![[OK]](Icons/tick.png) Basic Statistics
Basic Statistics
| Measure | Value |
|---|---|
| Filename | 140701_JD_C_18_CTTGTA_L005_R1_001.fastq.gz |
| File type | Conventional base calls |
| Encoding | Sanger / Illumina 1.9 |
| Total Sequences | 28842441 |
| Filtered Sequences | 0 |
| Sequence length | 51 |
| %GC | 43 |
![[OK]](Icons/tick.png) Per base sequence quality
Per base sequence quality
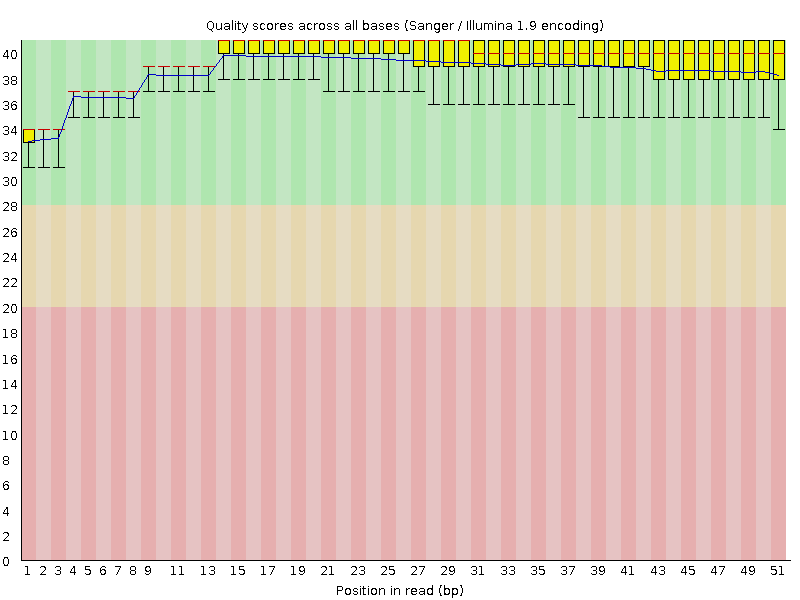
![[OK]](Icons/tick.png) Per sequence quality scores
Per sequence quality scores
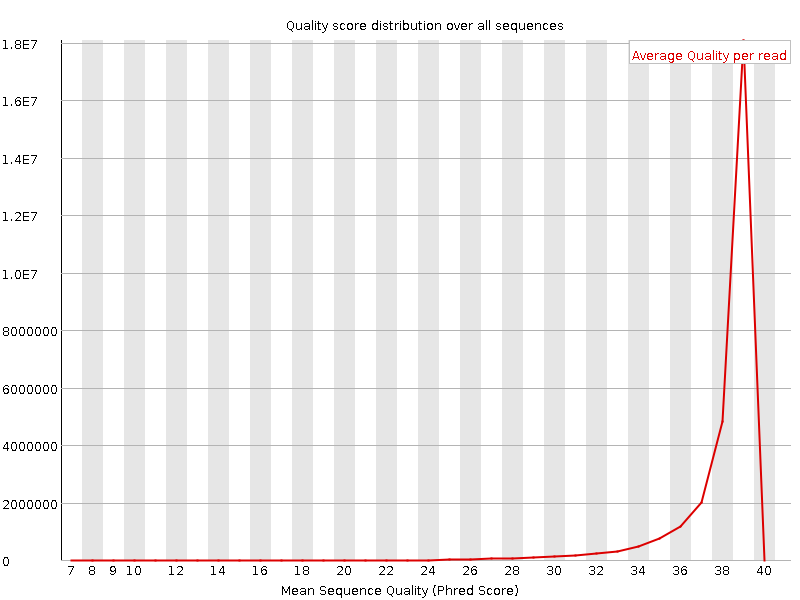
![[WARN]](Icons/warning.png) Per base sequence content
Per base sequence content

![[OK]](Icons/tick.png) Per base GC content
Per base GC content

![[OK]](Icons/tick.png) Per sequence GC content
Per sequence GC content

![[OK]](Icons/tick.png) Per base N content
Per base N content

![[OK]](Icons/tick.png) Sequence Length Distribution
Sequence Length Distribution
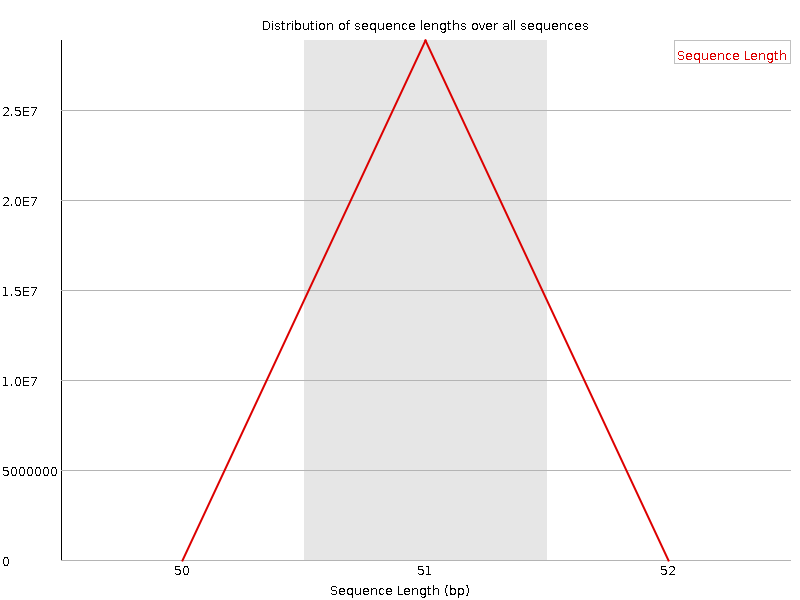
![[WARN]](Icons/warning.png) Sequence Duplication Levels
Sequence Duplication Levels
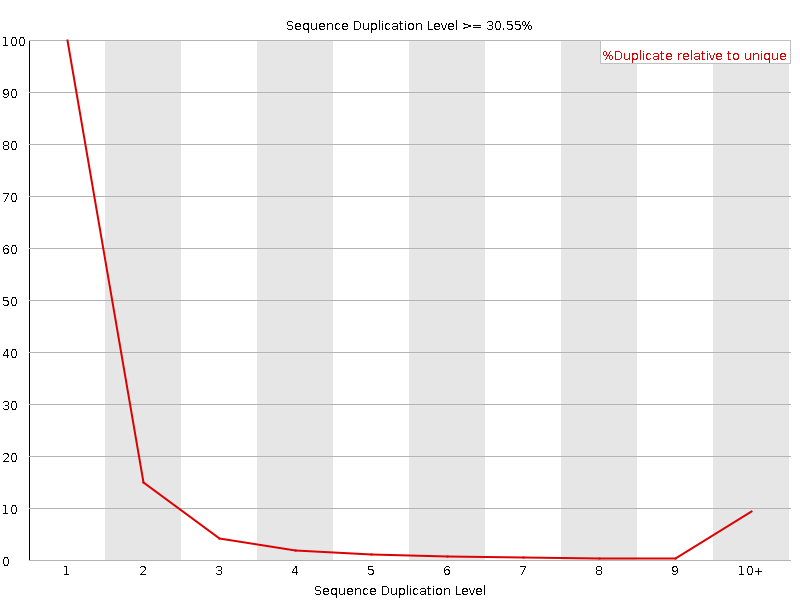
![[WARN]](Icons/warning.png) Overrepresented sequences
Overrepresented sequences
| Sequence | Count | Percentage | Possible Source |
|---|---|---|---|
| CTTTGTGTTTGATTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTT | 37393 | 0.129645753630908 | No Hit |
![[WARN]](Icons/warning.png) Kmer Content
Kmer Content
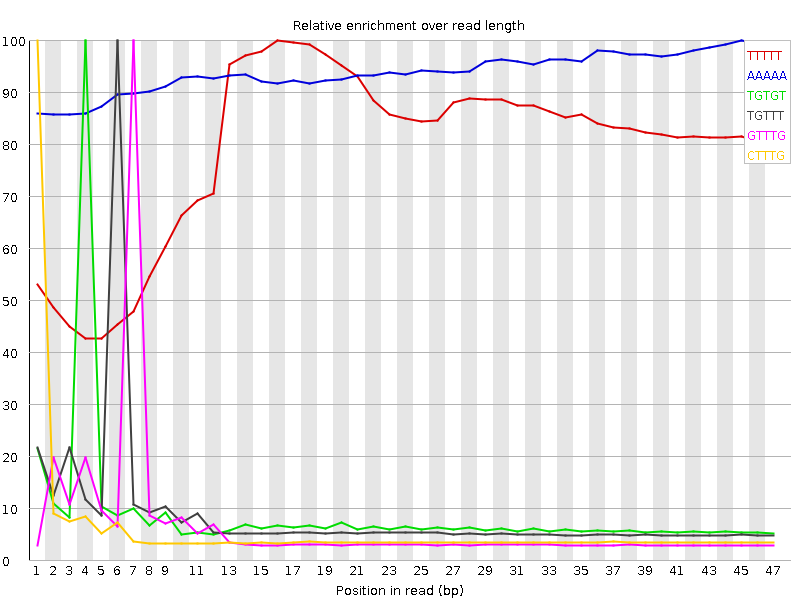
| Sequence | Count | Obs/Exp Overall | Obs/Exp Max | Max Obs/Exp Position |
|---|---|---|---|---|
| TTTTT | 12447790 | 4.4490943 | 5.617117 | 16 |
| AAAAA | 7851910 | 3.520836 | 3.7482657 | 45 |
| TGTGT | 5284815 | 3.258982 | 37.001495 | 4 |
| TGTTT | 5268855 | 2.4736145 | 28.33369 | 6 |
| GTTTG | 3943545 | 2.431862 | 36.32427 | 7 |
| CTTTG | 3526970 | 2.2778416 | 38.010525 | 1 |
| GTGTT | 3508740 | 2.1637313 | 36.268337 | 5 |
| TTTGA | 4399265 | 2.1611958 | 29.176056 | 8 |
| GAGGG | 1865135 | 2.0765073 | 11.029819 | 11 |
| TTTGT | 4266735 | 2.0031407 | 27.995132 | 2 |
| TTGTG | 3238635 | 1.997166 | 36.282413 | 3 |
| TGAGG | 2213125 | 1.8758268 | 14.002687 | 10 |
| TTGAG | 2774655 | 1.7904384 | 22.523243 | 9 |
| AGGGG | 1405990 | 1.5653281 | 6.147324 | 12 |
| TGAGC | 1643635 | 1.4590218 | 5.521914 | 10 |
| TGAGT | 1896040 | 1.2234828 | 6.7855687 | 10 |
| GAGTG | 1441410 | 1.2217275 | 5.4091654 | 11 |
| TTGAT | 2163650 | 1.0629212 | 7.3337665 | 9 |
| TTGAC | 1353160 | 0.91446936 | 5.616698 | 9 |