![[OK]](Icons/tick.png) Basic Statistics
Basic Statistics
| Measure | Value |
|---|---|
| Filename | 140701_JD_B_3_ACAGTG_L005_R2_001.fastq.gz |
| File type | Conventional base calls |
| Encoding | Sanger / Illumina 1.9 |
| Total Sequences | 29275164 |
| Filtered Sequences | 0 |
| Sequence length | 51 |
| %GC | 43 |
![[OK]](Icons/tick.png) Per base sequence quality
Per base sequence quality
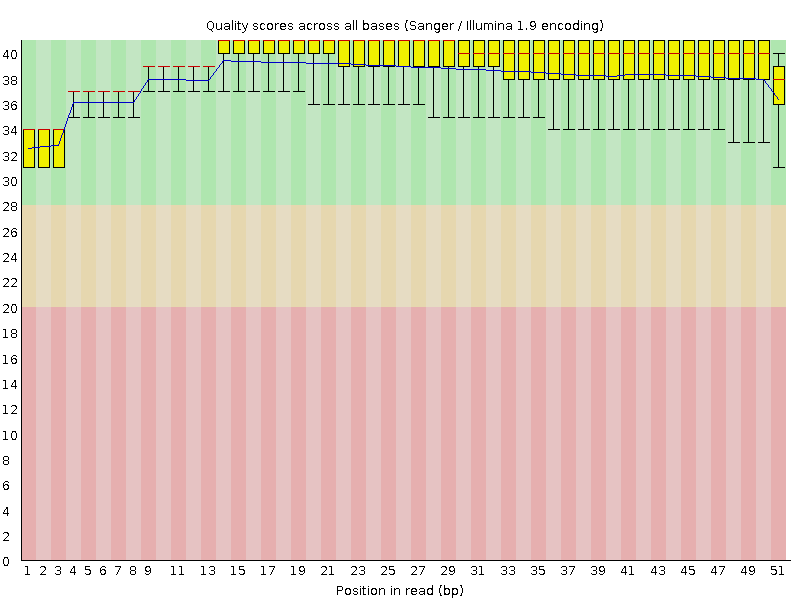
![[OK]](Icons/tick.png) Per sequence quality scores
Per sequence quality scores
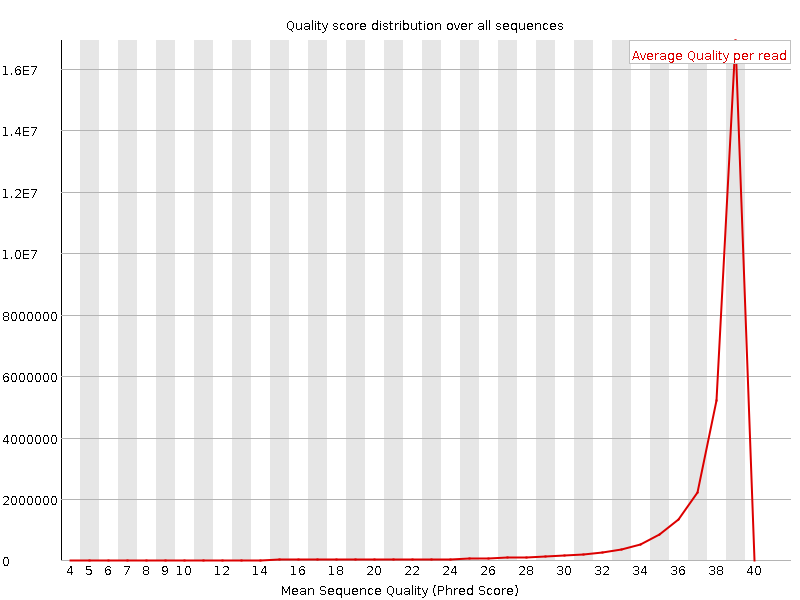
![[WARN]](Icons/warning.png) Per base sequence content
Per base sequence content

![[OK]](Icons/tick.png) Per base GC content
Per base GC content

![[OK]](Icons/tick.png) Per sequence GC content
Per sequence GC content

![[OK]](Icons/tick.png) Per base N content
Per base N content

![[OK]](Icons/tick.png) Sequence Length Distribution
Sequence Length Distribution
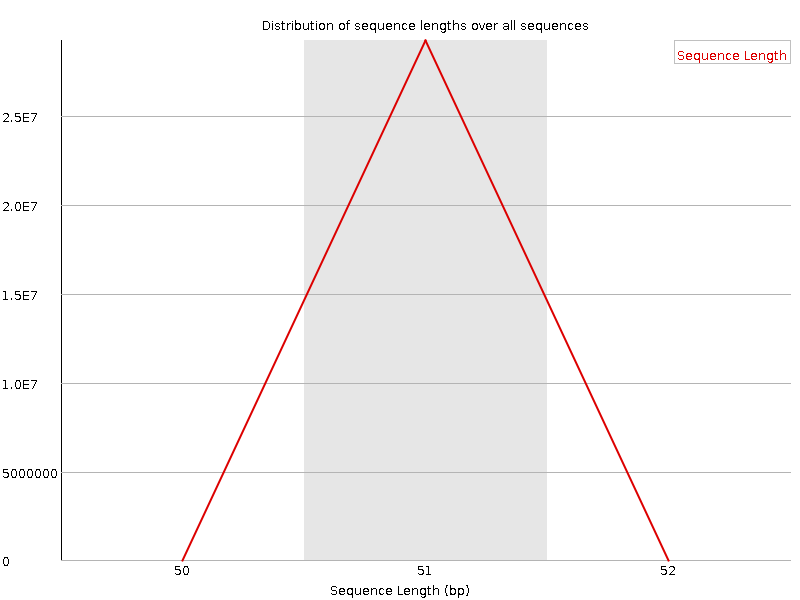
![[WARN]](Icons/warning.png) Sequence Duplication Levels
Sequence Duplication Levels
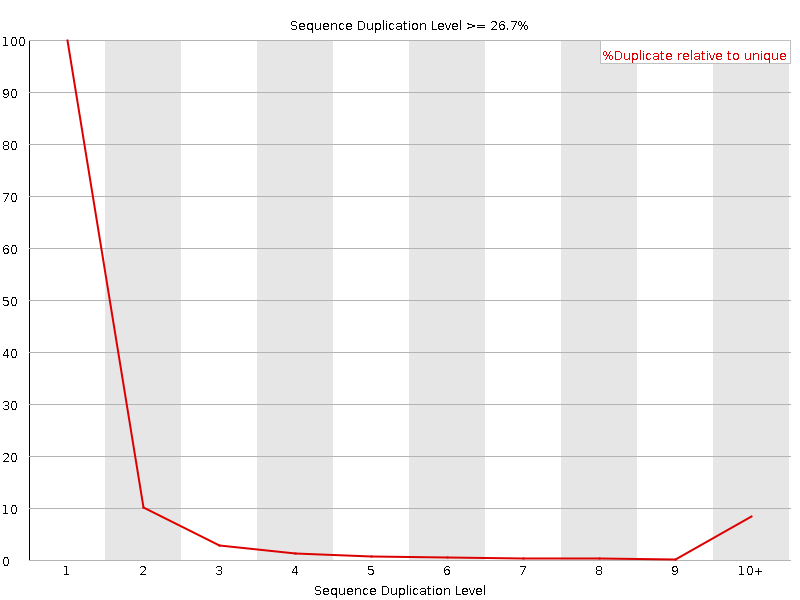
![[WARN]](Icons/warning.png) Overrepresented sequences
Overrepresented sequences
| Sequence | Count | Percentage | Possible Source |
|---|---|---|---|
| CTTTGTGTTTGATTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTT | 48459 | 0.16552938866542302 | No Hit |
| TTGTGTTTGATTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTT | 34493 | 0.11782342192856717 | No Hit |
![[WARN]](Icons/warning.png) Kmer Content
Kmer Content
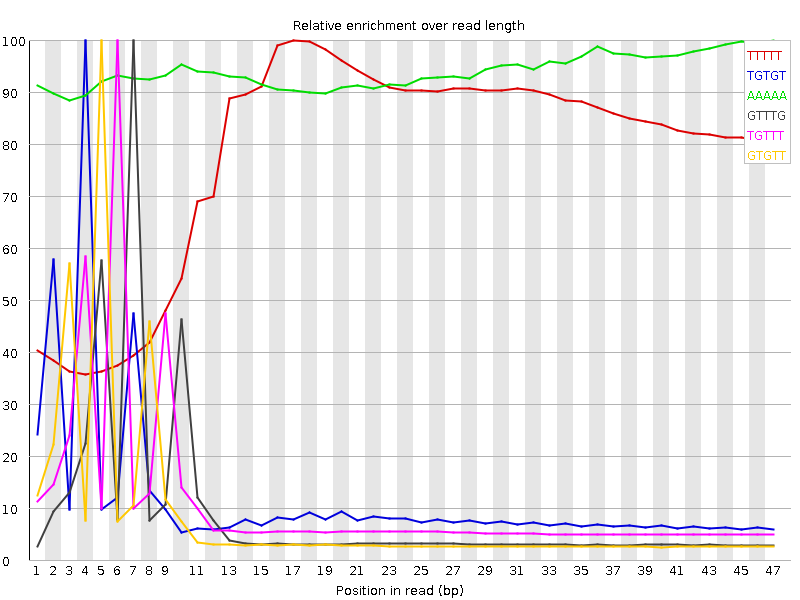
| Sequence | Count | Obs/Exp Overall | Obs/Exp Max | Max Obs/Exp Position |
|---|---|---|---|---|
| TTTTT | 15127995 | 5.240991 | 6.710867 | 17 |
| TGTGT | 6838425 | 3.9682212 | 33.406887 | 4 |
| AAAAA | 7551800 | 3.5464687 | 3.7683966 | 47 |
| GTTTG | 4939685 | 2.8664148 | 32.72591 | 7 |
| TGTTT | 6250565 | 2.8025618 | 25.95726 | 6 |
| GTGTT | 4695645 | 2.7248025 | 32.822052 | 5 |
| TTTGA | 5367030 | 2.5573647 | 27.087662 | 8 |
| CTTTG | 4047300 | 2.524663 | 35.443016 | 1 |
| TTGTG | 4328960 | 2.5120215 | 32.807 | 3 |
| GGGGG | 1977055 | 2.486967 | 5.3467803 | 13 |
| GAGGG | 2248345 | 2.322379 | 10.576637 | 11 |
| TTTGT | 4753505 | 2.131326 | 25.738682 | 2 |
| TTGAG | 3368185 | 2.077105 | 20.655539 | 9 |
| TGAGG | 2495875 | 1.9919991 | 12.698647 | 10 |
| AGGGG | 1658920 | 1.7135454 | 5.9576216 | 12 |
| AACTT | 2852565 | 1.5527987 | 14.58747 | 2 |
| ACTTT | 2987265 | 1.5301383 | 13.340598 | 3 |
| TGAGC | 1749535 | 1.5010227 | 5.2169113 | 10 |
| GAGTG | 1576005 | 1.2578356 | 5.0182834 | 11 |
| TGAGT | 2023405 | 1.247801 | 6.104101 | 10 |
| TAACT | 2092805 | 1.139222 | 14.295946 | 1 |
| TTGAT | 2338830 | 1.1144415 | 6.5352187 | 9 |
| TTGAC | 1465860 | 0.9717472 | 5.4746118 | 9 |