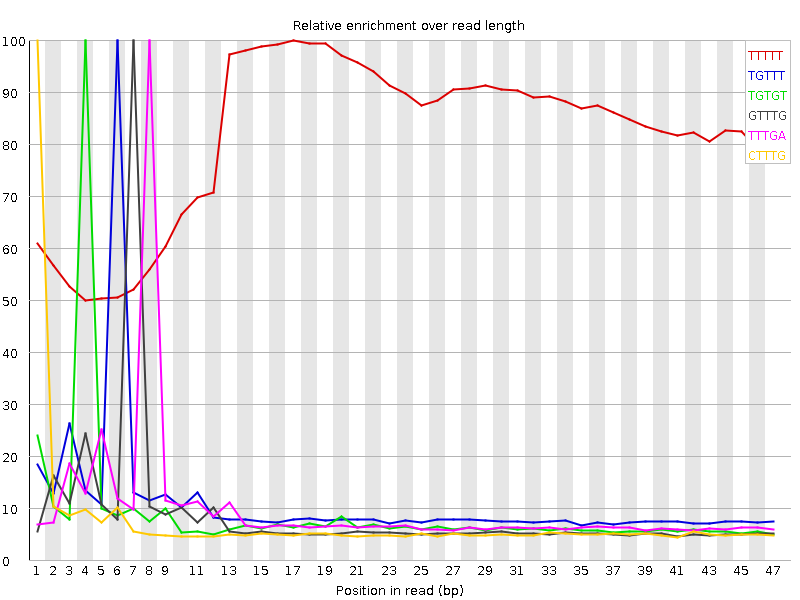
| Sequence | Count | Obs/Exp Overall | Obs/Exp Max | Max Obs/Exp Position |
|---|---|---|---|---|
| TTTTT | 1525150 | 4.0598474 | 4.9719462 | 17 |
| TGTTT | 653950 | 2.1126661 | 19.144915 | 6 |
| TGTGT | 515820 | 2.0224314 | 22.509382 | 4 |
| GTTTG | 499500 | 1.9584439 | 22.543749 | 7 |
| TTTGA | 574155 | 1.7751324 | 17.983385 | 8 |
| CTTTG | 431815 | 1.7379793 | 23.110313 | 1 |
| TTTGT | 523405 | 1.6909246 | 18.805862 | 2 |
| GAGGG | 293140 | 1.6201102 | 5.7184796 | 11 |
| GTGTT | 409905 | 1.6071593 | 22.342625 | 5 |
| TTGAG | 389960 | 1.4632244 | 12.409723 | 9 |
| TTGTG | 364790 | 1.430272 | 22.240696 | 3 |
| TGAGG | 313765 | 1.4288435 | 7.3824105 | 10 |
| TTGAT | 338460 | 1.046427 | 5.0229936 | 9 |