![[OK]](Icons/tick.png) Basic Statistics
Basic Statistics
| Measure | Value |
|---|---|
| Filename | 140701_JD_C_17_CAGATC_L004_R2_001.fastq.gz |
| File type | Conventional base calls |
| Encoding | Sanger / Illumina 1.9 |
| Total Sequences | 45160315 |
| Filtered Sequences | 0 |
| Sequence length | 51 |
| %GC | 43 |
![[OK]](Icons/tick.png) Per base sequence quality
Per base sequence quality
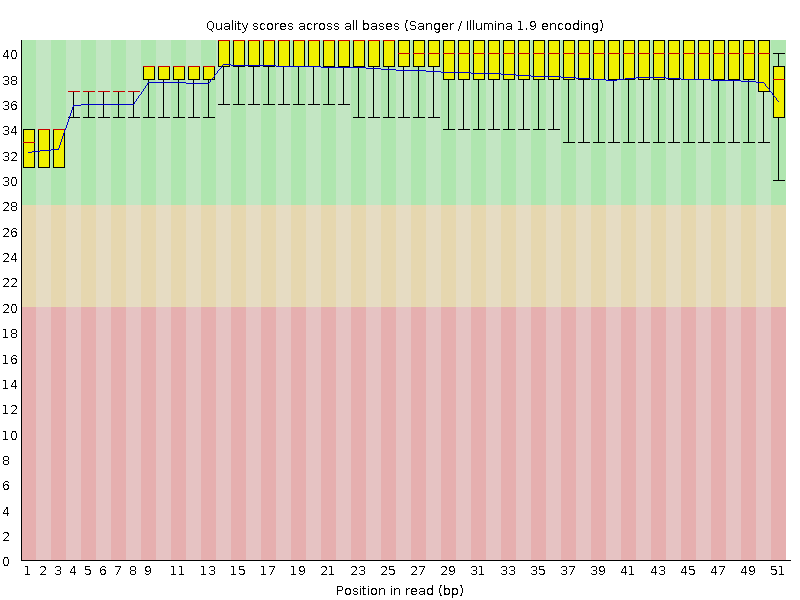
![[OK]](Icons/tick.png) Per sequence quality scores
Per sequence quality scores
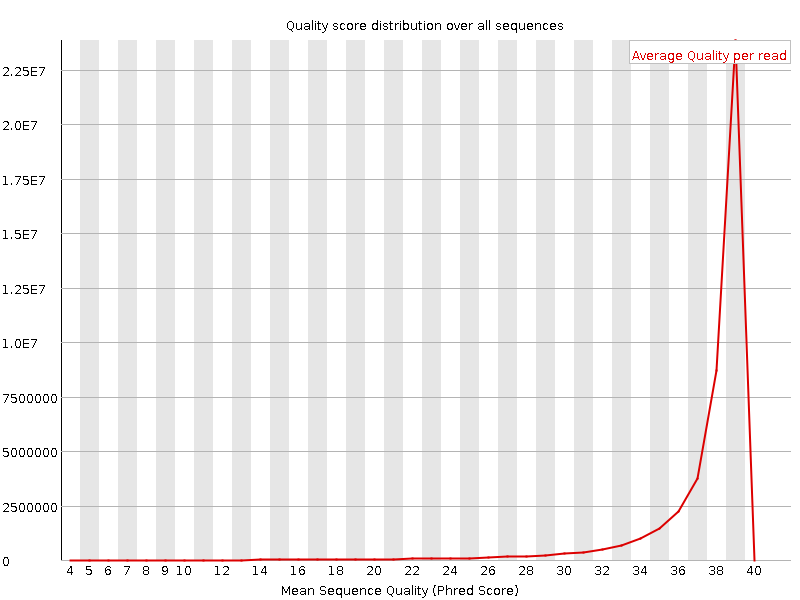
![[WARN]](Icons/warning.png) Per base sequence content
Per base sequence content

![[OK]](Icons/tick.png) Per base GC content
Per base GC content

![[OK]](Icons/tick.png) Per sequence GC content
Per sequence GC content

![[OK]](Icons/tick.png) Per base N content
Per base N content

![[OK]](Icons/tick.png) Sequence Length Distribution
Sequence Length Distribution
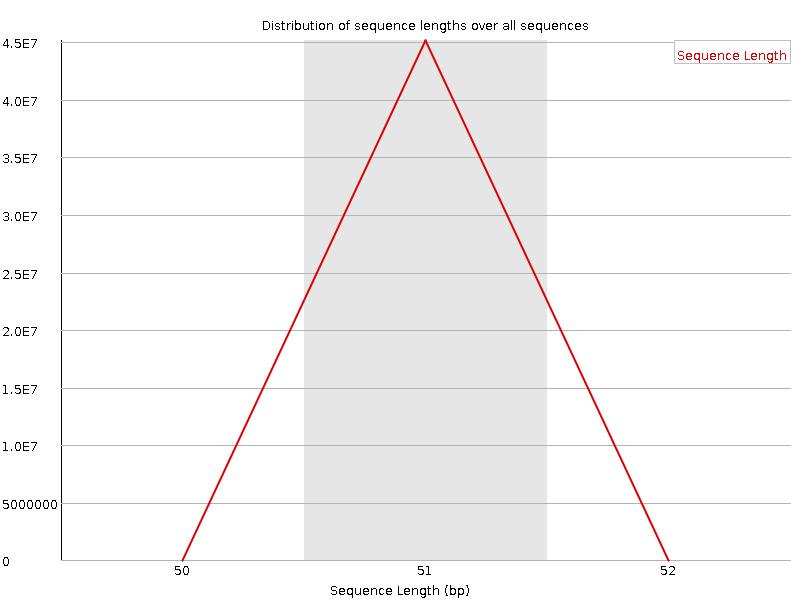
![[WARN]](Icons/warning.png) Sequence Duplication Levels
Sequence Duplication Levels
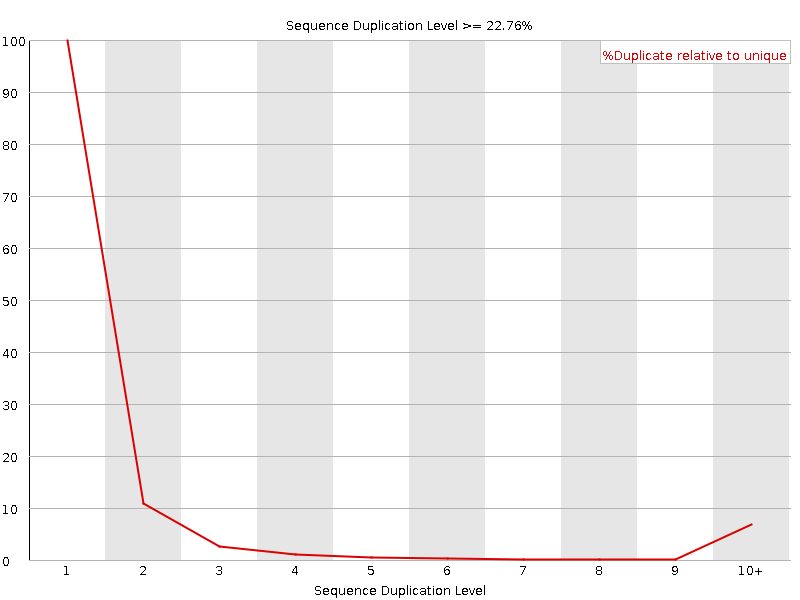
![[OK]](Icons/tick.png) Overrepresented sequences
Overrepresented sequences
No overrepresented sequences
![[FAIL]](Icons/error.png) Kmer Content
Kmer Content
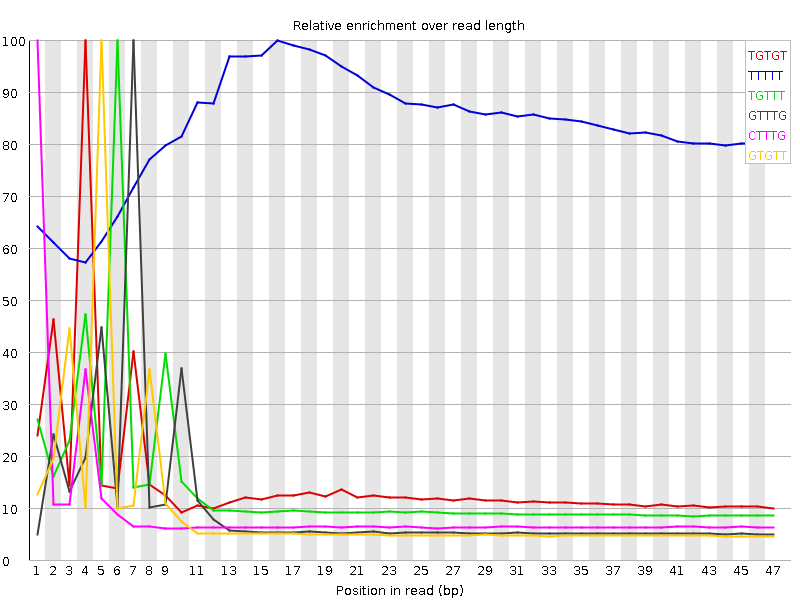
| Sequence | Count | Obs/Exp Overall | Obs/Exp Max | Max Obs/Exp Position |
|---|---|---|---|---|
| TGTGT | 8107525 | 3.3358748 | 21.935785 | 4 |
| TTTTT | 12196610 | 3.079676 | 3.6918783 | 16 |
| TGTTT | 7497360 | 2.4165843 | 17.371939 | 6 |
| GTTTG | 5316140 | 2.1873474 | 21.228823 | 7 |
| CTTTG | 4921595 | 2.1113474 | 22.290337 | 1 |
| GTGTT | 4902320 | 2.0170798 | 21.144596 | 5 |
| GAGGG | 2938455 | 2.0120118 | 6.766924 | 11 |
| TTTGA | 6102815 | 2.0089004 | 17.222536 | 8 |
| TTGTG | 4705560 | 1.9361222 | 21.193909 | 3 |
| TTTGT | 5984995 | 1.929112 | 17.034336 | 2 |
| TGAGG | 3346005 | 1.7947758 | 8.3294525 | 10 |
| TTGAG | 3981635 | 1.6730825 | 13.247025 | 9 |
| ACTTT | 3905375 | 1.3403658 | 6.585548 | 3 |
| AACTT | 3601425 | 1.2623202 | 6.88298 | 2 |
| TAACT | 2681580 | 0.93990916 | 6.6448364 | 1 |