![[OK]](Icons/tick.png) Basic Statistics
Basic Statistics
| Measure | Value |
|---|---|
| Filename | 140701_JD_D_17_TGACCA_L007_R2_001.fastq.gz |
| File type | Conventional base calls |
| Encoding | Sanger / Illumina 1.9 |
| Total Sequences | 33021066 |
| Filtered Sequences | 0 |
| Sequence length | 51 |
| %GC | 43 |
![[OK]](Icons/tick.png) Per base sequence quality
Per base sequence quality
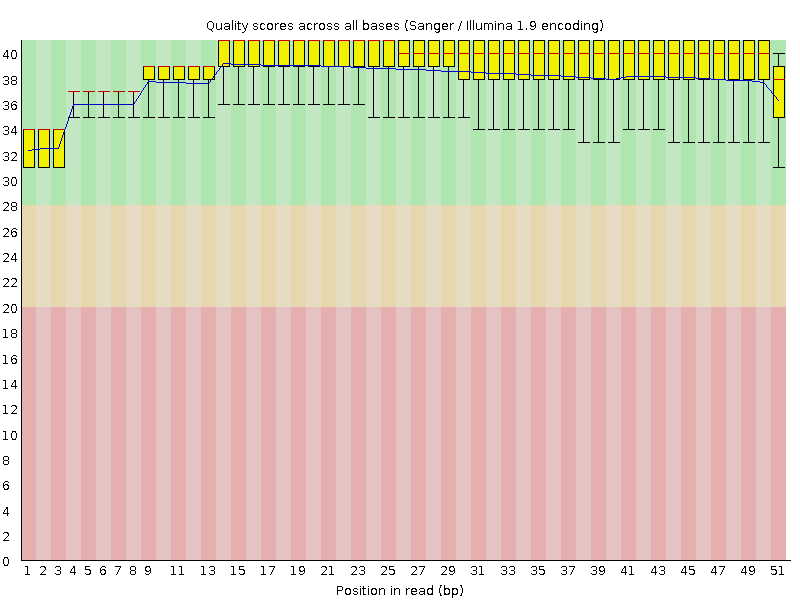
![[OK]](Icons/tick.png) Per sequence quality scores
Per sequence quality scores
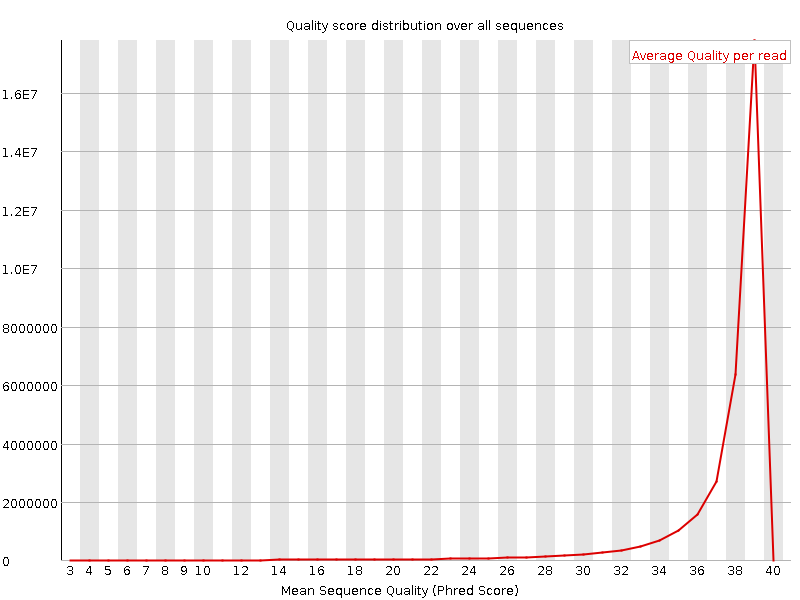
![[WARN]](Icons/warning.png) Per base sequence content
Per base sequence content

![[OK]](Icons/tick.png) Per base GC content
Per base GC content

![[OK]](Icons/tick.png) Per sequence GC content
Per sequence GC content

![[OK]](Icons/tick.png) Per base N content
Per base N content

![[OK]](Icons/tick.png) Sequence Length Distribution
Sequence Length Distribution
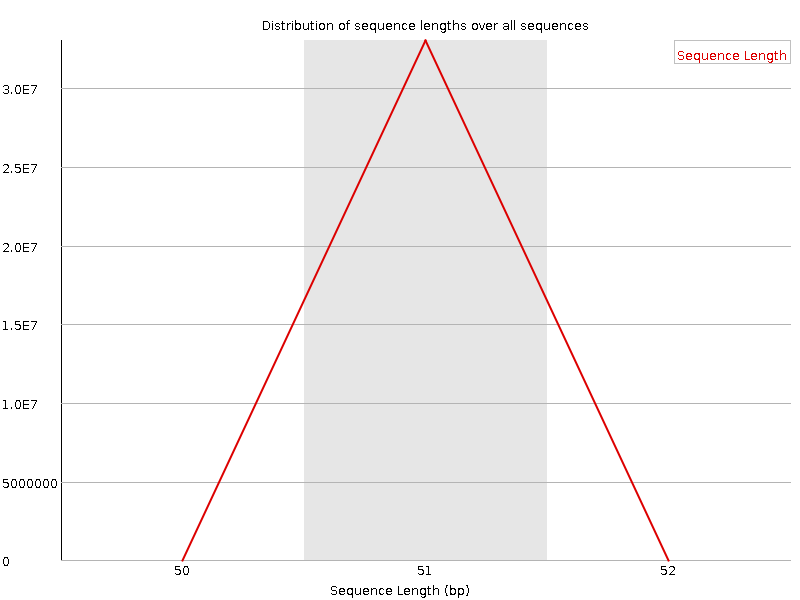
![[WARN]](Icons/warning.png) Sequence Duplication Levels
Sequence Duplication Levels
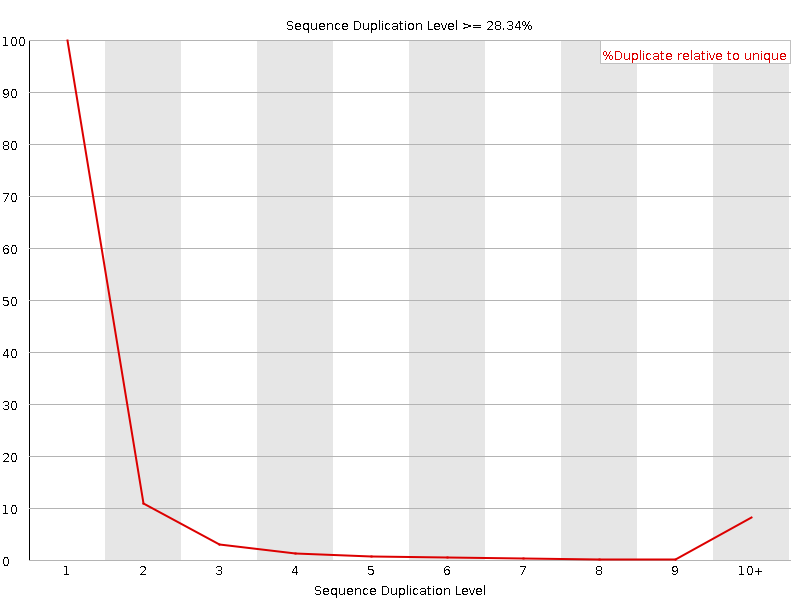
![[OK]](Icons/tick.png) Overrepresented sequences
Overrepresented sequences
No overrepresented sequences
![[WARN]](Icons/warning.png) Kmer Content
Kmer Content
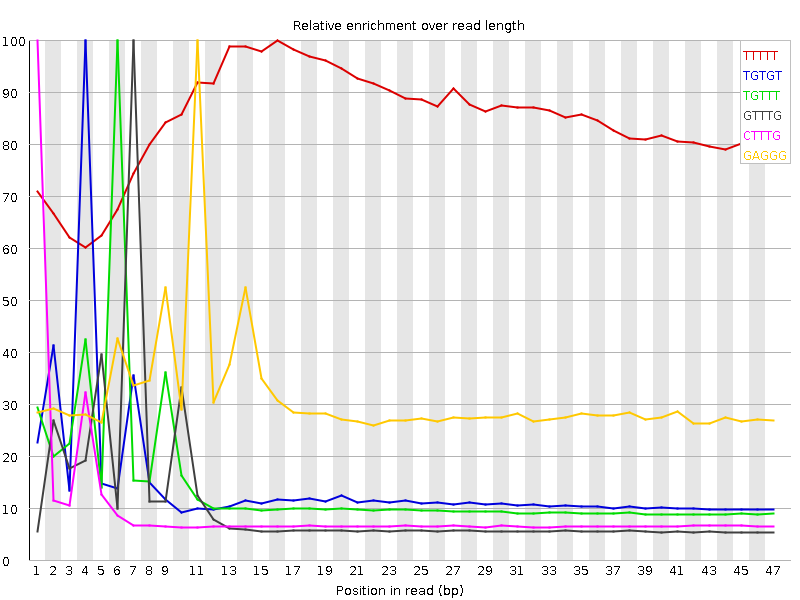
| Sequence | Count | Obs/Exp Overall | Obs/Exp Max | Max Obs/Exp Position |
|---|---|---|---|---|
| TTTTT | 8780345 | 3.019371 | 3.5683928 | 16 |
| TGTGT | 5310660 | 3.0037286 | 20.811852 | 4 |
| TGTTT | 5265120 | 2.3220246 | 16.398996 | 6 |
| GTTTG | 3770435 | 2.132572 | 20.153826 | 7 |
| CTTTG | 3427640 | 2.0181751 | 21.11576 | 1 |
| GAGGG | 2118400 | 2.0067277 | 6.4046197 | 11 |
| TTTGA | 4314450 | 1.9375145 | 16.259836 | 8 |
| GTGTT | 3353045 | 1.8964947 | 20.043062 | 5 |
| TTTGT | 4146155 | 1.8285383 | 16.042376 | 2 |
| TTGTG | 3157355 | 1.7858115 | 20.033112 | 3 |
| TGAGG | 2334835 | 1.7245787 | 7.825873 | 10 |
| TTGAG | 2802500 | 1.6140568 | 12.691837 | 9 |
| AACTT | 2872510 | 1.3673933 | 5.879932 | 2 |
| ACTTT | 2834880 | 1.3252738 | 5.4622545 | 3 |
| TAACT | 1987485 | 0.94609714 | 5.5824256 | 1 |