![[OK]](Icons/tick.png) Basic Statistics
Basic Statistics
| Measure | Value |
|---|---|
| Filename | 140701_JD_C_7_ACTTGA_L004_R1_001.fastq.gz |
| File type | Conventional base calls |
| Encoding | Sanger / Illumina 1.9 |
| Total Sequences | 31548134 |
| Filtered Sequences | 0 |
| Sequence length | 51 |
| %GC | 44 |
![[OK]](Icons/tick.png) Per base sequence quality
Per base sequence quality
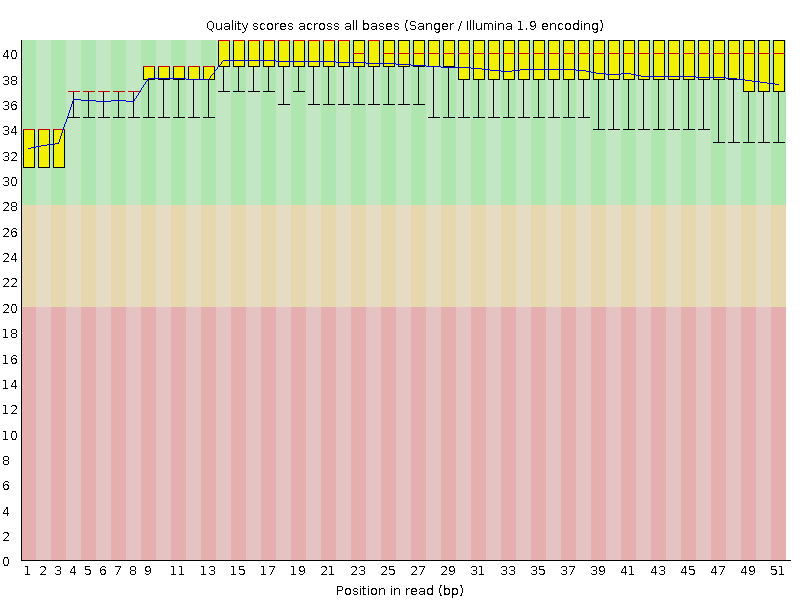
![[OK]](Icons/tick.png) Per sequence quality scores
Per sequence quality scores
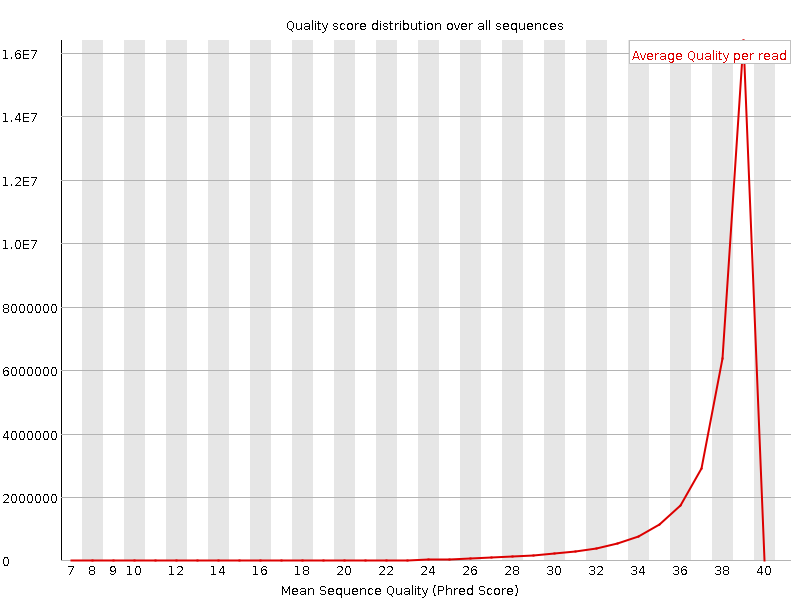
![[WARN]](Icons/warning.png) Per base sequence content
Per base sequence content

![[OK]](Icons/tick.png) Per base GC content
Per base GC content

![[OK]](Icons/tick.png) Per sequence GC content
Per sequence GC content

![[OK]](Icons/tick.png) Per base N content
Per base N content

![[OK]](Icons/tick.png) Sequence Length Distribution
Sequence Length Distribution
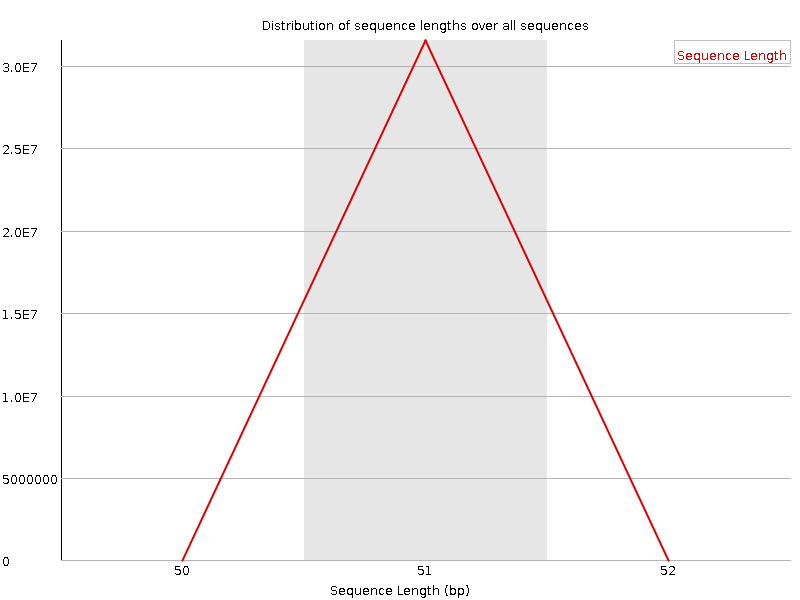
![[WARN]](Icons/warning.png) Sequence Duplication Levels
Sequence Duplication Levels
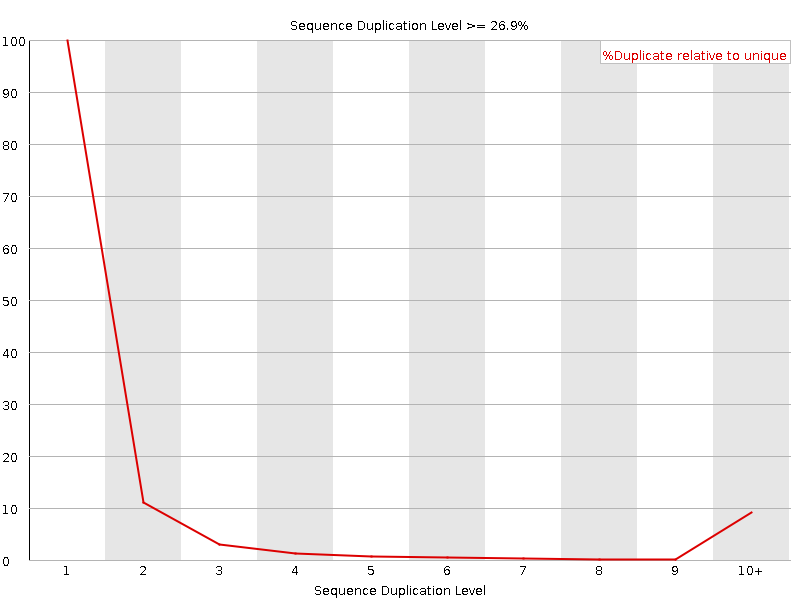
![[WARN]](Icons/warning.png) Overrepresented sequences
Overrepresented sequences
| Sequence | Count | Percentage | Possible Source |
|---|---|---|---|
| CTTTGTGTTTGATTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTT | 35120 | 0.11132195647450972 | No Hit |
![[WARN]](Icons/warning.png) Kmer Content
Kmer Content
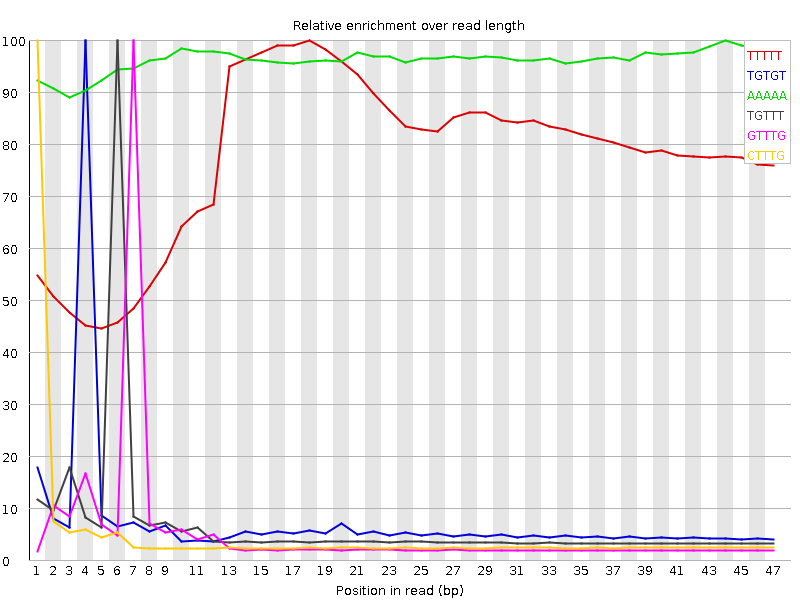
| Sequence | Count | Obs/Exp Overall | Obs/Exp Max | Max Obs/Exp Position |
|---|---|---|---|---|
| TTTTT | 11545230 | 4.0949516 | 5.273705 | 18 |
| TGTGT | 6884105 | 3.8547661 | 51.68089 | 4 |
| AAAAA | 7235455 | 3.3046505 | 3.4307594 | 44 |
| TGTTT | 6200060 | 2.7630851 | 41.081474 | 6 |
| GTTTG | 4849575 | 2.7155275 | 50.83543 | 7 |
| CTTTG | 4477120 | 2.6451914 | 53.70006 | 1 |
| GTGTT | 4548770 | 2.5470915 | 50.843197 | 5 |
| TTTGA | 5363010 | 2.5140254 | 42.81578 | 8 |
| TTGTG | 4163185 | 2.3311825 | 50.88441 | 3 |
| TTTGT | 5175940 | 2.3066814 | 40.798355 | 2 |
| GAGGG | 2324885 | 2.161818 | 13.765897 | 11 |
| GGGGG | 1920225 | 2.1328447 | 5.488521 | 13 |
| GAGAG | 2662265 | 2.0724277 | 5.036227 | 11 |
| TTGAG | 3367450 | 1.9834186 | 31.369003 | 9 |
| TGAGG | 2615525 | 1.9356388 | 18.572397 | 10 |
| AGGGG | 1703465 | 1.5839844 | 7.4644938 | 12 |
| TGAGA | 2349885 | 1.4558699 | 5.815376 | 10 |
| TGAGC | 1855510 | 1.4488937 | 6.871154 | 10 |
| GAGTG | 1731195 | 1.2811838 | 6.9969516 | 11 |
| TGAGT | 2163825 | 1.274487 | 8.929217 | 10 |
| TTGAT | 2364230 | 1.1082833 | 9.590013 | 9 |
| TTGAC | 1556655 | 0.96741635 | 8.717339 | 9 |