![[OK]](Icons/tick.png) Basic Statistics
Basic Statistics
| Measure | Value |
|---|---|
| Filename | 140701_JD_C_16_TGACCA_L004_R1_001.fastq.gz |
| File type | Conventional base calls |
| Encoding | Sanger / Illumina 1.9 |
| Total Sequences | 29390367 |
| Filtered Sequences | 0 |
| Sequence length | 51 |
| %GC | 42 |
![[OK]](Icons/tick.png) Per base sequence quality
Per base sequence quality
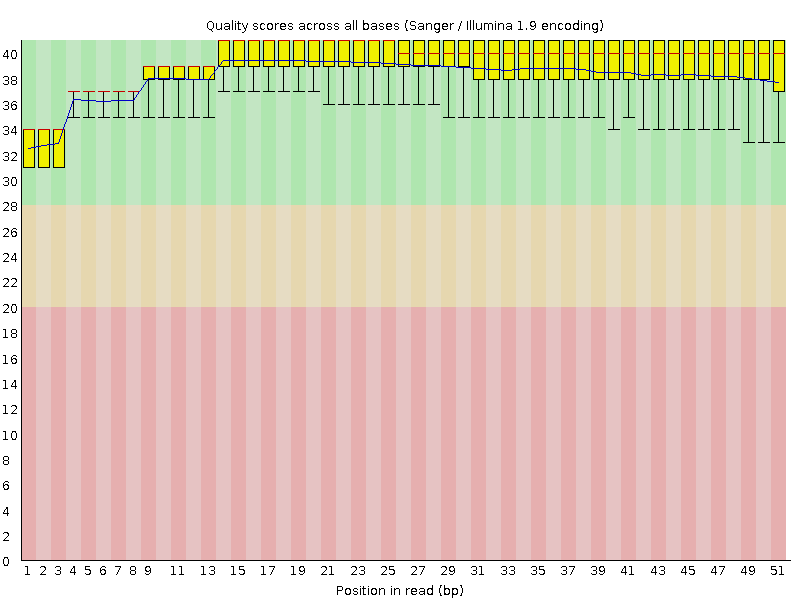
![[OK]](Icons/tick.png) Per sequence quality scores
Per sequence quality scores
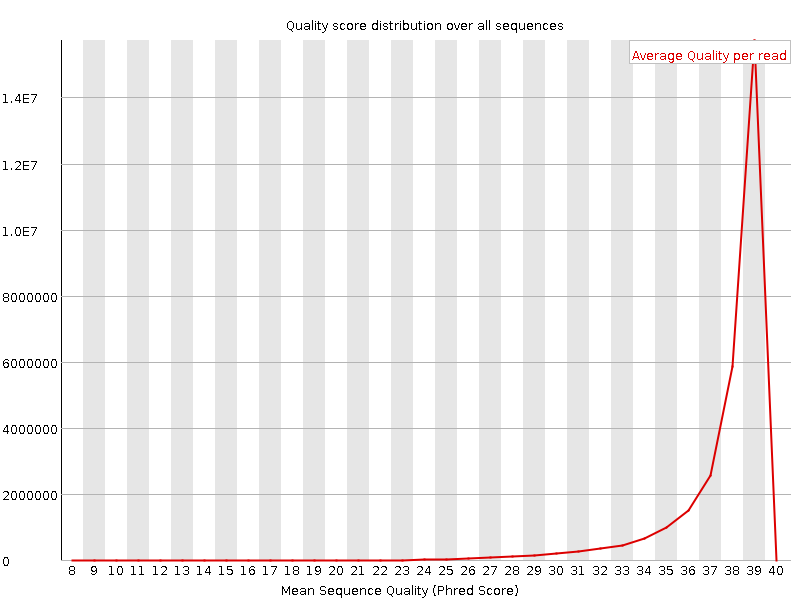
![[WARN]](Icons/warning.png) Per base sequence content
Per base sequence content

![[OK]](Icons/tick.png) Per base GC content
Per base GC content

![[OK]](Icons/tick.png) Per sequence GC content
Per sequence GC content

![[OK]](Icons/tick.png) Per base N content
Per base N content

![[OK]](Icons/tick.png) Sequence Length Distribution
Sequence Length Distribution
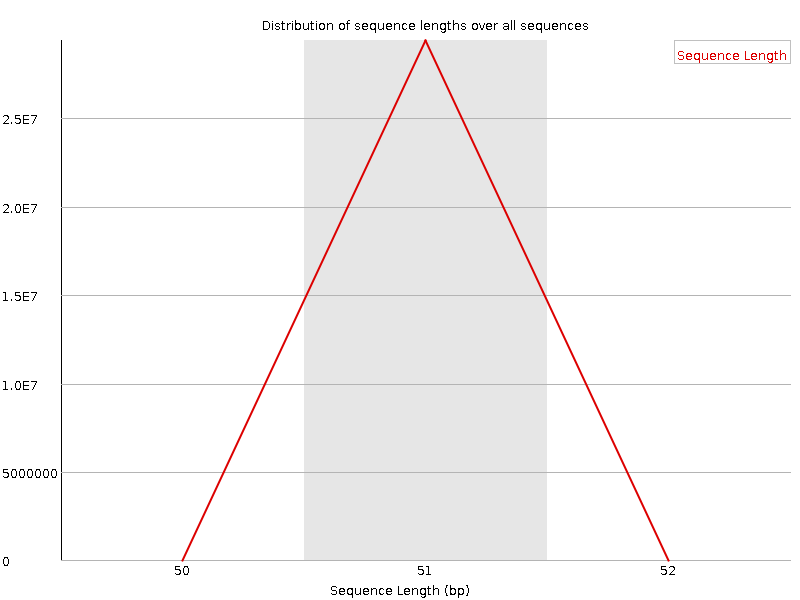
![[WARN]](Icons/warning.png) Sequence Duplication Levels
Sequence Duplication Levels
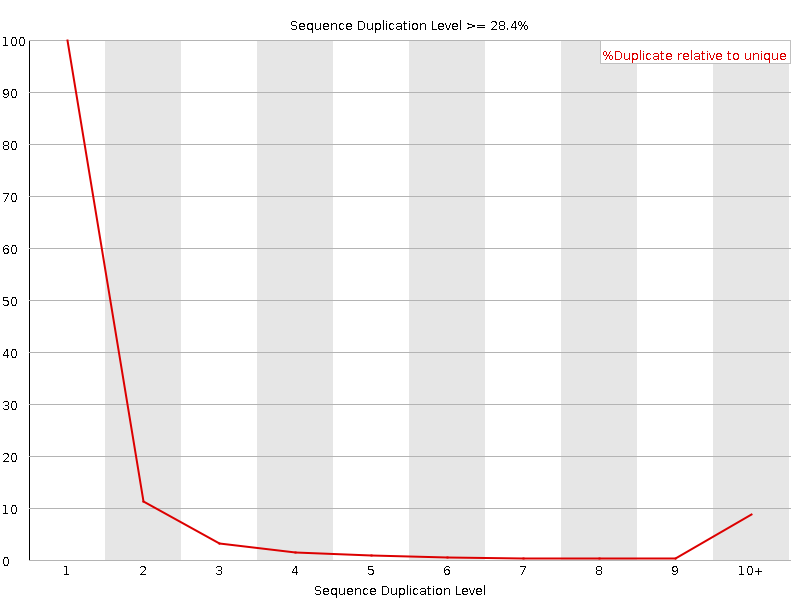
![[WARN]](Icons/warning.png) Overrepresented sequences
Overrepresented sequences
| Sequence | Count | Percentage | Possible Source |
|---|---|---|---|
| CTTTGTGTTTGATTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTT | 41361 | 0.14072978401392539 | No Hit |
![[WARN]](Icons/warning.png) Kmer Content
Kmer Content
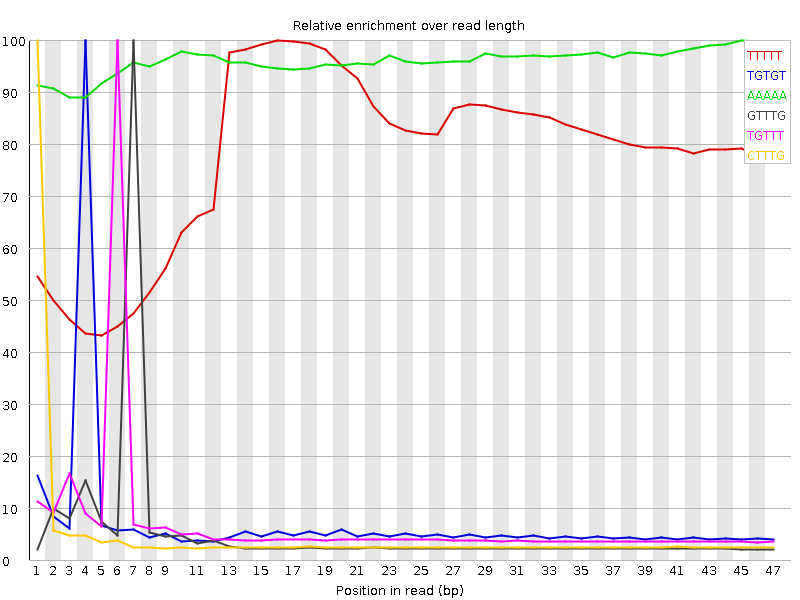
| Sequence | Count | Obs/Exp Overall | Obs/Exp Max | Max Obs/Exp Position |
|---|---|---|---|---|
| TTTTT | 12083735 | 4.0345116 | 5.178818 | 16 |
| TGTGT | 5825220 | 3.4907618 | 48.749527 | 4 |
| AAAAA | 7436355 | 3.1793666 | 3.3074615 | 45 |
| GTTTG | 4355935 | 2.6102931 | 48.07327 | 7 |
| TGTTT | 5609255 | 2.5090194 | 36.45368 | 6 |
| CTTTG | 3936040 | 2.4729514 | 50.45756 | 1 |
| GTGTT | 3909385 | 2.342698 | 47.966534 | 5 |
| TTTGA | 4784560 | 2.2486348 | 37.932205 | 8 |
| GAGGG | 1957880 | 2.2125351 | 14.703665 | 11 |
| TTGTG | 3609675 | 2.1630971 | 48.012054 | 3 |
| GGGGG | 1472145 | 2.1212196 | 5.9205713 | 13 |
| TTTGT | 4682100 | 2.094303 | 36.270355 | 2 |
| GAGAG | 2290875 | 2.0303674 | 5.0233054 | 11 |
| TGAGG | 2268500 | 1.9135238 | 18.598217 | 10 |
| TTGAG | 2976910 | 1.8743544 | 29.836739 | 9 |
| AGGGG | 1422540 | 1.6075653 | 7.718156 | 12 |
| TGAGC | 1619515 | 1.4322813 | 6.958151 | 10 |
| TGAGA | 2115010 | 1.39919 | 5.4884768 | 10 |
| TGAGT | 1991250 | 1.2537527 | 8.7317705 | 10 |
| GAGTG | 1456535 | 1.2286154 | 6.935703 | 11 |
| TTGAT | 2250100 | 1.0574961 | 8.994572 | 9 |
| TTGAC | 1482175 | 0.978439 | 7.5758476 | 9 |