![[OK]](Icons/tick.png) Basic Statistics
Basic Statistics
| Measure | Value |
|---|---|
| Filename | 140701_JD_B_3_ACAGTG_L005_R1_001.fastq.gz |
| File type | Conventional base calls |
| Encoding | Sanger / Illumina 1.9 |
| Total Sequences | 29275164 |
| Filtered Sequences | 0 |
| Sequence length | 51 |
| %GC | 43 |
![[OK]](Icons/tick.png) Per base sequence quality
Per base sequence quality
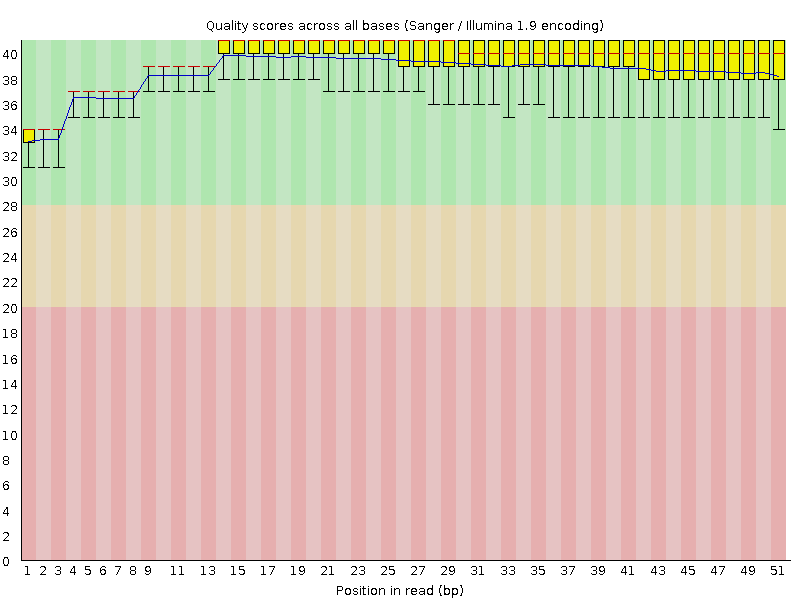
![[OK]](Icons/tick.png) Per sequence quality scores
Per sequence quality scores
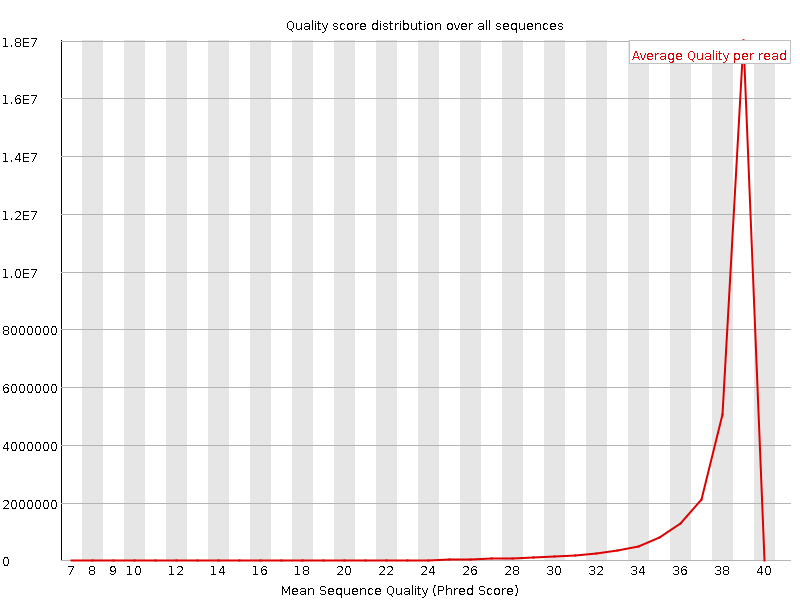
![[WARN]](Icons/warning.png) Per base sequence content
Per base sequence content

![[OK]](Icons/tick.png) Per base GC content
Per base GC content

![[OK]](Icons/tick.png) Per sequence GC content
Per sequence GC content

![[OK]](Icons/tick.png) Per base N content
Per base N content

![[OK]](Icons/tick.png) Sequence Length Distribution
Sequence Length Distribution
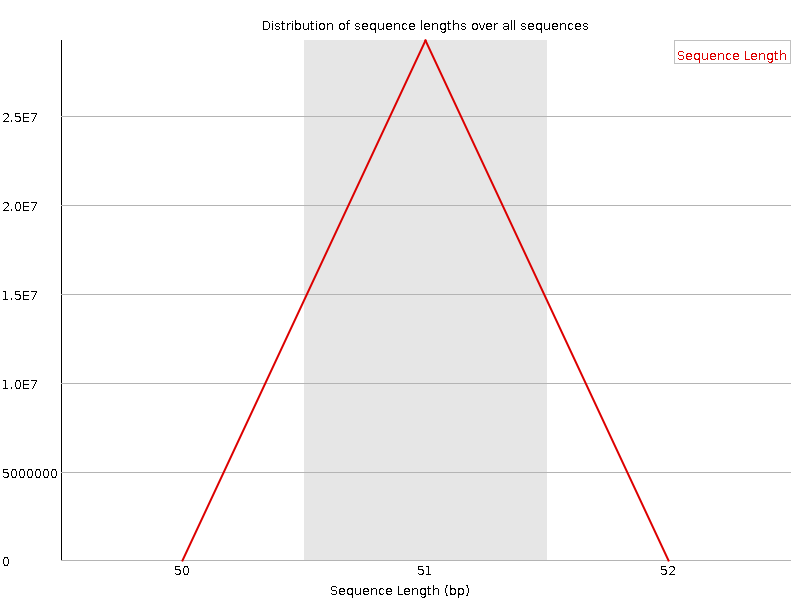
![[WARN]](Icons/warning.png) Sequence Duplication Levels
Sequence Duplication Levels
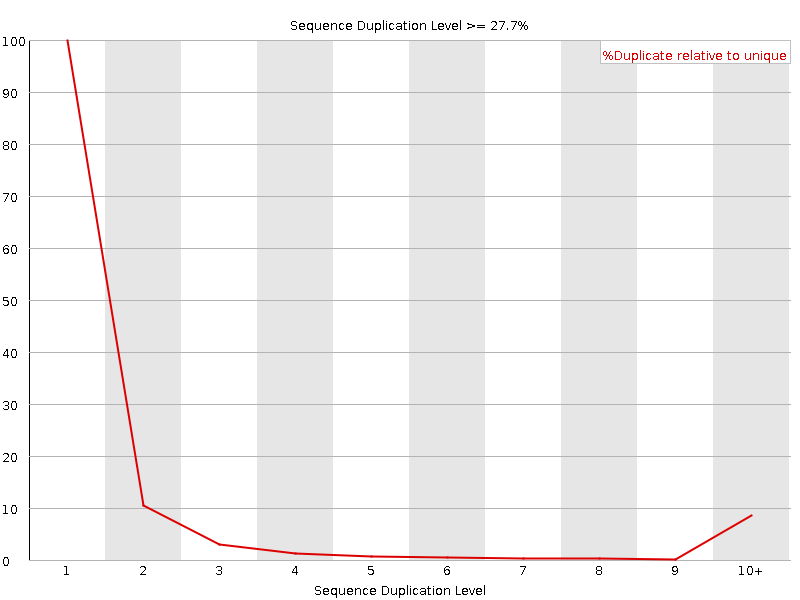
![[WARN]](Icons/warning.png) Overrepresented sequences
Overrepresented sequences
| Sequence | Count | Percentage | Possible Source |
|---|---|---|---|
| CTTTGTGTTTGATTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTT | 58490 | 0.19979392771292417 | No Hit |
![[WARN]](Icons/warning.png) Kmer Content
Kmer Content
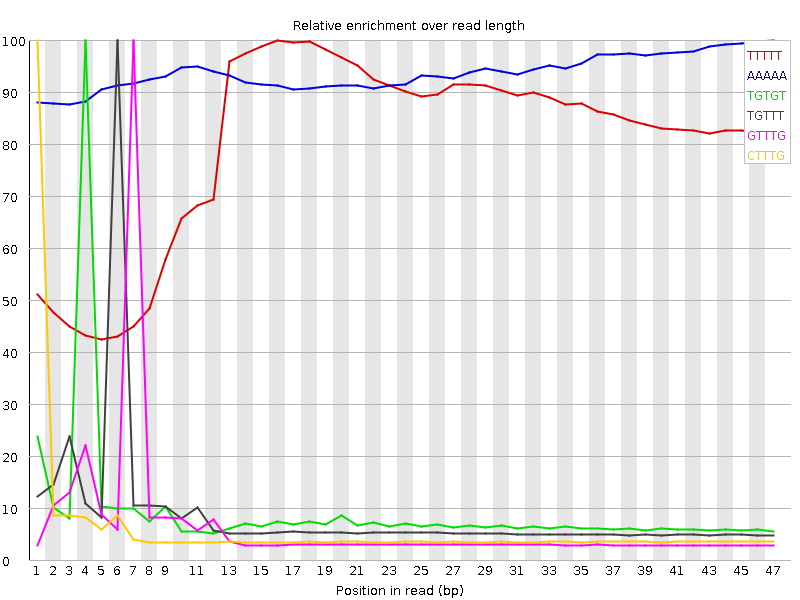
| Sequence | Count | Obs/Exp Overall | Obs/Exp Max | Max Obs/Exp Position |
|---|---|---|---|---|
| TTTTT | 13713115 | 4.9768157 | 6.1944494 | 16 |
| AAAAA | 7574750 | 3.4290724 | 3.6544847 | 47 |
| TGTGT | 5427015 | 3.3205366 | 35.608963 | 4 |
| TGTTT | 5094470 | 2.4006584 | 27.521038 | 6 |
| GTTTG | 3771435 | 2.307565 | 34.84878 | 7 |
| CTTTG | 3529480 | 2.254184 | 36.431866 | 1 |
| GTGTT | 3508235 | 2.146525 | 34.82546 | 5 |
| GAGGG | 1972425 | 2.1265626 | 10.547272 | 11 |
| TTTGA | 4229500 | 2.0831423 | 28.305283 | 8 |
| TTTGT | 4207440 | 1.9826647 | 27.216198 | 2 |
| TTGTG | 3174770 | 1.9424934 | 34.813103 | 3 |
| TGAGG | 2182325 | 1.8120961 | 13.104851 | 10 |
| TTGAG | 2673205 | 1.7095356 | 21.161634 | 9 |
| AGGGG | 1472455 | 1.5875218 | 5.9487576 | 12 |
| TGAGC | 1661760 | 1.4403281 | 5.2454877 | 10 |
| GAGTG | 1464465 | 1.21602 | 5.007458 | 11 |
| TGAGT | 1864670 | 1.192471 | 6.2322745 | 10 |
| TTGAT | 2106380 | 1.0374488 | 7.194327 | 9 |
| TTGAC | 1329925 | 0.887778 | 5.6463776 | 9 |