![[OK]](Icons/tick.png) Basic Statistics
Basic Statistics
| Measure | Value |
|---|---|
| Filename | 140701_JD_C_18_CTTGTA_L005_R2_001.fastq.gz |
| File type | Conventional base calls |
| Encoding | Sanger / Illumina 1.9 |
| Total Sequences | 28842441 |
| Filtered Sequences | 0 |
| Sequence length | 51 |
| %GC | 43 |
![[OK]](Icons/tick.png) Per base sequence quality
Per base sequence quality
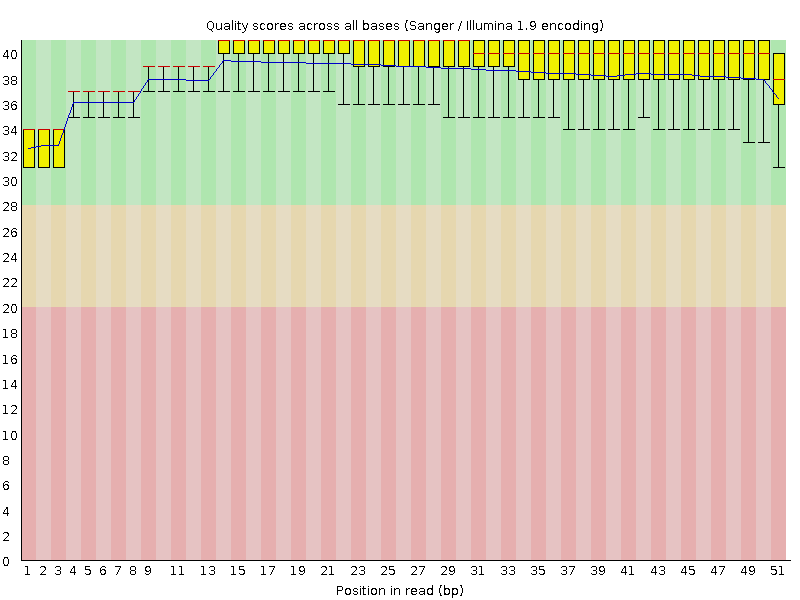
![[OK]](Icons/tick.png) Per sequence quality scores
Per sequence quality scores
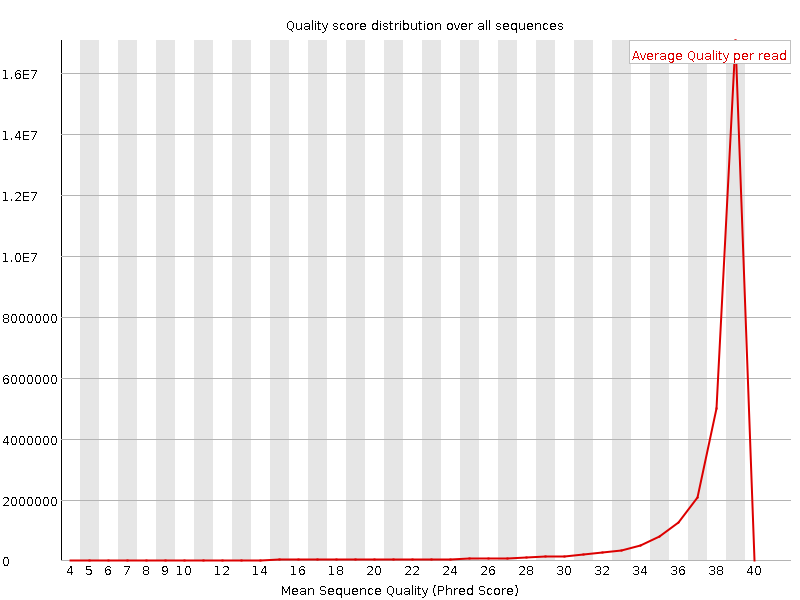
![[WARN]](Icons/warning.png) Per base sequence content
Per base sequence content

![[OK]](Icons/tick.png) Per base GC content
Per base GC content

![[OK]](Icons/tick.png) Per sequence GC content
Per sequence GC content

![[OK]](Icons/tick.png) Per base N content
Per base N content

![[OK]](Icons/tick.png) Sequence Length Distribution
Sequence Length Distribution
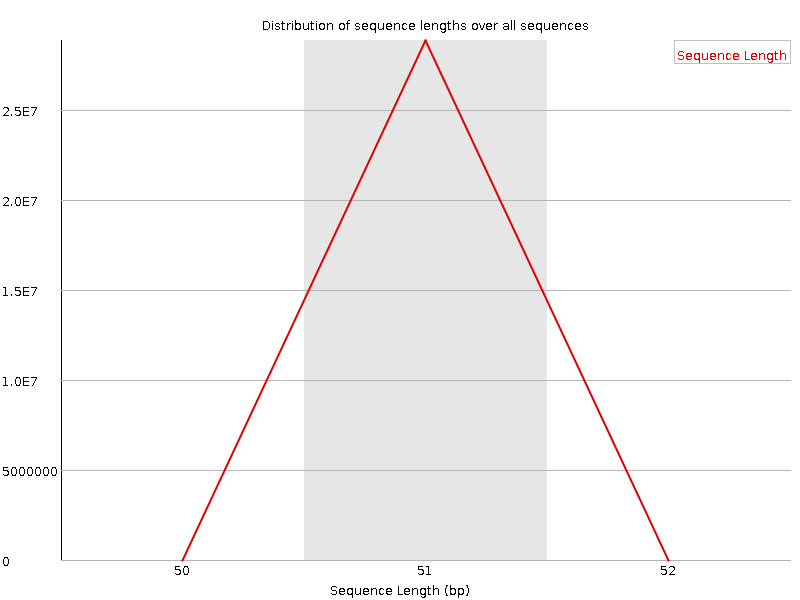
![[WARN]](Icons/warning.png) Sequence Duplication Levels
Sequence Duplication Levels
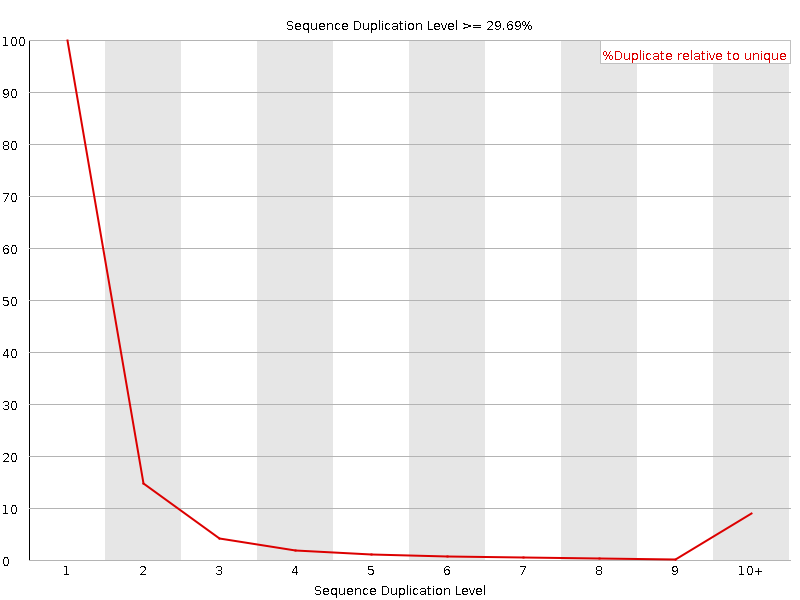
![[WARN]](Icons/warning.png) Overrepresented sequences
Overrepresented sequences
| Sequence | Count | Percentage | Possible Source |
|---|---|---|---|
| CTTTGTGTTTGATTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTT | 32565 | 0.11290653242560156 | No Hit |
![[WARN]](Icons/warning.png) Kmer Content
Kmer Content
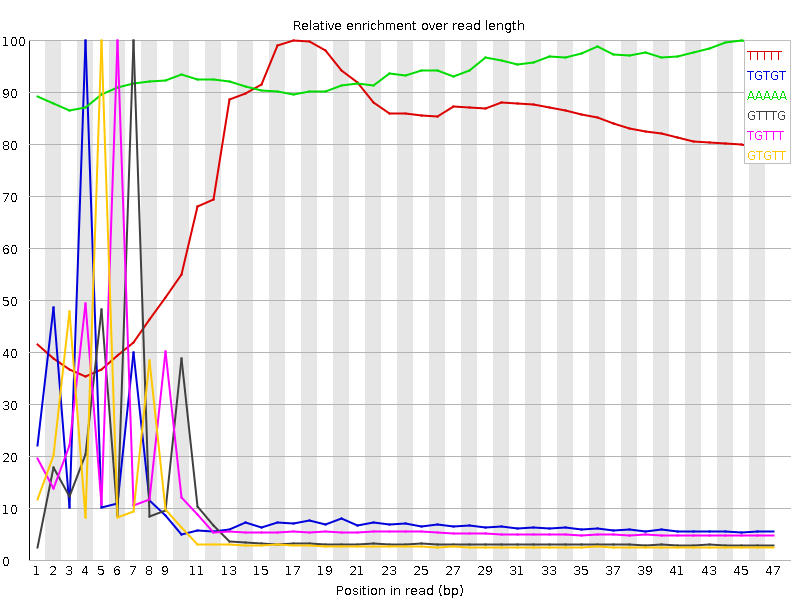
| Sequence | Count | Obs/Exp Overall | Obs/Exp Max | Max Obs/Exp Position |
|---|---|---|---|---|
| TTTTT | 13805935 | 4.73674 | 6.165843 | 17 |
| TGTGT | 6437265 | 3.7986817 | 35.283005 | 4 |
| AAAAA | 8080220 | 3.7302437 | 3.9706059 | 45 |
| GTTTG | 4968680 | 2.9320579 | 34.777077 | 7 |
| TGTTT | 6275370 | 2.823657 | 27.137852 | 6 |
| GTGTT | 4547815 | 2.6837022 | 34.679607 | 5 |
| GGGGG | 1977570 | 2.6323283 | 5.5611587 | 13 |
| TTTGA | 5394395 | 2.5757008 | 28.309273 | 8 |
| CTTTG | 3969615 | 2.5132213 | 37.304142 | 1 |
| TTGTG | 4225765 | 2.4936578 | 34.69292 | 3 |
| GAGGG | 2094790 | 2.2561595 | 10.6581 | 11 |
| TTTGT | 4752020 | 2.1382124 | 26.81942 | 2 |
| TTGAG | 3355470 | 2.1011882 | 21.712708 | 9 |
| TGAGG | 2460345 | 2.0205355 | 13.240094 | 10 |
| AGGGG | 1585100 | 1.7072061 | 5.9497776 | 12 |
| TGAGC | 1710270 | 1.5069054 | 5.563806 | 10 |
| AACTT | 2739640 | 1.4892826 | 12.715404 | 2 |
| ACTTT | 2851375 | 1.4606894 | 11.625981 | 3 |
| TGAGT | 2036170 | 1.2750454 | 6.584356 | 10 |
| GAGTG | 1527050 | 1.2540755 | 5.23143 | 11 |
| TTGAT | 2384710 | 1.1386448 | 7.0603747 | 9 |
| TAACT | 2002380 | 1.0885041 | 12.459257 | 1 |
| TTGAC | 1466445 | 0.9852085 | 5.6442294 | 9 |