![[OK]](Icons/tick.png) Basic Statistics
Basic Statistics
| Measure | Value |
|---|---|
| Filename | 140701_JD_D_2_ATCACG_L002_R2_001.fastq.gz |
| File type | Conventional base calls |
| Encoding | Sanger / Illumina 1.9 |
| Total Sequences | 40336508 |
| Filtered Sequences | 0 |
| Sequence length | 51 |
| %GC | 43 |
![[OK]](Icons/tick.png) Per base sequence quality
Per base sequence quality
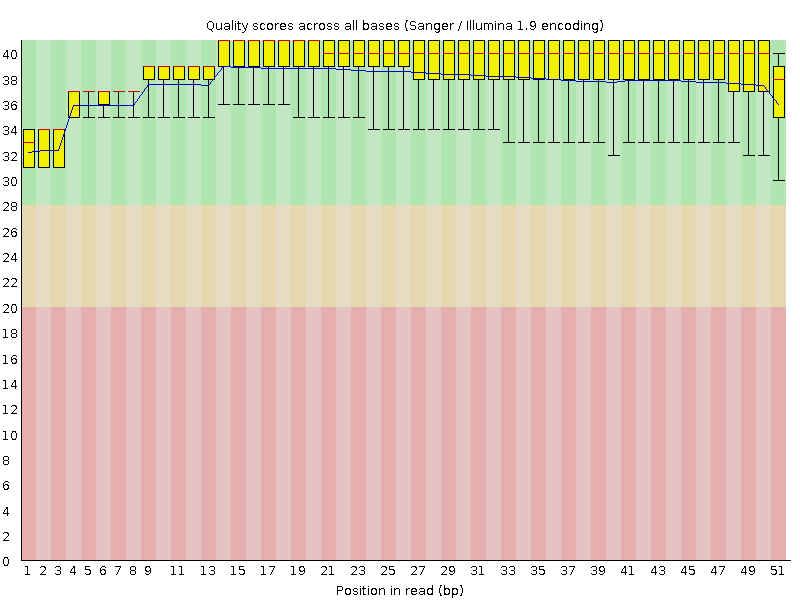
![[OK]](Icons/tick.png) Per sequence quality scores
Per sequence quality scores
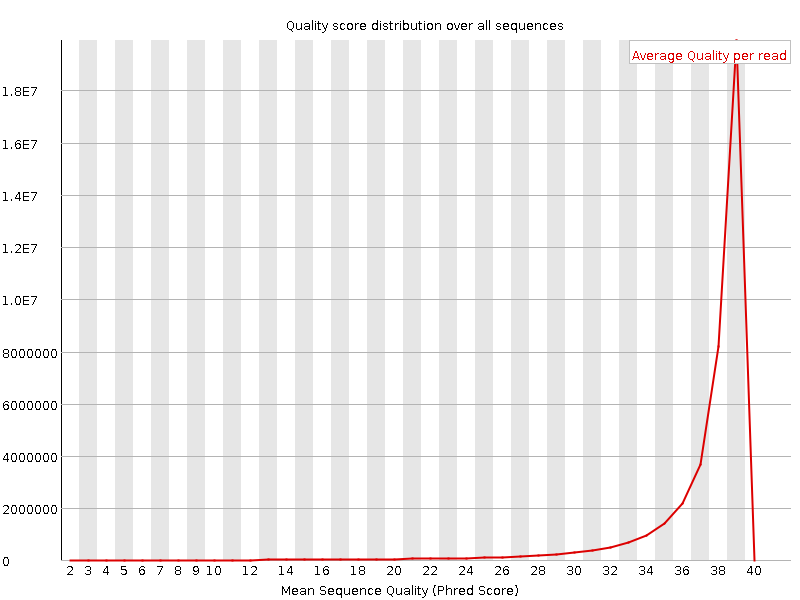
![[WARN]](Icons/warning.png) Per base sequence content
Per base sequence content

![[OK]](Icons/tick.png) Per base GC content
Per base GC content

![[OK]](Icons/tick.png) Per sequence GC content
Per sequence GC content

![[OK]](Icons/tick.png) Per base N content
Per base N content

![[OK]](Icons/tick.png) Sequence Length Distribution
Sequence Length Distribution
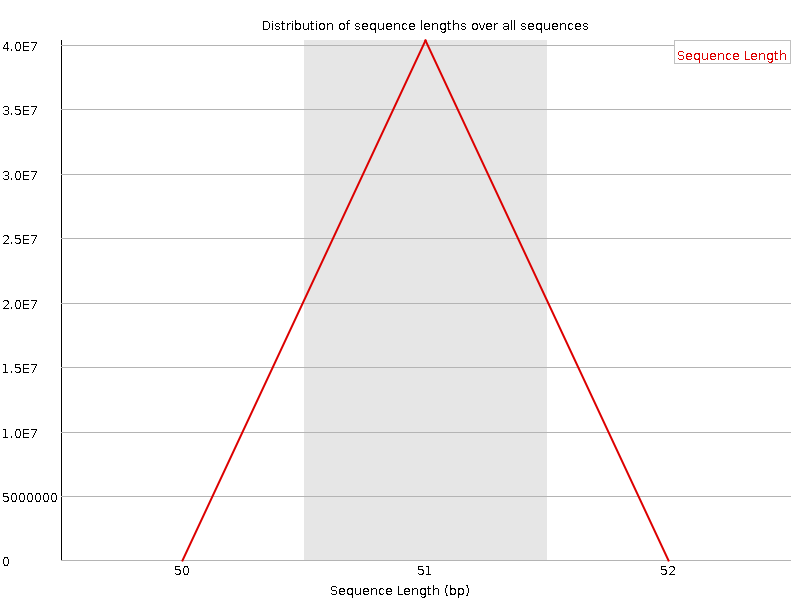
![[WARN]](Icons/warning.png) Sequence Duplication Levels
Sequence Duplication Levels
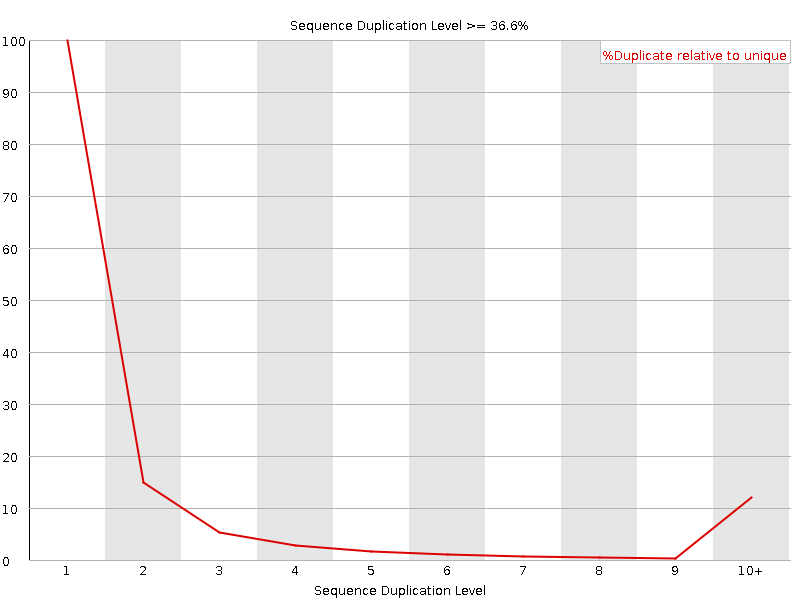
![[OK]](Icons/tick.png) Overrepresented sequences
Overrepresented sequences
No overrepresented sequences
![[WARN]](Icons/warning.png) Kmer Content
Kmer Content
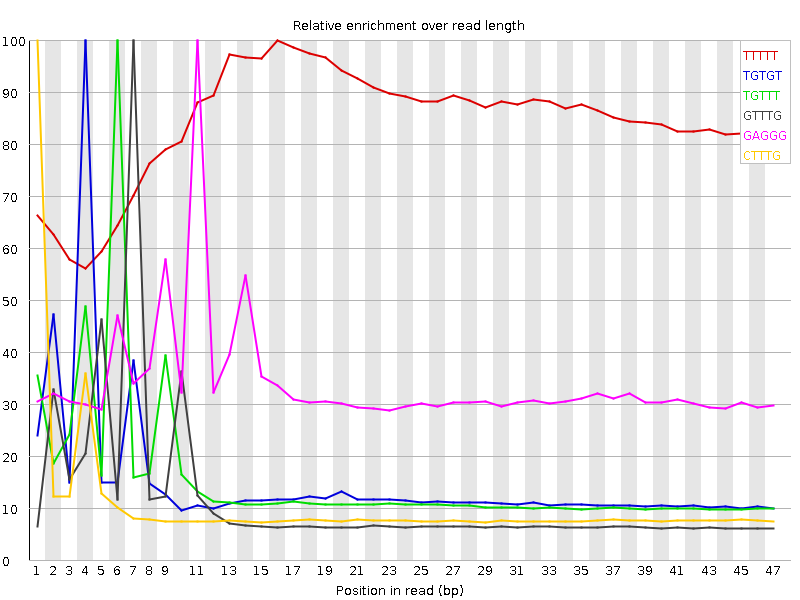
| Sequence | Count | Obs/Exp Overall | Obs/Exp Max | Max Obs/Exp Position |
|---|---|---|---|---|
| TTTTT | 11391775 | 3.273765 | 3.8806098 | 16 |
| TGTGT | 5798815 | 2.7050114 | 18.050505 | 4 |
| TGTTT | 6085240 | 2.2280285 | 14.454892 | 6 |
| GTTTG | 4350960 | 2.0296211 | 17.499365 | 7 |
| GAGGG | 2585460 | 1.9932886 | 5.8576584 | 11 |
| CTTTG | 4043395 | 1.9521725 | 18.324532 | 1 |
| TTTGA | 5043925 | 1.8803573 | 14.281598 | 8 |
| GTGTT | 3826375 | 1.7849144 | 17.412888 | 5 |
| TTTGT | 4808075 | 1.7604115 | 14.1223955 | 2 |
| TTGTG | 3611080 | 1.6844844 | 17.397736 | 3 |
| TGAGG | 2761770 | 1.6712177 | 6.8447704 | 10 |
| TTGAG | 3202600 | 1.5211127 | 10.701814 | 9 |
| AACTT | 3502500 | 1.376008 | 5.6872883 | 2 |
| ACTTT | 3435900 | 1.3257283 | 5.319982 | 3 |
| TAACT | 2395070 | 0.94093806 | 5.351978 | 1 |