![[OK]](Icons/tick.png) Basic Statistics
Basic Statistics
| Measure | Value |
|---|---|
| Filename | 140701_JD_C_16_TGACCA_L004_R2_001.fastq.gz |
| File type | Conventional base calls |
| Encoding | Sanger / Illumina 1.9 |
| Total Sequences | 29390367 |
| Filtered Sequences | 0 |
| Sequence length | 51 |
| %GC | 42 |
![[OK]](Icons/tick.png) Per base sequence quality
Per base sequence quality
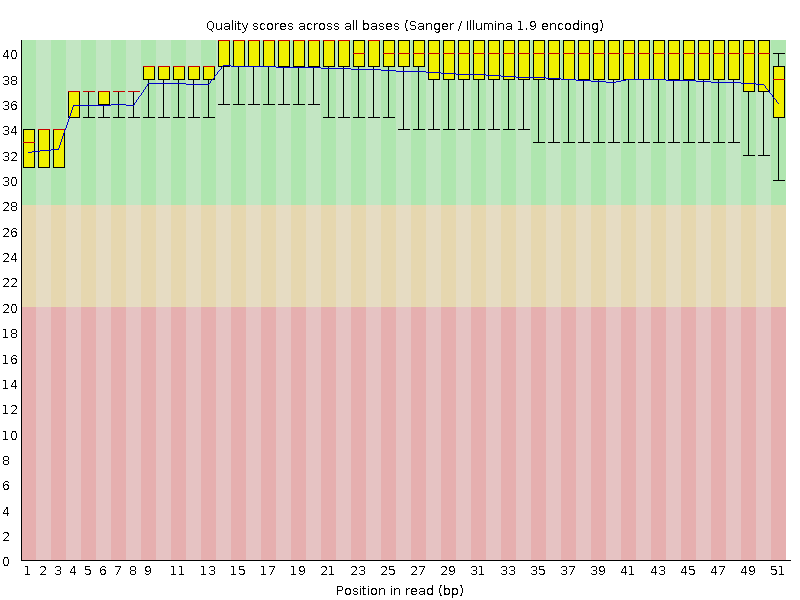
![[OK]](Icons/tick.png) Per sequence quality scores
Per sequence quality scores
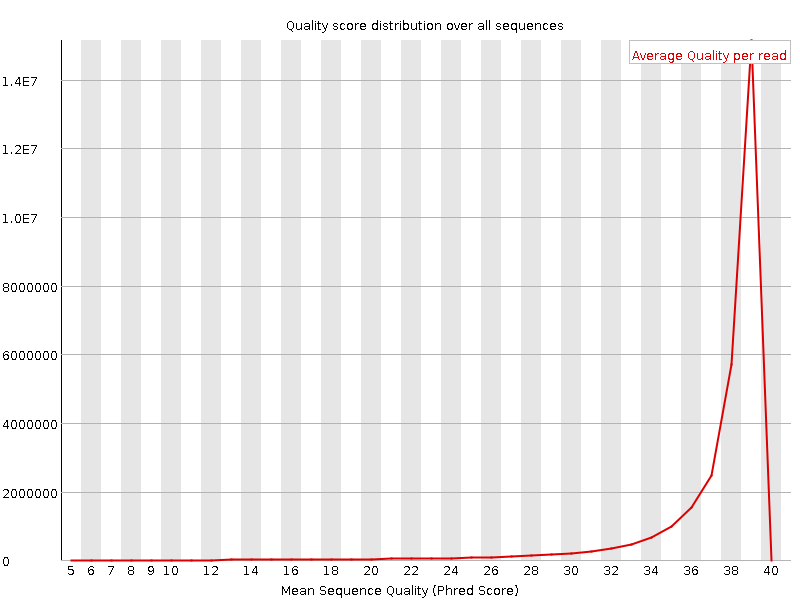
![[WARN]](Icons/warning.png) Per base sequence content
Per base sequence content

![[OK]](Icons/tick.png) Per base GC content
Per base GC content

![[OK]](Icons/tick.png) Per sequence GC content
Per sequence GC content

![[OK]](Icons/tick.png) Per base N content
Per base N content

![[OK]](Icons/tick.png) Sequence Length Distribution
Sequence Length Distribution
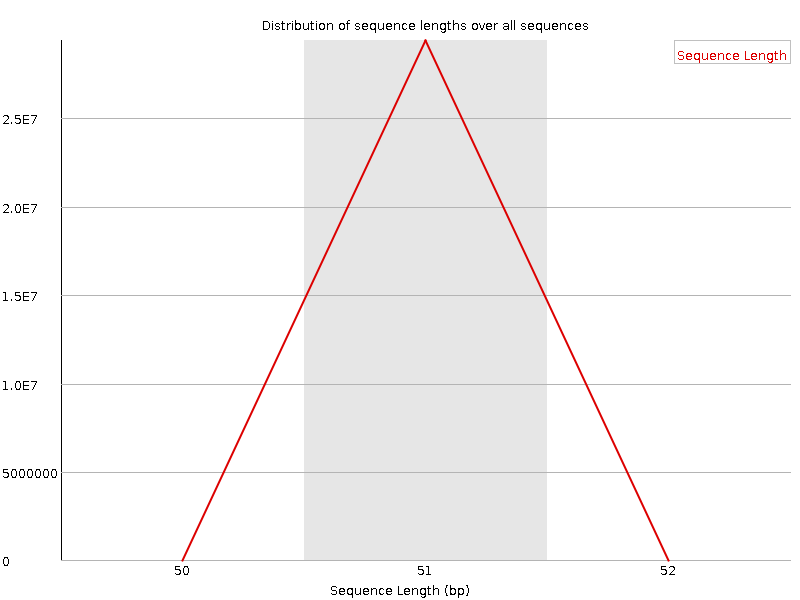
![[WARN]](Icons/warning.png) Sequence Duplication Levels
Sequence Duplication Levels
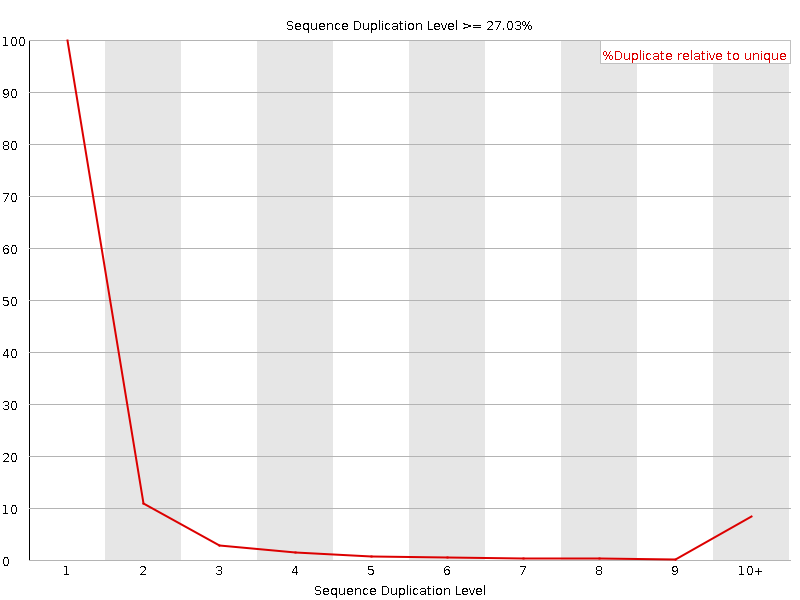
![[WARN]](Icons/warning.png) Overrepresented sequences
Overrepresented sequences
| Sequence | Count | Percentage | Possible Source |
|---|---|---|---|
| CTTTGTGTTTGATTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTT | 33061 | 0.11248923839569612 | No Hit |
![[WARN]](Icons/warning.png) Kmer Content
Kmer Content
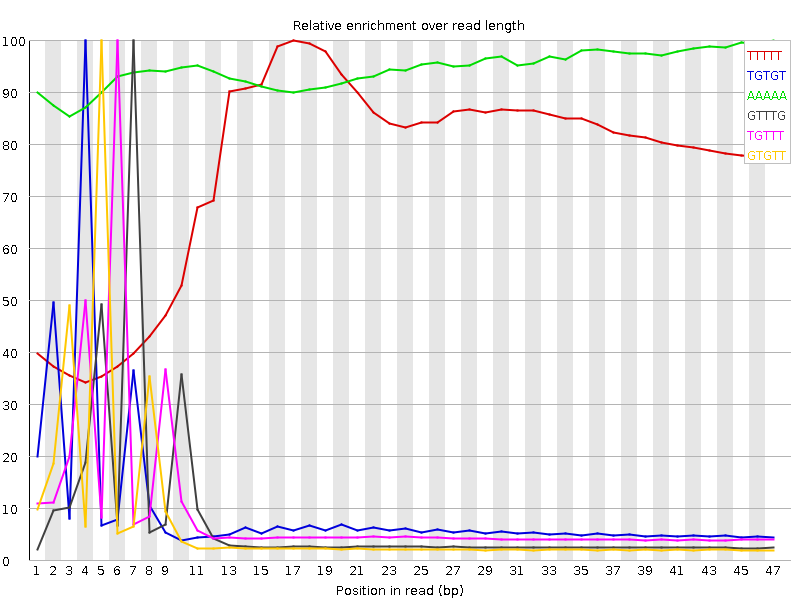
| Sequence | Count | Obs/Exp Overall | Obs/Exp Max | Max Obs/Exp Position |
|---|---|---|---|---|
| TTTTT | 13213270 | 4.2656226 | 5.644533 | 17 |
| TGTGT | 7195160 | 4.077572 | 42.57654 | 4 |
| AAAAA | 7511820 | 3.379186 | 3.5725029 | 47 |
| GTTTG | 5576400 | 3.1602032 | 42.13406 | 7 |
| TGTTT | 6827135 | 2.9201496 | 32.316326 | 6 |
| GTGTT | 5151530 | 2.9194255 | 41.992237 | 5 |
| TTTGA | 5958340 | 2.7233965 | 34.135765 | 8 |
| TTGTG | 4762965 | 2.6992214 | 42.05024 | 3 |
| CTTTG | 4388785 | 2.6951737 | 45.791046 | 1 |
| GGGGG | 2000025 | 2.6362102 | 6.24407 | 13 |
| GAGGG | 2249815 | 2.3917503 | 13.432961 | 11 |
| TTGAG | 3683225 | 2.2305315 | 26.834738 | 9 |
| TTTGT | 5164090 | 2.2088203 | 32.11286 | 2 |
| TGAGG | 2586425 | 2.0752692 | 16.473072 | 10 |
| AGGGG | 1628720 | 1.731472 | 7.0562005 | 12 |
| AACTT | 2868525 | 1.518254 | 14.259825 | 2 |
| TGAGC | 1720720 | 1.4961188 | 6.510825 | 10 |
| ACTTT | 2998915 | 1.4853576 | 13.023549 | 3 |
| TGAGA | 2206505 | 1.4279201 | 5.0663056 | 10 |
| TGAGT | 2146230 | 1.2997394 | 7.810207 | 10 |
| GAGTG | 1578185 | 1.266288 | 6.2968545 | 11 |
| TTGAT | 2499540 | 1.1424723 | 7.9535713 | 9 |
| TAACT | 2122045 | 1.1231569 | 14.017698 | 1 |
| TTGAC | 1596670 | 1.0477957 | 6.8774643 | 9 |