![[OK]](Icons/tick.png) Basic Statistics
Basic Statistics
| Measure | Value |
|---|---|
| Filename | 140701_JD_D_2_ATCACG_L002_R1_001.fastq.gz |
| File type | Conventional base calls |
| Encoding | Sanger / Illumina 1.9 |
| Total Sequences | 40336508 |
| Filtered Sequences | 0 |
| Sequence length | 51 |
| %GC | 43 |
![[OK]](Icons/tick.png) Per base sequence quality
Per base sequence quality
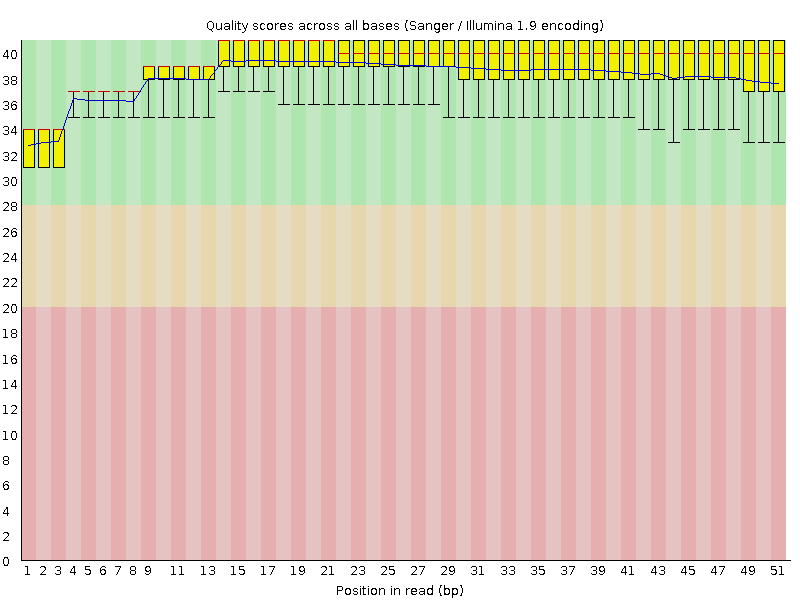
![[OK]](Icons/tick.png) Per sequence quality scores
Per sequence quality scores
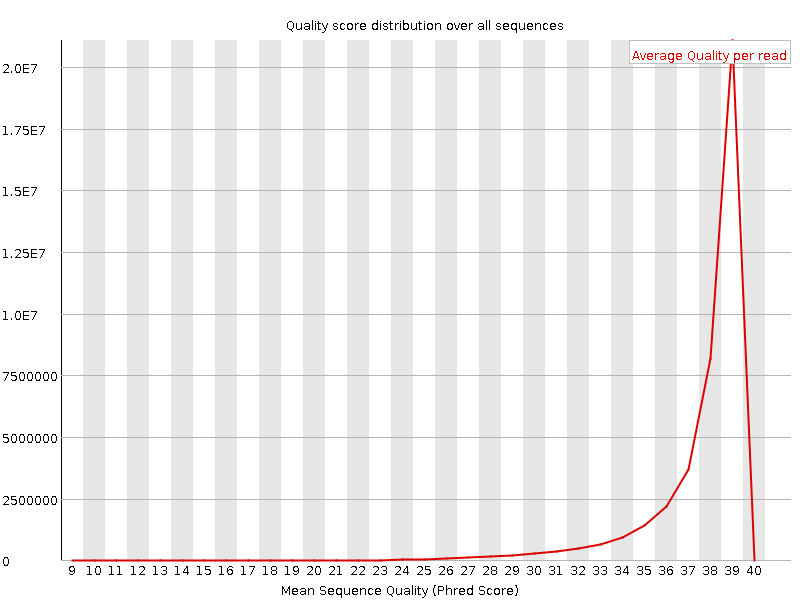
![[WARN]](Icons/warning.png) Per base sequence content
Per base sequence content

![[OK]](Icons/tick.png) Per base GC content
Per base GC content

![[OK]](Icons/tick.png) Per sequence GC content
Per sequence GC content

![[OK]](Icons/tick.png) Per base N content
Per base N content

![[OK]](Icons/tick.png) Sequence Length Distribution
Sequence Length Distribution
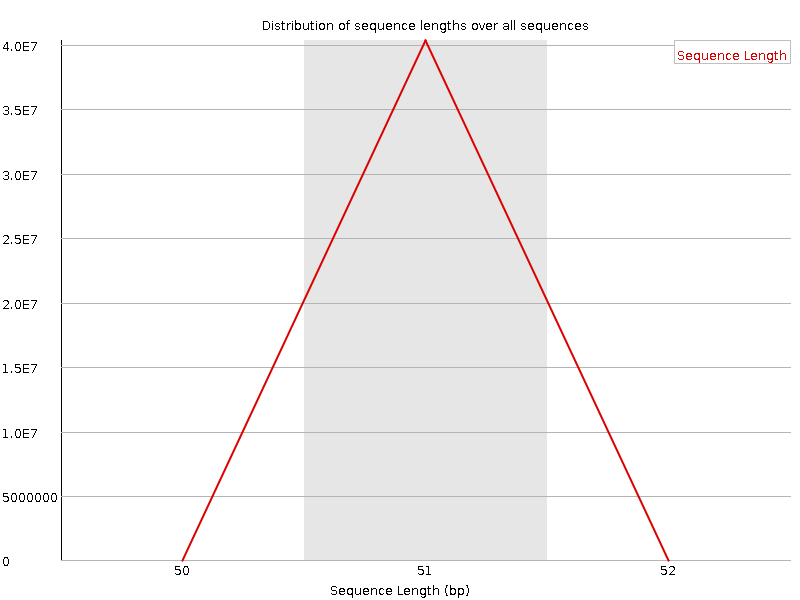
![[WARN]](Icons/warning.png) Sequence Duplication Levels
Sequence Duplication Levels
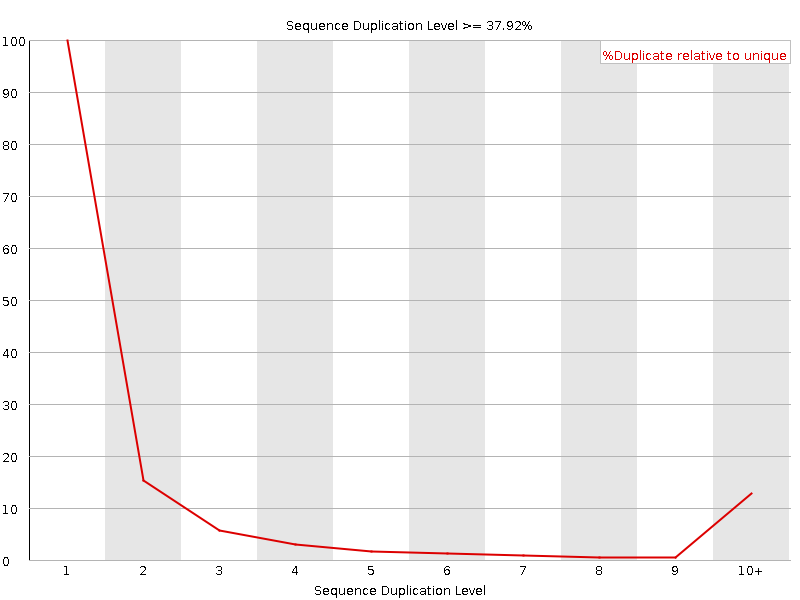
![[OK]](Icons/tick.png) Overrepresented sequences
Overrepresented sequences
No overrepresented sequences
![[WARN]](Icons/warning.png) Kmer Content
Kmer Content
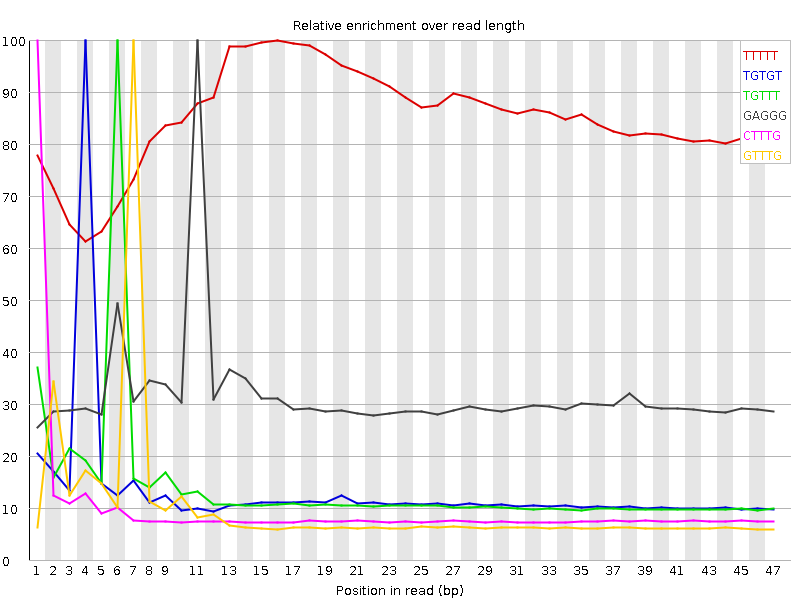
| Sequence | Count | Obs/Exp Overall | Obs/Exp Max | Max Obs/Exp Position |
|---|---|---|---|---|
| TTTTT | 11331025 | 3.1975772 | 3.7580895 | 16 |
| TGTGT | 5210550 | 2.4596438 | 18.5864 | 4 |
| TGTTT | 5575180 | 2.0348346 | 14.628735 | 6 |
| GAGGG | 2405140 | 1.9332016 | 6.0914125 | 11 |
| CTTTG | 3825575 | 1.8437835 | 18.543947 | 1 |
| GTTTG | 3798710 | 1.7931838 | 17.964563 | 7 |
| TTTGT | 4614370 | 1.6841574 | 14.35534 | 2 |
| TTTGA | 4532995 | 1.6840959 | 14.435703 | 8 |
| TGAGG | 2613100 | 1.6239564 | 7.2309923 | 10 |
| GTGTT | 3271880 | 1.5444933 | 17.890717 | 5 |
| TTGTG | 3126605 | 1.4759161 | 17.907974 | 3 |
| TTGAG | 2881810 | 1.3847307 | 10.966075 | 9 |